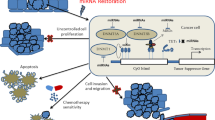
Abstract
Endometrial cancer is the most common gynecological cancer in the United States. We wanted to identify epigenetic aberrations involving microRNAs (miRNAs), whose genes become hypermethylated in endometrial primary tumors. By integrating known miRNA sequences from the miRNA database (miRBase) with DNA methylation data from methyl-CpG-capture sequencing, we identified 111 differentially methylated regions (DMRs) associated with CpG islands (CGIs) and miRNAs. Among them, 22 DMRs related to 29 miRNAs and within 8 kb of CGIs were hypermethylated in endometrial tumors but not in normal endometrium. miR-137 was further validated in additional endometrial primary tumors. Hypermethylation of miR-137 was found in both endometrioid and serous endometrial cancer (P < 0.01), and it led to the loss of miR-137 expression. Treating hypermethylated endometrial cancer cells with epigenetic inhibitors reactivated miR-137. Moreover, genetic overexpression of miR-137 suppressed cancer cell proliferation and colony formation in vitro. When transfected cancer cells were implanted into nude mice, the cells that overexpressed miR-137 grew more slowly and formed smaller tumors (P < 0.05) than vector transfectants. Histologically, xenograft tumors from cancer cells expressing miR-137 were less proliferative (P < 0.05), partly due to inhibition of EZH2 and LSD1 expression (P < 0.01) in both the transfected cancer cells and tumors. Reporter assays indicated that miR-137 targets EZH2 and LSD1. These results suggest that miR-137 is a tumor suppressor that is repressed in endometrial cancer because the promoter of its gene becomes hypermethylated.
Similar content being viewed by others
Introduction
Endometrial cancer is the fourth most common malignancy in women and the most common gynecologic cancer in the United States [1, 2]. The disease can be broadly classified into two subtypes, endometrioid endometrial cancer or EEC (type I) and non-EEC (type II), which has primarily serous histology [3]. In EEC tumors, aberrant phosphoinositide 3-kinase (PI3K) signaling is the most commonly activated pathway. Activation of PI3K signaling is frequently due to loss of PTEN protein or mutations in PIK3CA, PIK3R1, PIK3R2 (genes coding for catalytic or regulatory subunits of P13K), or AKT1 [2, 4]. In non-EEC tumors, mutations in TP53, PIK3CA, and PPP2R1A are the drivers for oncogenesis [2, 5, 6].
We reported that in addition to well-known oncogenic events (such as genetic amplification, mutation, and translocation), dysregulation of miRNAs can lead to oncogenic activation [7, 8]. miRNAs are a class of small noncoding RNAs (18–25 nucleotides) that can target certain mRNAs by pairing imperfectly at the 3′-end of the untranslated region (UTR). As a result, the mRNA is degraded or not translated [9]. For example, miRNAs can silence certain protooncogene mRNAs to prevent unwanted cell division, thereby acting as tumor suppressors [10].
In contrast, dysregulation of miRNAs through epigenetic mechanisms contributes to tumorigenesis. One well-studied epigenetic phenomenon is DNA methylation, frequently observed in the promoter CpG island regions of genes [11]. In endometrial cancer, we found that repression of miR-129-2 or miR-203 by DNA hypermethylation in the promoters of their genes, correlated with overexpression of an oncogene, SOX4, a transcription factor that normally helps regulate embryonic development [11, 12]. Re-expression of miR-129 or miR-203 by knockin or pharmacologic treatments partially re-suppressed SOX4 expression.
The Cancer Genome Atlas (TCGA) conducted a large-scale survey of DNA methylation, using Illumina Infinium DNA methylation arrays [2], a microarray-based methodology. By coupling methyl-CpG-capture with next-generation sequencing, we identified 2302 loci in 67 primary endometrial tumors that were hypermethylated when compared with 10 control samples [13]. Those regions were found in transcriptional start sites (TSSs), CpG island cores, flanking regions of TSSs, or so-called CpG shores next to intergenic and intragenic CpG islands and non-CpG island promoters. Pathways associated with many of those loci relate to aggressive growth and metastasis, and include factors such as the EFG receptor (EGFR), mitogen-activated protein kinase (MAPK), Wnt, and gap junction. In this study, we integrated methyl-CpG-capture sequencing with miRNA database searches to identify hypermethylated miRNAs in endometrial primary tumors [13]. We further characterized one miRNA, miR-137, that has not been investigated in endometrial cancer. We demonstrated that miR-137 is hypermethylated in human endometrial tumors and confirmed that it is a tumor suppressor that acts through epigenetic silencing.
Materials and methods
Patient sample collection
Primary tumor specimens were obtained through the Cooperative Human Tissue Network (CHTN) following approval by the Medical College of Wisconsin (MCW) Institutional Review Board. Clinical endometrial tumor subtype, staging, and tumor grade were assigned on the basis of the International Federation of Gynecology and Obstetrics (FIGO) 1988 criteria and summarized in Supplementary Table 1. Tumor specimens had high neoplastic cellularity (mean 89.2%; median 90%), whereas normal tissue displayed no malignant portions under the microscope. All the cancer samples tested were primary gynecological tumors. Genomic DNA was isolated by using the AllPrep DNA/RNA Mini Kit (Qiagen, Valencia CA) and following the manufacturer’s protocol. DNA methylation by methyl-CpG-capture sequencing has been described in our publications [13, 14]. Expression and promoter methylation of miR-137 were acquired from TCGA’s miRNA sequences, and Illumina Infinium DNA methylation was described previously [2].
Cell culture
Endometrial normal cells (EM-E6/E7/TERT) and cancer cells (AN3CA, HEC1A, KLE, RL-95-2) were kindly provided by Dr. Paul Goodfellow (The Ohio State University). Hec50co and Ishikawa H were kindly gifted by Dr. Kimberly K. Leslie (University of Iowa) [8]. HEC1B, EFE-184, MFE-280, and MFE-296 were purchased from ATCC (Manassas, VA) or the DSMZ German Collection of Microorganisms and Cell Cultures (Braunschweig, Germany). All cell lines were cultured in a 37 °C incubator with 5% CO2, according to the suppliers’ instructions, and were passaged for no more than 6 months after receipt or thawing. Cell line authentication was performed by IDEXX (Westbrook, ME), which utilizes short tandem repeat (STR) profiling. To assess the role of DNA methylation in miR-137 expression, endometrial cancer cells were treated with 5-aza-2′-deoxycytidine (DAC; 5 μmol/L) for 48 h, with or without trichostatin A (TSA, 0.5 μmol/L) (Sigma-Aldrich, St. Louis MO) for 24 h.
Reverse transcription and quantitative PCR (RT-qPCR)
We performed reverse transcription on total RNA (1 µg), using iScript RT Supermix (Bio-Rad, Hercules, CA). PCR was performed as described previously [7]. SYBR assays used Pre-/Pri-miR-137 and U6 primers, as published previously [15]. Mature miR-137 and U6 primers were obtained from Thermo Fisher Scientific using TaqMan assay kits. The relative expression of a gene in cells was determined by comparing the threshold cycle (Ct) of the gene against the Ct of a housekeeping gene, U6.
Western blotting
Western blot analysis was performed as described previously [7, 13]. Briefly, 15 μg of protein lysate was loaded onto 4–20% Mini-PROTEAN TGX Gel (Bio-Rad) and transferred to polyvinylidene difluoride (PVDF) membranes. After blocking the membranes with bovine serum albumin and incubating them with primary and secondary antibodies, we exposed them to ECL-plus (GE Healthcare) and visualized the protein bands with the ChemiDoc imaging system (Bio-Rad). Antibodies against EZH2, LSD1, and β-actin were obtained from Abcam (Cambridge, MA) or Cell Signaling Technology (Danvers, MA).
Combined bisulfite restriction analysis
The combined bisulfite restriction analysis (COBRA) assay was described previously [7, 12, 13]. Briefly, genomic DNA was treated with bisulfite using the EZ DNA Methylation kit (Zymo Research, Irvine CA), amplified by PCR, and digested with the methylation-sensitive enzyme AciI (New England Biolabs, Ipswich MA). Digested and non-digested PCR products were resolved on 2% agarose gels and stained with ethidium bromide. Smaller DNA fragments digested by AciI were identified as “methylated” in a given sample. COBRA primers are described in Supplementary Table 2.
Bisulfite pyrosequencing
DNA methylation was measured using the PyroMark MD system (Qiagen) as described previously [12]. Oligonucleotide primers for the nearby miR-137 CpG island published previously [16] were purchased from Thermo Fisher Scientific. Primer sequences for the distal CpG island of miR-137 in the P1 and P2 regions are given in Supplementary Table 2. Methylation was quantified using the software provided (Qiagen).
Cell proliferation and colony formation
Cell proliferation was monitored using CellTiter 96 AQueous One Solution (Promega, Madison, WI) as described previously [7]. Endometrial cancer cells (2000 per well) transfected with miR-137 or empty vector were seeded into 96-well plates. Cell proliferation was documented at 3, 48, or 96 h after seeding. Colony formation was reported previously [17]. Transfected cells (200/well) were seeded into six-well plates. Two weeks later, colonies were fixed with 4% formaldehyde, stained with crystal violet (0.5% w/v), and counted under a microscope.
Knocking in miR-137, LSD1, and EZH2
HEC1A and Ishikawa H cells (2 × 105) were seeded into six-well plates with transfected plasmids of human miR-137 or empty vector (2 μg/well; Origene, Rockville MD), using the TurboFectin Transfection Reagent (Origene) according to the manufacturer’s instructions. Stable clones were selected by G418 (1 mg/ml) after transfection. miR-137 stable cancer cells were transfected with EZH2, LSD1, and their relative empty plasmids (OriGene and Addgene, Cambridge, MA) by TurboFectin. Proliferation of the transfected cells was measured, and cell lysates were isolated for mRNA and protein assays.
3′-UTR luciferase reporter assay
Cancer cells were seeded into 96-well white plates and co-transfected, using Lipofectamine RNAiMAX with miR-137 mimic or anti-miR-137 (15 ng/well) (Thermo Fisher Scientific, Waltham, MA) along with pMirTarget 3′-UTR plasmids of EZH2 and LSD1 (7.5 ng/well, Origene). Forty-eight hours after transfection, luciferase activity was measured using the Steady-Glo Luciferase kit (Promega) according to the manufacturer’s instructions. The fold-change values were calculated back to the cells that had been co-transfected with the negative control miRNA and the control plasmid.
Xenograft tumors
Tumors were generated by subcutaneous injection of cancer cells (2 × 106) into 6-week-old female athymic nude mice (n = 5, Jackson Laboratories, Bar Harbor, ME) as described previously [18]. All mice were housed in identical conditions in a pathogen-free animal facility under 14-h light/10-h dark cycles. Food and water were supplied ad libitum. Experimental procedures were conducted in accordance with the guidelines of the MCW Institutional Animal Care and Use Committee (IACUC). The mice were monitored for tumor development twice weekly, and tumor volume was calculated from external caliper measurements: volume = ½ (L × W2), where L is the greatest longitudinal diameter and W is the greatest transverse diameter. When tumors reached 2 cm3, mice were euthanized and tumors were collected (fresh-freezing and formalin fixation) for molecular analyses and hematoxylin and eosin (H&E) staining.
Immunohistochemical staining and scoring
Paraffin-embedded slides were placed in a 60 °C oven for 1 h, cooled, deparaffinized, and rehydrated through xylene and graded ethanol solutions to water. All slides were quenched for 5 min in a 3% hydrogen peroxide aqueous solution to inhibit endogenous peroxidases. The slides were then stained, using a Dako Autostainer (Carpinteria, CA). Primary antibodies and antibody dilutions were as follows: Ki-67 (1:400, Dako), cleaved caspase-3 (Asp175) (1:200, Cell Signaling), EZH2 (1:50), and LSD1 (1:100). Following a 1-h incubation at room temperature, the slides were stained with their respective secondary antibodies, counterstained with hematoxylin, dehydrated through a graded ethanol series, and then cover-slipped. Five fields (at least 400 cells/field) per tumor were selected at random to determine the percentages of positively stained cells relative to the total cell count.
Statistical analyses
mRNA expression, methylation levels, UTR reporters, colony number and size, cell and tumor growth, and tumor weights were compared using Student’s t-test or Fisher’s exact test. A P-value of <0.05 was considered significant. All tests were two-sided, and all statistical analyses were performed using GraphPad Prism 7 software (GraphPad Software, La Jolla CA).
Results
Hypermethylated miRNA identified in human endometrial tumors
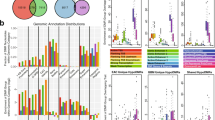
To identify miRNAs that associate with DNA hypermethylation in human endometrial tumors, we screened global methylation profiles of 705 miRNAs from the miRBase 13.0 database (Fig. 1). The number screened was based on our previous sequencing data from 10 normal control and 67 EEC primary tumors [13, 14]. When we limited our search for differentially methylated regions (DMRs) to within CpG islands, we found 111 DMRs. In a further analysis, we found 44 DMRs that could distinguish tumors from normal controls. Among these, 29 miRNAs were associated with 22 DMR CpG islands within an 8-kb region (Fig. 1a and Supplementary Figure 1). These miRNAs were miR-9-1, 9-3, 34b, 34c, 124-1, 124-2, 124-3, 129-2, 132, 212, 135b, 182, 96, 183, 181c, 181d, 195, 497, 196a-1, 203, 935, 1179, 7-2, 1224, 1247, 1469, and 196a-2. In addition, we found miR-9-2 and 199a-2 to be hypermethylated in EEC tumors, but those miRNAs were not associated with CpG islands (Supplementary Figure 1).
DNA methylation of miR-137 in primary endometrial tissues. a Outline for identifying hypermethylated miRNA in endometrial cancer. b Methylation profiles of 10 normal endometrial tissues (N) and 67 primary tumors (Ca) created after methyl-capture sequencing analysis. Red and white dashed-line squares represent methylated and unmethylated regions, respectively. Green bar: the nearby CpG island
miR-137 is hypermethylated and loses gene expression in human endometrial tumors
One above mentioned miRNA, miR-137, had not been investigated in endometrial cancer previously. We limited our screening to identify DNA methylation associated with miRNA to 8 kb upstream of known miRNA sequences. When we extended CpG island screening to 100 kb, we found two CpG islands within 10 kb of miR-137 (termed the distal and nearby CpG islands, Supplementary Figure 2a). Using bisulfite pyrosequencing analyses and two sets of primers (P1 and P2, Supplementary Figure 2b), we first assessed DNA methylation levels in the distal CpG island in the samples from our clinical cohort (Supplementary Table 1). Interestingly, the P1 region was hypermethylated in both normal controls and primary tumors (Supplementary Figure 2b). In contrast, the P2 region was considerably less methylated in both groups. We found little or no difference in both regions between 10 paired endometrial tissues and different subtypes of primary tumors (Supplementary Figure 2). These results suggest that DNA methylation of the distal CpG island of miR-137 cannot distinguish normal endometrium from cancer specimens.
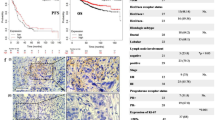
We next determined whether DNA methylation of the nearby CpG island (Fig. 2a) might distinguish normal from tumor tissue. Using two controls known methylation levels (100% and 0%), we premixed and generated gradient standards. Bisulfite pyrosequencing methylation analysis was well optimized to achieve good associations between expected methylation levels and those measured using known samples (gradient standards) (Supplementary Figure 3a). We assayed and identified hypermethylation of miR-137 in the nearby CpG island of our EEC tumors, detecting higher levels than in paired normal adjacent tissues (P < 0.05, Fig. 2b). We also found that miR-137 was hypermethylated in our primary serous endometrial tumors (P < 0.01, Fig. 2c), and the degree of hypermethylation did not differ significantly from that in the EEC samples. DNA methylation levels were less intensive in cervical and ovary tumors (Supplementary Figure 3b). Using TCGA methylation data obtained with Humanmethylation450 [2], we found that miR-137 was hypermethylated in tumors compared with their paired counterparts (P < 0.001, Fig. 2d). We categorized samples as normal, EEC tumors, or serous tumors based on histology types, and then performed group comparisons. We found miR-137 hypermethylation in both EEC and serous primary tumors compared with normal endometria (P < 0.001, Fig. 2e). In agreement with our DNA methylation data, miR-137 expression was lost and silenced in most endometrial tumors (P < 0.01), including both EEC (P < 0.001) and serous (P < 0.05) tumor subtypes (Fig. 2f). In a further analysis of DNA methylation data from TCGA, lower levels of miR-137 methylation correlated with microsatellite instability (MSI, P = 0.01, Supplementary Figure 4). However, miR-137 methylation levels did not associate with race, body mass index (BMI), disease recurrence, grade, stage, MLH1 methylation, or overall and disease-free survival (Supplementary Figure 4). Taken together, these DNA methylation data unambiguously suggest that miR-137 is hypermethylated in endometrial cancer, including in both EEC and serous subtypes of tumors, leading to loss of miR-137 expression.
miR-137 is hypermethylated and shows loss of expression in human primary endometrial tumors. a Genomic map of the CpG island nearby miR-137 (black bar), COBRA and bisulfite pyrosequencing (Pyro) amplicons and TCGA methylation probes. CpG sites are lines under bars. b Dot plot showing miR-137 methylation levels in 10 pairs of endometrioid endometrial cancer (EEC) and normal adjacent tissues (NAT) quantified by bisulfite pyrosequencing. c DNA methylation was determined by bisulfite pyrosequencing in human endometrial tissues. NAT: normal adjacent tissue, Serous: serous endometrial tumor. Each dot represents one specimen. Each horizontal line indicates the mean of methylation within each group. DNA methylation of miR-137 by probes in paired samples (d) and subtypes of endometrial tumors (e) in TCGA cohort. Pair (n = 33); N: normal endometrium (n = 13); EEC (n = 312); serous (n = 98). f Expression of miR-137 in TCGA endometrial cohort. Gray portions: miR-137 not expressed; white portions: miR-137 expressed. Number in each bar indicates sample size. *: P < 0.05; **: P < 0.01; ***: P < 0.001
miR-137 is hypermethylated and reactivated by epigenetic inhibitors in endometrial cancer cells
Using COBRA assays, we first examined DNA methylation in endometrial cell lines in the distal CpG island, using two sets of primers (P1 and P2, Supplementary Figure 2a). Interestingly, the P1 region was methylated in a normal endometrial cell line (EM-E6/E7/TERT) and in 11 endometrial cancer cell lines. In contrast, the P2 region was substantially less methylated than in primary tissues (Supplementary Figures 5a-b). We next assessed whether the miR-137 gene in the nearby CpG island was methylated in either tissue type. Both COBRA and bisulfite pyrosequencing assays confirmed that this region is indeed significantly methylated in endometrial cancer cells relative to the level in a normal cell line (EM-E6/E7/TERT, P < 0.05, Fig. 3a and Supplementary Figure 5c). Thus, hypermethylation of miR-137 results in loss of miR-137 expression in endometrial cancer cells except in RL95-2 cells (Fig. 3b).
miR-137 is hypermethylated and reactivated by epigenetic inhibitors in endometrial cancer cells. a DNA methylation levels determined by bisulfite pyrosequencing in a normal endometrial cell line (EM-E6/E7/TERT) and 11 endometrial cancer cell lines. b, c Reactivation of pre-/pri- and mature miR-137 after epigenetic inhibitors in endometrial cancer cells after endometrial cancer cells were treated with epigenetic inhibitors. Cancer cells (HEC1A, Ishikawa H, and Hec50co) were treated with 5-aza-2’-deoxycytidine (DAC) and/or trichostatin A (TSA). Gene expression was determined by RT-qPCR and compared to untreated controls (CTR). U6 served as an internal control. Bars: means ± SD; *: P < 0.05
Next, we determined whether hypermethylation in the nearby CpG island helps regulate the expression of this locus. Hypermethylation of this promoter CpG island was detected in HEC1A and Hec50co cells (50 and 83% methylation) and, to a lesser extent, in Ishikawa H cells (14% methylation) (Fig. 3a and Supplementary Figure 5c). When these cells were treated with a demethylating agent, DAC (Decitabine), a histone deacetylase inhibitor, TSA (Trichostatin A), or both, they all displayed reactivation of both pre-/pri- and mature miR-137 (Fig. 3c, d). Re-expression was more marked when the three cell lines were treated with both agents. These results suggest that, in endometrial cancer cells, hypermethylation in the promoter of the miR-137 gene associates with loss of miR-137 expression.
miR-137 suppresses both endometrial cancer cell proliferation and colony formation in vitro
We next investigated the functional implications of miR-137 in endometrial cancer cells. We overexpressed miR-137 in HEC1A and Ishikawa H cells by transfecting them with miR-137 plasmid (miR-137) or vector (pCMV-MIR) only. mRNA expression of miR-137 (detected by RT-qPCR) confirmed that miR-137-transfected cells showed higher expression of miR-137 than those transfected with empty vector (pCMV-MIR) or parental cells (control) (Fig. 4a). This overexpression inhibited cell proliferation in the miR-137-transfected cells but not in the control transfectants (Fig. 4b). This observation was more highly significant in the HEC1A cells than in the Ishikawa H cells, which perhaps contain endogenous unmethylated and therefore expressed miR-137. Colony formation analysis indicated that miR-137 transfectants formed fewer and smaller clones (P < 0.05, Fig. 4c, d). These data suggest that miR-137 impairs cancer cell proliferation. We investigated two target genes of miR-137, enhancer zeste homolog 2 (EZH2), which participates in histone methylation, and lysine-specific demethylase 1 (LSD1), a histone demethylation enzyme [16, 19, 20]. RT-qPCR and western blotting confirmed that EZH2 and LSD1 are suppressed in cell transfected with miR-137 (Fig. 4e, f).
Functional analysis of miR-137 in endometrial cancer cells. a Relative expression of miR-137 in stably transfected clones of HEC1A and Ishikawa H cancer cells by RT-qPCR. pCMV-MIR: mock transfection with vector, miR-137: miR-137 transfection. U6 served as an internal control. Bars: means ± SD. b Cellular proliferation in transfected HEC1A and Ishikawa H cells at different time points, as determined by MTS assays. c, d Colony number and size in miR-137-transfected HEC1A and Ishikawa H endometrial cancer cells. e, f Gene expression by RT-qPCR and western blotting in transfected cells. Bars: means ± SD. *: P < 0.05; **: P < 0.01; ***: P < 0.001
miR-137 inhibits xenograft growth and tumor weight in vivo
To explore the in vivo effects of miR-137 transfectants, we implanted the HEC1A cells that overexpressed miR-137 into nude mice. Cells transfected with miR-137 grew more slowly (Fig. 5a) and formed smaller tumors (based on absolute tumor weight) than those that received vector only (Fig. 5b). Western blot analysis revealed that EZH2 and LSD1 were suppressed in the miR-137 transfectants relative to the parental HEC1A cells and the vector control (pCMV-MIR) (Fig. 5c).
miR-137 impairs xenograft tumor formation. a Xenograft tumor growth curves in miR-137 stably transfected clones of HEC1A cells. pCMV-MIR: mock transfection with vector, miR-137: miR-137 transfection. b Tumor weights of miR-137- or mock-transfected HEC1A cells. c Protein expression of EZH2 and LSD1 in transfected cells by western blotting. β-actin serves as a loading control. d, e Representative sections and summary of H&E and IHC staining of xenograft tumors. Bars: means ± SE. *: P < 0.05; **: P < 0.01
H&E staining showed that the xenograft tumors were moderately well-differentiated adenocarcinomas, consistent with the grade 2 of the original tumor [21], and we found no difference between vector and miR-137 transfectants (Fig. 5d). We found less cell proliferation (quantified by Ki-67) in the miR-137 transfectants than in the pCMV-MIR cells but no difference in apoptosis (quantified by cleaved caspase-3 staining) (Fig. 5d, e). As described above, IHC staining for EZH2 and LSD1 was also lower in the miR-137 xenograft tumors than in the pCMV-MIR tumors (P < 0.01). Taken together, our data imply that miR-137 suppresses endometrial cancer cell proliferation and tumor formation in nude mice by inhibiting the expression of EZH2 and LSD1.
miR-137 targets EZH2 and LSD1, promoting cell proliferation in miR-137-transfected cancer cells
To show that miR-137 specifically targets LSD1 and EZH2, we performed 3′-UTR reporter assays in HEC1A and Ishikawa H cells. Cells transfected with miR-137 displayed less luciferase activity than the negative miRNA control (Fig. 6a). Interestingly, reporter activities were similar in cells co-transfected with miR-137 and anti-miR-137 to those in the negative miRNA control. To confirm that miR-137 suppresses EZH2 and LSD1, we conducted a rescuing experiment by transiently transfecting EZH2 and LSD1 plasmids into miR-137 expressed stable clone of HEC1A cancer cells. RT-qPCR revealed mRNA overexpression of EZH2 and LSD1 after transfection (Fig. 6b, c), and cell proliferation assays indicated increased cell growth in both the EZH2 and LSD1 transfectants (Fig. 6d). These data support the notion that miR-137 suppresses cell proliferation in part by targeting EZH2 and LSD1.
miR-137 targets EZH2 amd LSD1. a 3′-UTR reporter assays were conducted in cells co-transfected with reporter plasmids, negative contol miRNA, miR-137, or anti-miR-137. Fold changes were normalized to plamsid and negative control miRNA transfections. Gene expression of EZH2 (b) and LSD1 (c) in miR-137-transfected HEC1A cancer cells by RT-qPCR. The stably transfected cells were subsequently transfected with EZH2 and control vector pCMV-HA (HA), or with LSD1 and its empty vector pCMV6 (PS). GAPDH served as an internal control. d Cellular proliferation in transfected cells at different time points was determined by MTS assays. *: P < 0.05; ***: P < 0.001
Discussion
Our current study shows that miR-137 is highly hypermethylated in both EEC and serous endometrial primary tumors and that this type of hypermethylation is less common in normal endometrium. Thus, repression of miR-137 might contribute to endometrial tumorigenesis. The hypermethylation of miR-137 associated with microsatellite instability (MSI) but did not correlate with patient survival rate in TCGA cohort. MSI is caused by deregulation of DNA mismatch repair genes such as MLH1, MSH2, PMS1, and PMS2, and miR-137 does not directly target those genes. Instead, EZH2 and LSD1 help regulate MLH1 expression [22, 23]. Upregulation of EZH2 suppressed MSH2 expression [24]. Those findings suggest that miR-137 methylation is indirectly associated with MSI status. Functional analysis suggested that miR-137 acts as a tumor suppressor by inhibiting cell proliferation and targeting EZH2 and LSD1 proteins to prevent endometrial cancer.
Most primary miRNAs (pri-miRNAs) are transcribed by RNA polymerase II (RNA Pol II). The farthest transcription start site for miRNA is ~9.8 kb upstream of the mature miRNA, as identified by the binding pattern of RNA Pol II [25]. In this study, we found only two CpG islands within 10 kb of mature miR-137, termed the distal and nearby CpG islands, even though we screened up to 100 kb of DNA. However, we found similar methylation levels at the distal CpG site when we compared adjacent normal controls with tumors or normal cells with cancer cell lines. In contrast, we found a significant difference in the nearby CpG site in our cohort, and we confirmed this finding in the endometrial specimens of TCGA.
DNA hypermethylation of miR-137 has been reported in several cancers [16, 19, 26,27,28], in each instance with loss of miR-137 expression in tumors. For example, hypermethylation of miR-137 in bowel lavage fluid is a prognostic marker for colorectal cancer, in oral rinses for head and neck squamous cell carcinoma, and in urine for bladder cancer [26, 29, 30]. Promoter methylation of miR-137 associates with poorer overall survival and female gender in gastric and head and neck cancer, inversely with BMI in head and neck squamous cell carcinoma, and with a worse prognosis in diffuse gastric cancer [26,27,28].
Functional analysis indicates that miR-137 suppresses cell proliferation, induces G1 cell cycle arrest and apoptosis, inhibits cell migration and invasion, and affects brain tumor stem cell differentiation [16, 19, 20, 26,27,28]. Our results agree with previous findings that overexpression of miR-137 inhibits cell proliferation and colony formation in vitro and reduces xenograft tumor growth and tumor weight in vivo. miR-137 reportedly targets both EZH2 and LSD1 proteins [19, 20]. We show here that overexpression of miR-137 suppresses protein levels, but not mRNA levels of both EZH2 and LSD1 in cells and xenograft tumors.
The enhancer of zeste homolog 2 (EZH2) is a histone methyltransferase that mediates gene silencing by catalyzing histone H3 lysine 27 trimethylation (H3K27me3). EZH2 is subject to genetic alterations in hematological malignancies, and is overexpressed in a wide range of solid tumors, including endometrial cancer [31,32,33]. It may contribute to tumorigenesis by promoting neoplastic transformation of immortalized epithelial cells, increasing cell proliferation, and inhibiting senescence and differentiation [34, 35]. Overexpression of EZH2 has been associated with high-grade and end-stage tumors, myometrial invasion, and poor overall survival in endometrial cancer [31,32,33].
LSD1, lysine-specific demethylase 1, also known as KDM1A and AOF2, belongs to the flavin monoamine oxidase (FMAO) family. This enzyme specifically demethylates histone H3 at sites where lysine 4 has undergone mono- or di-methylation (H3K4me and H3K4me2) to repress transcription at H3K9me and H3K9me2, thereby activating transcription. The final action appears to depend on LSD1’s interacting partners [36, 37]. LSD1 interacts with transcriptional repression complexes such as CoREST and NuRD to inhibit transcription or with androgen (AR) and estrogen (ER) receptors to activate transcription. Knocking out the Lsd1 locus is embryonically lethal, suggesting that Lsd1 is required for gastrulation during mouse embryogenesis [38]. Increased expression of LSD1 has been found in several types of solid tumors and associates with poor prognosis. Inhibition of LSD1 by either pharmacologic or genetic means suppresses cancer cell proliferation, migration, and invasion [39,40,41,42].
Both EZH2 and LSD1 are overexpressed [32, 33, 43] in endometrial cancer, but the mechanisms are not completely known. Genetic alternations (mutation, deletion, and amplification) of EZH2 account for 6% of TCGA’s human endometrial tumors [2]. Also, EZH2 is overexpressed in up to 25% of all human endometrial tumors, suggesting that its gene could be a potential prognostic marker [31,32,33]. Our study extends existing evidence that overexpression of EZH2 and LSD1 results partly from hypermethylation of miR-137. This notion supports our previous finding that epigenetic dysregulation of miRNA activates oncogenes [7, 12].
In summary, we identified hypermethylated promoters of miRNA genes in human endometrial primary tumors and verified that miR-137 is epigenetically silenced by DNA methylation in EEC tumors serous endometrial tumors, and endometrial cancer cell lines. This conclusion was confirmed through pharmacologic and genetic evidence, as treatment with epigenetic inhibitors restored miR-137 expression in endometrial cancer cells. Overexpression of miR-137 inhibited cancer cell proliferation and colony formation in vitro and xenograft tumor growth in vivo. Suppression of cancer cell growth associated with targeted and reduced EZH2 and LSD1 expression. Future studies could examine the therapeutic potential of miR-137 and identify additional genome-wide targets of this microRNA.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67:7–30.
The Cancer Genome Atlas Research Network, Kandoth C, Schultz N, et al. Integrated genomic characterization of endometrial carcinoma. Nature. 2013;497:67–73.
Lax SF, Kurman RJ. A dualistic model for endometrial carcinogenesis based on immunohistochemical and molecular genetic analyses. Verh Dtsch Ges Pathol. 1997;81:228–32.
Cheung LWT, Hennessy BT, Li J, et al. High frequency of PIK3R1 and PIK3R2 mutations in endometrial cancer elucidates a novel mechanism for regulation of PTEN protein stability. Cancer Discov. 2011;1:170–85.
Kuhn E, Wu RC, Guan B, et al. Identification of molecular pathway aberrations in uterine serous carcinoma by genome-wide analyses. J Natl Cancer Inst. 2012;104:1503–13.
Le Gallo M, O’Hara AJ, Rudd ML, et al. Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes. Nat Genet. 2012;44:1310–5.
Huang YW, Liu JC, Deatherage DE, et al. Epigenetic repression of microRNA-129-2 leads to overexpression of SOX4 oncogene in endometrial cancer. Cancer Res. 2009;69:9038–46.
Albitar L, Pickett G, Morgan M, et al. Models representing type I and type II human endometrial cancers: Ishikawa H and Hec50co cells. Gynecol Oncol. 2007;106:52–64.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Saito Y, Liang G, Egger G, et al. Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell. 2006;9:435–43.
Jones PA, Baylin SB. The epigenomics of cancer. Cell. 2007;128:683–92.
Huang YW, Kuo CT, Chen JH, et al. Hypermethylation of miR-203 in endometrial carcinomas. Gynecol Oncol. 2014;133:340–5.
Hsu YT, Gu F, Huang YW, et al. Promoter hypomethylation of EpCAM-regulated bone morphogenetic protein gene family in recurrent endometrial cancer. Clin Cancer Res. 2013;19:6272–85.
Gu F, Doderer MS, Huang YW, et al. CMS: a web-based system for visualization and analysis of genome-wide methylation data of human cancers. PLoS ONE. 2013;8:e60980.
Jiang J, Lee EJ, Gusev Y, et al. Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res. 2005;33:5394–403.
Balaguer F, Link A, Lozano JJ, et al. Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res. 2010;70:6609–18.
Franken NAP, Rodermond HM, Stap J, et al. Clonogenic assay of cells in vitro. Nat Protoc. 2006;1:2315–9.
Morton CL, Houghton PJ. Establishment of human tumor xenografts in immunodeficient mice. Nat Protoc. 2007;2:247–50.
Althoff K, Beckers A, Odersky A, et al. miR-137 functions as a tumor suppressor in neuroblastoma by downregulating KDM1A. Int J Cancer. 2013;133:1064–73.
Szulwach KE, Li X, Smrt RD, et al. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J Cell Biol. 2010;189:127–41.
Kuramoto H, Tamura S, Notake Y. Establishment of a cell line of human endometrial adenocarcinoma in vitro. Am J Obstet Gynecol. 1972;114:1012–9.
Wang J, Yu L, Cai J, et al. The role of EZH2 and DNA methylation in hMLH1 silencing in epithelial ovarian cancer. Biochem Biophys Res Commun. 2013;433:470–6.
Lu Y, Wajapeyee N, Turker Mitchell S, et al. Silencing of the DNA mismatch repair gene MLH1 induced by hypoxic stress in a pathway dependent on the histone demethylase LSD1. Cell Rep. 2014;8:501–13.
Yang Q, Laknaur A, Elam L, et al. Identification of polycomb group protein EZH2-mediated DNA mismatch repair gene MSH2 in human uterine fibroids. Reprod Sci. 2016;23:1314–25.
Wang G, Wang Y, Shen C, et al. RNA polymerase II binding patterns reveal genomic regions involved in microRNA gene regulation. PLoS ONE. 2010;5:e13798.
Langevin SM, Stone RA, Bunker CH, et al. MicroRNA-137 promoter methylation in oral rinses from patients with squamous cell carcinoma of the head and neck is associated with gender and body mass index. Carcinogenesis. 2010;31:864–70.
Langevin SM, Stone RA, Bunker CH, et al. MicroRNA-137 promoter methylation is associated with poorer overall survival in patients with squamous cell carcinoma of the head and neck. Cancer. 2011;117:1454–62.
Steponaitiene R, Kupcinskas J, Langner C, et al. Epigenetic silencing of miR-137 is a frequent event in gastric carcinogenesis. Mol Carcinog. 2016;55:376–86.
Harada T, Yamamoto E, Ho Yamano, et al. Analysis of DNA methylation in bowel lavage fluid for detection of colorectal cancer. Cancer Prev Res. 2014;7:1002–10.
Shimizu T, Suzuki H, Nojima M, et al. Methylation of a panel of microRNA genes is a novel biomarker for detection of bladder cancer. Eur Urol. 2013;63:1091–1100.
Bachmann IM, Halvorsen OJ, Collett K, et al. EZH2 expression is associated with high proliferation rate and aggressive tumor subgroups in cutaneous melanoma and cancers of the endometrium, prostate, and breast. J Clin Oncol. 2006;24:268–73.
Eskander RN, Ji T, Huynh B, et al. Inhibition of enhancer of zeste homolog 2 (EZH2) expression is associated with decreased tumor cell proliferation, migration, and invasion in endometrial cancer cell lines. Int J Gynecol Cancer. 2013;23:997–1005.
Zhou J, Roh J-W, Bandyopadhyay S, et al. Overexpression of enhancer of zeste homolog 2 (EZH2) and focal adhesion kinase (FAK) in high grade endometrial carcinoma. Gynecol Oncol. 2013;128:344–8.
Varambally S, Dhanasekaran SM, Zhou M, et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature. 2002;419:624–9.
Kleer CG, Cao Q, Varambally S, et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc Natl Acad Sci USA. 2003;100:11606–11.
Shi Y, Lan F, Matson C, et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–53.
Metzger E, Wissmann M, Yin N, et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature. 2005;437:436–9.
Wang J, Hevi S, Kurash JK, et al. The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat Genet. 2009;41:125–9.
Kahl P, Gullotti L, Heukamp LC, et al. Androgen receptor coactivators lysine-specific histone demethylase 1 and four and a half LIM domain protein 2 predict risk of prostate cancer recurrence. Cancer Res. 2006;66:11341–7.
Wang Y, Zhang H, Chen Y, et al. LSD1 is a subunit of the nuRD complex and targets the metastasis programs in breast cancer. Cell. 2009;138:660–72.
Ferrari-Amorotti G, Chiodoni C, Shen F, et al. Suppression of invasion and metastasis of triple-negative breast cancer lines by pharmacological or genetic inhibition of slug activity. Neoplasia. 2014;16:1047–58.
Ketscher A, Jilg CA, Willmann D, et al. LSD1 controls metastasis of androgen-independent prostate cancer cells through PXN and LPAR6. Oncogenesis. 2014;3:e120.
Liu YD, Dai M, Yang SS, et al. Overexpression of lysine-specific demethylase 1 is associated with tumor progression and unfavorable prognosis in Chinese patients with endometrioid endometrial adenocarcinoma. Int J Gynecol Cancer. 2015;25:1453–60.
Acknowledgements
This study was supported by funds from the Women’s Health Research Program and Faculty Affairs Committee at the Medical College of Wisconsin, Institutional Research Grant from the American Cancer Society, the Foundation for Women’s Cancer (to Y.-W.H.), and the National Cancer Institute (R01CA172279 to T.H.-M.H.). We thank Kristin Miller at The Ohio State University for her immunohistological services.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, W., Chen, JH., Shan, T. et al. miR-137 is a tumor suppressor in endometrial cancer and is repressed by DNA hypermethylation. Lab Invest 98, 1397–1407 (2018). https://doi.org/10.1038/s41374-018-0092-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-018-0092-x
This article is cited by
-
PDIA6, which is regulated by TRPM2-AS/miR-424-5p axis, promotes endometrial cancer progression via TGF-beta pathway
Cell Death & Disease (2023)
-
Oral squamous cell carcinomas: state of the field and emerging directions
International Journal of Oral Science (2023)
-
The long and short non-coding RNAs modulating EZH2 signaling in cancer
Journal of Hematology & Oncology (2022)
-
Prognostic Value of miR-137 in Children with Medulloblastoma and its Regulatory Effect on Tumor Progression
NeuroMolecular Medicine (2022)
-
miR-29a-3p inhibits endometrial cancer cell proliferation, migration and invasion by targeting VEGFA/CD C42/PAK1
BMC Cancer (2021)