Abstract
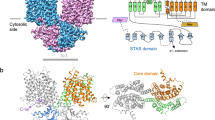
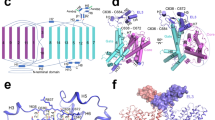
The SLC26 family of membrane proteins combines a variety of functions within a conserved molecular scaffold. Its members, besides coupled anion transporters and channels, include the motor protein Prestin, which confers electromotility to cochlear outer hair cells. To gain insight into the architecture of this protein family, we characterized the structure and function of SLC26Dg, a facilitator of proton-coupled fumarate symport, from the bacterium Deinococcus geothermalis. Its modular structure combines a transmembrane unit and a cytoplasmic STAS domain. The membrane-inserted domain consists of two intertwined inverted repeats of seven transmembrane segments each and resembles the fold of the unrelated transporter UraA. It shows an inward-facing, ligand-free conformation with a potential substrate-binding site at the interface between two helix termini at the center of the membrane. This structure defines the common framework for the diverse functional behavior of the SLC26 family.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
12 April 2016
In the version of this article initially published, the Protein Data Bank accession code for the coordinates and structure factors for SLC26DgΔSTAS (PDB 5IOF) was not included. The error has been corrected in the HTML and PDF versions of the article.
References
Alper, S.L. & Sharma, A.K. The SLC26 gene family of anion transporters and channels. Mol. Aspects Med. 34, 494–515 (2013).
Dorwart, M.R., Shcheynikov, N., Yang, D. & Muallem, S. The solute carrier 26 family of proteins in epithelial ion transport. Physiology (Bethesda) 23, 104–114 (2008).
Price, G.D. & Howitt, S.M. The cyanobacterial bicarbonate transporter BicA: its physiological role and the implications of structural similarities with human SLC26 transporters. Biochem. Cell Biol. 89, 178–188 (2011).
Ohana, E., Yang, D., Shcheynikov, N. & Muallem, S. Diverse transport modes by the solute carrier 26 family of anion transporters. J. Physiol. (Lond.) 587, 2179–2185 (2009).
Oliver, D. et al. Intracellular anions as the voltage sensor of prestin, the outer hair cell motor protein. Science 292, 2340–2343 (2001).
Dallos, P. & Fakler, B. Prestin, a new type of motor protein. Nat. Rev. Mol. Cell Biol. 3, 104–111 (2002).
Price, G.D., Woodger, F.J., Badger, M.R., Howitt, S.M. & Tucker, L. Identification of a SulP-type bicarbonate transporter in marine cyanobacteria. Proc. Natl. Acad. Sci. USA 101, 18228–18233 (2004).
Karinou, E., Compton, E.L., Morel, M. & Javelle, A. The Escherichia coli SLC26 homologue YchM (DauA) is a C4-dicarboxylic acid transporter. Mol. Microbiol. 87, 623–640 (2013).
Sharma, A.K., Rigby, A.C. & Alper, S.L. STAS domain structure and function. Cell. Physiol. Biochem. 28, 407–422 (2011).
Sharma, A.K. et al. Solution structure of the guanine nucleotide-binding STAS domain of SLC26-related SulP protein Rv1739c from Mycobacterium tuberculosis. J. Biol. Chem. 286, 8534–8544 (2011).
Babu, M. et al. Structure of a SLC26 anion transporter STAS domain in complex with acyl carrier protein: implications for E. coli YchM in fatty acid metabolism. Structure 18, 1450–1462 (2010).
Pasqualetto, E. et al. Structure of the cytosolic portion of the motor protein prestin and functional role of the STAS domain in SLC26/SulP anion transporters. J. Mol. Biol. 400, 448–462 (2010).
Geertsma, E.R. & Dutzler, R. A versatile and efficient high-throughput cloning tool for structural biology. Biochemistry 50, 3272–3278 (2011).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Geertsma, E.R., Groeneveld, M., Slotboom, D.J. & Poolman, B. Quality control of overexpressed membrane proteins. Proc. Natl. Acad. Sci. USA 105, 5722–5727 (2008).
Detro-Dassen, S. et al. Conserved dimeric subunit stoichiometry of SLC26 multifunctional anion exchangers. J. Biol. Chem. 283, 4177–4188 (2008).
Compton, E.L. et al. Conserved structure and domain organization among bacterial Slc26 transporters. Biochem. J. 463, 297–307 (2014).
Shibagaki, N. & Grossman, A.R. The role of the STAS domain in the function and biogenesis of a sulfate transporter as probed by random mutagenesis. J. Biol. Chem. 281, 22964–22973 (2006).
Chernova, M.N. et al. Acute regulation of the SLC26A3 congenital chloride diarrhoea anion exchanger (DRA) expressed in Xenopus oocytes. J. Physiol. (Lond.) 549, 3–19 (2003).
Pardon, E. et al. A general protocol for the generation of Nanobodies for structural biology. Nat. Protoc. 9, 674–693 (2014).
Compton, E.L., Karinou, E., Naismith, J.H., Gabel, F. & Javelle, A. Low resolution structure of a bacterial SLC26 transporter reveals dimeric stoichiometry and mobile intracellular domains. J. Biol. Chem. 286, 27058–27067 (2011).
Forrest, L.R., Kramer, R. & Ziegler, C. The structural basis of secondary active transport mechanisms. Biochim. Biophys. Acta 1807, 167–188 (2011).
Lu, F. et al. Structure and mechanism of the uracil transporter UraA. Nature 472, 243–246 (2011).
Wong, F.H. et al. The amino acid-polyamine-organocation superfamily. J. Mol. Microbiol. Biotechnol. 22, 105–113 (2012).
Västermark, A. & Saier, M.H. Jr. Evolutionary relationship between 5+5 and 7+7 inverted repeat folds within the amino acid-polyamine-organocation superfamily. Proteins 82, 336–346 (2014).
Gorbunov, D. et al. Molecular architecture and the structural basis for anion interaction in prestin and SLC26 transporters. Nat. Commun. 5, 3622 (2014).
Barneaud-Rocca, D., Etchebest, C. & Guizouarn, H. Structural model of the anion exchanger 1 (SLC4A1) and identification of transmembrane segments forming the transport site. J. Biol. Chem. 288, 26372–26384 (2013).
Cordat, E. & Reithmeier, R.A. Structure, function, and trafficking of SLC4 and SLC26 anion transporters. Curr. Top. Membr. 73, 1–67 (2014).
He, J.J. & Quiocho, F.A. Dominant role of local dipoles in stabilizing uncompensated charges on a sulfate sequestered in a periplasmic active transport protein. Protein Sci. 2, 1643–1647 (1993).
Dutzler, R., Campbell, E.B. & MacKinnon, R. Gating the selectivity filter in ClC chloride channels. Science 300, 108–112 (2003).
Lobet, S. & Dutzler, R. Ion-binding properties of the ClC chloride selectivity filter. EMBO J. 25, 24–33 (2006).
Kalli, A.C., Sansom, M.S. & Reithmeier, R.A. Molecular dynamics simulations of the bacterial UraA H+-uracil symporter in lipid bilayers reveal a closed state and a selective interaction with cardiolipin. PLOS Comput. Biol. 11, e1004123 (2015).
Lee, C. et al. A two-domain elevator mechanism for sodium/proton antiport. Nature 501, 573–577 (2013).
Paulino, C., Wohlert, D., Kapotova, E., Yildiz, O. & Kuhlbrandt, W. Structure and transport mechanism of the sodium/proton antiporter MjNhaP1. eLife 3, e03583 (2014).
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005).
Casadaban, M.J. & Cohen, S.N. Analysis of gene control signals by DNA fusion and cloning in Escherichia coli. J. Mol. Biol. 138, 179–207 (1980).
Ehrnstorfer, I.A., Geertsma, E.R., Pardon, E., Steyaert, J. & Dutzler, R. Crystal structure of a SLC11 (NRAMP) transporter reveals the basis for transition-metal ion transport. Nat. Struct. Mol. Biol. 21, 990–996 (2014).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
Pape, T. & Schneider, T.R. HKL2MAP: a graphical user interface for phasing with SHELX programs. J. Appl. Crystallogr. 37, 843–844 (2004).
de La Fortelle, E. & Bricogne, G. in Methods in Enzymology (eds. Carter, C.W. & Sweet, R.M.) 492–494 (Academic, New York, 1997).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Geertsma, E.R., Nik Mahmood, N.A., Schuurman-Wolters, G.K. & Poolman, B. Membrane reconstitution of ABC transporters and assays of translocator function. Nat. Protoc. 3, 256–266 (2008).
Acknowledgements
This research was supported by a grant of the Swiss National Science Foundation through the National Center of Competence in Research (NCCR) Structural Biology (R.D.), a long-term fellowship from the Human Frontier Science Program (LT-00899/2008) (E.R.G.), the German Research Foundation through the Cluster of Excellence Frankfurt 'Macromolecular Complexes' (E.R.G.), INSTRUCT as part of the European Strategy Forum on Research Infrastructures (ESFRI) (J.S.) and the Hercules Foundation Flanders (J.S.). We thank B. Fakler and R.J.C. Hilf for input at initial stages of the project; the staff of the X06SA beamline for support during data collection; B. Blattman and C. Stutz-Ducommun (Protein Crystallization Center at University of Zurich) for their support with crystallization; B. Dreier for help with MALS experiments; and N. Buys and K. Willibal for help with nanobody selection. All members of the Dutzler laboratory are acknowledged for help in all stages of the project.
Author information
Authors and Affiliations
Contributions
F.R.S. and E.R.G. screened homologs, established purification of SLC26Dg and crystallized SLC26DgΔSTAS. E.P. performed immunization, cloned and expressed nanobodies, and guided E.R.G. during the initial selections. J.S. supervised nanobody production. E.R.G. and Y.N. established nanobody overexpression and purification for crystallization. Y.-N.C. and E.R.G. crystallized the SLC26Dg–nanobody complex and carried out crystallographic experiments and transport assays. R.D. assisted E.R.G. in structure determination. E.R.G. and R.D. jointly planned the experiments, analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Sequence alignment.
Sequence alignment of SLC26Dg (GI:499845065), the E. coli transporter DauA (GI:115512548) and human Prestin (SLC26A5, GI:30348882). The Alignment was improved based on the SLC26Dg structure and the structures of the STAS domains of Prestin and DauA. Secondary structure elements of SLC26Dg are shown below. The two halves of the transmembrane domain are colored in green and beige respectively. The STAS domain is colored in red. The numbering corresponds to SLC26Dg. Selected residues in the aqueous cavity in proximity to the potential substrate binding site are indicated as blue or beige dots, the start of the STAS domain as red dot.
Supplementary Figure 2 Functional properties.
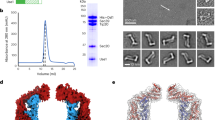
Gel filtration and light scattering results for different protein constructs purified in the detergent DM. Continuous black traces correspond to the UV280 elution profiles. The respective molecular weight of the protein component obtained from light scattering is shown at its corresponding position on the chromatogram in red. Panels show (a), SLC26Dg, and (b), the SLC26Dg-Nb5776 complex. (c) Glutaraldehyde (GA) crosslinking of reconstituted SLC26Dg. Samples were separated by SDS PAGE, the respective molecular weight of marker proteins is indicated. The band at around 130 kDa corresponds to a cross-linked SLC26Dg dimer, the band at around 55 kDa to monomeric SLC26Dg observed upon solubilization of proteoliposomes in SDS prior to glutaraldehyde addition. Left panel, SDS-PAGE gel of proteoliposomes reconstituted at a protein:lipid ratio of 1:50 (wt/wt); right panel, Western blot of proteoliposomes reconstituted at 1:250 and 1:500 indicating that crosslinking of SLC26Dg dimers also occurred at very low protein concentrations in the membrane. (d) Substrate selectivity. Competition of 14C-fumarate uptake by a 70-fold excess of other unlabeled dicarboxylates. (e) Concentration dependence of fumarate transport. The solid curve shows a fit to a Michaelis-Menten equation with a linear component due to the passive uptake of fumarate into liposomes. The dashed line shows the slope of the linear component. Specific activities were calculated assuming a complete incorporation of the protein into liposomes during reconstitution. (f) Membrane orientation of SLC26Dg (WT) and SLC26DgΔSTAS (TM) in proteoliposomes. Both proteins were expressed and reconstituted as N-terminal fusions to GFP. The membrane orientation was investigated by probing the accessibility of the HRV 3C protease cleavage site between the membrane transporter and GFP. In-gel fluorescence was measured after separation of the cleaved products on SDS PAGE. – refers to the untreated samples, p to samples treated with HRV 3C protease, and p/d to samples treated with HRV 3C protease and detergent. * indicates uncleaved protein. For both constructs, incubation with the protease decreased the GFP fluorescence of the uncleaved protein by approximately 50% indicating a similar random distribution of both orientations within the membrane. Molecular weight of markers (kDa) are indicated on the left. (g) Transport properties of SLC26DgΔSTAS. Time dependent uptake of 14C-fumarate into proteoliposomes containing SLC26DgΔSTAS (TM). ΔpH refers to an outside pH of 6.0 and a pH of 7.5 inside the liposomes, ΔNa+ to an external Na+ concentration of 50 mM and no Na+ inside the liposomes. Traces of WT and liposomes not containing any protein (with data presented in Fig. 1a) are shown for comparison. Proteoliposomes were prepared with equimolar amounts of protein. Equimolar protein amounts were also used for each data point. Data in d and g represents mean and s.e.m of 3 technical replicates, data in e represent mean and s.e.m of 6 technical replicates from two independent batches of proteoliposomes.
Supplementary Figure 3 Electron density of the SLC26Dg–Nb5776 complex.
Proteins are shown as sticks with carbons colored in green. 2Fo-Fc density (calculated at 3.2 Å, sharpened with b=90 and contoured at 1σ) calculated with phases obtained from the refined model is superimposed on the structure. Sections of different regions of the complex are shown: (a) Stereo view of the substrate binding site of the transmembrane domain with a bound DM molecule (with carbon atoms of the detergent colored in blue). The detergent binds to a similar location as a bound detergent in the UraA structure. (b) STAS domain viewed from the nanobody binding interface with α-helices 2 and 3 labeled. (c) View of the nanobody. The electron density for the nanobody is less well defined than the density of the transmembrane domain of SLC26Dg, which indicates a higher degree of disorder. The entire nanobody is shown. Supplementary Figs. 3, 4, 5, 6 were prepared with DINO (http://www.dino3d.org).
Supplementary Figure 4 STAS-domain structure.
(a) Ribbon representation of the STAS domain of SLC26Dg with secondary structure elements labeled. (b) Superposition of Cα-traces of the SLC26Dg structure (red) with the equivalent domain from the SLC26 transporter from Rhodobacter sphaeroides (SLC26Rs, blue, PDB code 3oiz), which shares 50% of identical residues. (c) Superposition of the SLC26Dg STAS domain on a structure of the equivalent domain of rat Prestin that lacks a long unstructured loop connecting α1 and β312 (SLC26A5, blue, PDB code 3llo). (d) Superposition of the SLC26Dg STAS domain on the STAS domain ACP complex of the E. coli transporter DauA11 (PDB code 3ny7). The DauA STAS domain is colored in blue, ACP in green. ACP binds to a region on the C-terminus of the STAS domain of DauA that is not defined in the electron density of SLC26Dg. The binding interface does not overlap with the nanobody-binding epitope on the STAS domain of SLC26Dg.
Supplementary Figure 5 SLC26Dg-Nb5776 interactions.
(a) Ribbon representation of the nanobody Nb5776 (blue) and the STAS domain of SLC26Dg (red). Both proteins interact via a parallel β-sheet formed by β5 of Nb5776 (containing the variable region CDR2) and β4 of the STAS domain and via side-chain interactions with a long loop of the nanobody connecting β-strands 8 and 9 (part of the variable region CDR3, *) that has changed its conformation upon binding to the transport protein. (b) Side-chain interactions between the Nb5776 and the STAS domain of SLC26Dg. (c) Ribbon representation of the Nb5776 structure. (d) Superposition of Cα-traces of the isolated Nb5776 (beige) and of its structure observed in the complex (blue). The difference in the conformation of the CDR3 region is apparent. (e) Stereo view of 2Fo-Fc density (calculated at 2.4 Å and contoured at 1σ) of a Nb5776 crystal superimposed on the structure. An asterisk indicates the region of CDR3 that has changed its conformation upon binding to SLC26Dg. (f) Interactions of the nanobody and the STAS domain between symmetry-related molecules in the crystals of the SLC26Dg-Nb5776 complex. Transmembrane and STAS domains of SLC26Dg are colored in green and red respectively, the nanobody is colored in blue. (g) SLC26Dg structure placed in a model of a lipid bilayer (obtained from http://www.lobos.nih.gov/mbs/coords.shtml). The molecular surface of the protein is shown with polar residues colored in green, acidic residues in red and basic residues in blue. The observed conformation of SLC26Dg would place the hydrophilic STAS domain (indicated by brackets) within the hydrophobic core of the lipid bilayer.
Supplementary Figure 6 TM-domain structure and functional behavior of a mutant.
(a) Stereo view of the SLC26Dg transmembrane domain. The protein is shown as ribbon with its N-terminal half colored in green and its C-terminal half in beige. (b) Stereo view of a superposition of the pseudo-symmetry related N- and C-terminal halves of the SLC26Dg transmembrane domain. The protein sub-domains are shown as Cα representation, the sidechains of the two pseudo-symmetry related glutamates facing the intracellular aqueous cavity as sticks. (c) Model of the SLC26Dg transmembrane domain in a lipid bilayer. The molecular surface of the protein is displayed with aromatic residues colored in beige and hydrophobic residues in yellow. Lipids are shown as sticks. (d) Arrangement of molecules in the unit cell of the SLC26DgΔSTAS crystal. The crystal is of space group P1 and contains 4 copies of the transport domain. (e) Stereo view of 2Fo-Fc electron density (calculated at 4.2 Å, sharpened with b=90 and contoured at 1σ) superimposed on a Cα-trace of the SLC26DgΔSTAS structure. The model was improved by refinement maintaining strong structural constraints. (f) Transport properties of the SLC26Dg double mutant E38A E241A. Time dependent uptake of 14C-fumarate (136 μM) into proteoliposomes. ΔpH refers to a pH of 6.0 outside and 7.5 inside the liposomes, pH 7.5 refers to symmetrical pH conditions. (g) Concentration dependence of fumarate transport in the mutant E38A E241A. Left panel: quantification of the protein reconstituted in the proteoliposomes. Proteoliposomes were solubilized with 1% DDM and samples before and after ultracentrifugation were taken to determine the fraction membrane-inserted proteins. Densitometric analysis of the samples after ultracentrifugation indicates a 7 times lower protein concentration in proteoliposomes containing the mutant E38A E241A. Right panel: the solid curve shows a fit to a Michaelis-Menten equation with a linear component due to the passive uptake of fumarate into liposomes. The dashed line shows the slope of the linear component. Specific activities were calculated in comparison to WT based on a quantification of the reconstituted protein in proteoliposomes (left panel). Data in f and g represent mean and s.e.m of 3 technical replicates.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 1852 kb)
Rights and permissions
About this article
Cite this article
Geertsma, E., Chang, YN., Shaik, F. et al. Structure of a prokaryotic fumarate transporter reveals the architecture of the SLC26 family. Nat Struct Mol Biol 22, 803–808 (2015). https://doi.org/10.1038/nsmb.3091
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3091
This article is cited by
-
Mechanism of anion exchange and small-molecule inhibition of pendrin
Nature Communications (2024)
-
The Remarkable Outer Hair Cell: Proceedings of a Symposium in Honour of W. E. Brownell
Journal of the Association for Research in Otolaryngology (2023)
-
Cryo-EM structures of thermostabilized prestin provide mechanistic insights underlying outer hair cell electromotility
Nature Communications (2022)
-
Characterization of pendrin in urinary extracellular vesicles in a rat model of aldosterone excess and in human primary aldosteronism
Hypertension Research (2021)
-
Progress in Structural Biology of Solute Carriers
Current Molecular Biology Reports (2021)