Abstract
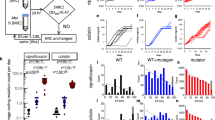
Predicting the future is difficult, especially for evolutionary processes that are influenced by numerous unknown factors. Still, this is what is required of drug developers when they assess the risk of resistance arising against a new antibiotic candidate during preclinical development. In this Opinion article, we argue that the traditional procedures that are used for the prediction of antibiotic resistance today could be markedly improved by including a broader analysis of bacterial fitness, infection dynamics, horizontal gene transfer and other factors. This will lead to more informed preclinical decisions for continuing or discontinuing the development of drug candidates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
[No authors listed.] Tackling drug-resistant infections globally: final report and recommendations (review on antimicrobial resistance, 2016). AMR Review https://amr-review.org/sites/default/files/160525_Final%20paper_with%20cover.pdf (2016).
Gilmore, M. S., Lebreton, F. & van Schaik, W. Genomic transition of enterococci from gut commensals to leading causes of multidrug-resistant hospital infection in the antibiotic era. Curr. Opin. Microbiol. 16, 10–16 (2013).
zur Wiesch, P. A., Kouyos, R., Engelstädter, J., Regoes, R. R. & Bonhoeffer, S. Population biological principles of drug-resistance evolution in infectious diseases. Lancet Infect. Dis. 11, 236–247 (2011).
Smith, M. R. & Wood, W. B. An experimental analysis of the curative action of penicillin in acute bacterial infections. III. The effect of suppuration upon the antibacterial action of the drug. J. Exp. Med. 103, 509–522 (1956).
Palaci, M. et al. Cavitary disease and quantitative sputum bacillary load in cases of pulmonary tuberculosis. J. Clin. Microbiol. 45, 4064–4066 (2007).
Feldman, W. E. Concentrations of bacteria in cerebrospinal fluid of patients with bacterial meningitis. J. Pediatr. 88, 549–552 (1976).
Canetti, G. Present aspects of bacterial resistance in tuberculosis. Am. Rev. Respir. Dis. 92, 687–703 (1965).
Canetti, G. Dynamic aspects of the pathology and bacteriology of tuberculous lesions. Am. Rev. Tuberc. 74, 13–21 (1956).
Foster, P. L. Methods for determining spontaneous mutation rates. Methods Enzymol. 409, 195–213 (2006).
Drake, J. W., Charlesworth, B., Charlesworth, D. & Crow, J. F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998).
Lynch, M. Evolution of the mutation rate. Trends Genet. 26, 345–352 (2010).
Tubulekas, I., Buckingham, R. H. & Hughes, D. Mutant ribosomes can generate dominant kirromycin resistance. J. Bacteriol. 173, 3635–3643 (1991).
Lofton, H., Pränting, M., Thulin, E. & Andersson, D. I. Mechanisms and fitness costs of resistance to antimicrobial peptides LL-37, CNY100HL and wheat germ histones. PLoS ONE 8, e68875 (2013).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
Nilsson, A. I., Berg, O. G., Aspevall, O., Kahlmeter, G. & Andersson, D. I. Biological costs and mechanisms of fosfomycin resistance in Escherichia coli. Antimicrob. Agents Chemother. 47, 2850–2858 (2003).
Thulin, E., Sundqvist, M. & Andersson, D. I. Amdinocillin (mecillinam) resistance mutations in clinical isolates and laboratory-selected mutants of Escherichia coli. Antimicrob. Agents Chemother. 59, 1718–1727 (2015).
Drusano, G. L., Louie, A., MacGowan, A. & Hope, W. Suppression of emergence of resistance in pathogenic bacteria: keeping our powder dry, part 1. Antimicrob. Agents Chemother. 60, 1183–1193 (2015).
Drusano, G. L., Hope, W., MacGowan, A. & Louie, A. Suppression of emergence of resistance in pathogenic bacteria: keeping our powder dry, part 2. Antimicrob. Agents Chemother. 60, 1194–1201 (2015).
Chancey, S. T., Zähner, D. & Stephens, D. S. Acquired inducible antimicrobial resistance in Gram-positive bacteria. Future Microbiol. 7, 959–978 (2012).
Chevereau, G. et al. Quantifying the determinants of evolutionary dynamics leading to drug resistance. PLoS Biol. 13, e1002299 (2015).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Andersson, D. I. & Levin, B. R. The biological cost of antibiotic resistance. Curr. Opin. Microbiol. 2, 489–493 (1999).
Wiser, M. J., Ribeck, N. & Lenski, R. E. Long-term dynamics of adaptation in asexual populations. Science 342, 1364–1367 (2013).
Hughes, D. & Andersson, D. I. Evolutionary consequences of drug resistance: shared principles across diverse targets and organisms. Nat. Rev. Genet. 16, 459–471 (2015).
Brandis, G., Pietsch, F., Alemayehu, R. & Hughes, D. Comprehensive phenotypic characterization of rifampicin resistance mutations in Salmonella provides insight into the evolution of resistance in Mycobacterium tuberculosis. J. Antimicrob. Chemother. 70, 680–685 (2015).
O'Neill, A. J., Huovinen, T., Fishwick, C. W. G. & Chopra, I. Molecular genetic and structural modeling studies of Staphylococcus aureus RNA polymerase and the fitness of rifampin resistance genotypes in relation to clinical prevalence. Antimicrob. Agents Chemother. 50, 298–309 (2006).
Bottger, E. C., Springer, B., Pletschette, M. & Sander, P. Fitness of antibiotic-resistant microorganisms and compensatory mutations. Nat. Med. 4, 1343–1344 (1998).
Sander, P. et al. Fitness cost of chromosomal drug resistance-conferring mutations. Antimicrob. Agents Chemother. 46, 1204–1211 (2002).
Shcherbakov, D. et al. Directed mutagenesis of Mycobacterium smegmatis 16S rRNA to reconstruct the in vivo evolution of aminoglycoside resistance in Mycobacterium tuberculosis. Mol. Microbiol. 77, 830–840 (2010).
Foucault, M.-L., Depardieu, F., Courvalin, P. & Grillot-Courvalin, C. Inducible expression eliminates the fitness cost of vancomycin resistance in enterococci. Proc. Natl Acad. Sci. USA 107, 16964–16969 (2010).
Andersson, D. I. & Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 12, 465–478 (2014).
Gullberg, E., Albrecht, L. M., Karlsson, C., Sandegren, L. & Andersson, D. I. Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. mBio 5, e01918-14 (2014).
Oz, T. et al. Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution. Mol. Biol. Evol. 31, 2387–2401 (2014).
De Visser, J. A. G. M. & Krug, J. Empirical fitness landscapes and the predictability of evolution. Nat. Rev. Genet. 15, 480–490 (2014).
Kondrashov, D. A. & Kondrashov, F. A. Topological features of rugged fitness landscapes in sequence space. Trends Genet. 31, 24–33 (2015).
Brandis, G. & Hughes, D. Genetic characterization of compensatory evolution in strains carrying rpoB Ser531Leu, the rifampicin resistance mutation most frequently found in clinical isolates. J. Antimicrob. Chemother. 68, 2493–2497 (2013).
Brandis, G., Wrande, M., Liljas, L. & Hughes, D. Fitness-compensatory mutations in rifampicin-resistant RNA polymerase. Mol. Microbiol. 85, 142–151 (2012).
Lannergård, J. et al. Genetic complexity of fusidic acid-resistant small colony variants (SCV) in Staphylococcus aureus. PLoS ONE 6, e28366 (2011).
Marcusson, L. L., Frimodt-Møller, N. & Hughes, D. Interplay in the selection of fluoroquinolone resistance and bacterial fitness. PLoS Pathog. 5, e1000541 (2009).
Schrag, S. J., Perrot, V. & Levin, B. R. Adaptation to the fitness costs of antibiotic resistance in Escherichia coli. Proc. Biol. Sci. 264, 1287–1291 (1997).
Angst, D. C. & Hall, A. R. The cost of antibiotic resistance depends on evolutionary history in Escherichia coli. BMC Evol. Biol. 13, 163 (2013).
Komp Lindgren, P., Marcusson, L. L., Sandvang, D., Frimodt-Møller, N. & Hughes, D. Biological cost of single and multiple norfloxacin resistance mutations in Escherichia coli implicated in urinary tract infections. Antimicrob. Agents Chemother. 49, 2343–2351 (2005).
Trindade, S. et al. Positive epistasis drives the acquisition of multidrug resistance. PLoS Genet. 5, e1000578 (2009).
Björkman, J., Samuelsson, P., Andersson, D. I. & Hughes, D. Novel ribosomal mutations affecting translational accuracy, antibiotic resistance and virulence of Salmonella typhimurium. Mol. Microbiol. 31, 53–58 (1999).
Hall, A. R. & MacLean, R. C. Epistasis buffers the fitness effects of rifampicin-resistance mutations in Pseudomonas aeruginosa. Evolution 70, 1161–1161 (2016).
Rozen, D. E., McGee, L., Levin, B. R. & Klugman, K. P. Fitness costs of fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob. Agents Chemother. 51, 412–416 (2007).
Vogwill, T., Kojadinovic, M. & MacLean, R. C. Epistasis between antibiotic resistance mutations and genetic background shape the fitness effect of resistance across species of Pseudomonas. Proc. Biol. Sci. 283, 20160151 (2016).
Vogwill, T. & MacLean, R. C. The genetic basis of the fitness costs of antimicrobial resistance: a meta-analysis approach. Evol. Appl. 8, 284–295 (2015).
Johanson, U., Ævarsson, A., Liljas, A. & Hughes, D. The dynamic structure of EF-G studied by fusidic acid resistance and internal revertants. J. Mol. Biol. 258, 420–432 (1996).
Nagaev, I., Bjorkman, J., Andersson, D. I. & Hughes, D. Biological cost and compensatory evolution in fusidic acid-resistant Staphylococcus aureus. Mol. Microbiol. 40, 433–439 (2001).
Salverda, M. L. M. et al. Initial mutations direct alternative pathways of protein evolution. PLoS Genet. 7, e1001321 (2011).
Weinreich, D. M., Delaney, N. F., Depristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
Dahlberg, C. & Chao, L. Amelioration of the cost of conjugative plasmid carriage in Eschericha coli K12. Genetics 165, 1641–1649 (2003).
Loftie-Eaton, W. et al. Evolutionary paths that expand plasmid host-range: implications for spread of antibiotic resistance. Mol. Biol. Evol. 33, 885–897 (2016).
San Millan, A., Heilbron, K. & MacLean, R. C. Positive epistasis between co-infecting plasmids promotes plasmid survival in bacterial populations. ISME J. 8, 601–612 (2014).
San Millan, A. et al. Positive selection and compensatory adaptation interact to stabilize non-transmissible plasmids. Nat. Commun. 5, 5208–5211 (2014).
Silva, R. F. et al. Pervasive sign epistasis between conjugative plasmids and drug-resistance chromosomal mutations. PLoS Genet. 7, e1002181 (2011).
Porse, A., Schønning, K., Munck, C. & Sommer, M. O. A. Survival and evolution of a large multidrug resistance plasmid in new clinical bacterial hosts. Mol. Biol. Evol. 33, 2860–2873 (2016).
Alekshun, M. N. & Levy, S. B. Molecular mechanisms of antibacterial multidrug resistance. Cell 128, 1037–1050 (2007).
Palmer, A. C. & Kishony, R. Understanding, predicting and manipulating the genotypic evolution of antibiotic resistance. Nat. Rev. Genet. 14, 243–248 (2013).
Garcia, L. G. et al. Antibiotic activity against small-colony variants of Staphylococcus aureus: review of in vitro, animal and clinical data. J. Antimicrob. Chemother. 68, 1455–1464 (2013).
Munck, C., Gumpert, H. K., Wallin, A. I. N., Wang, H. H. & Sommer, M. O. A. Prediction of resistance development against drug combinations by collateral responses to component drugs. Sci. Transl Med. 6, 262ra156 (2014).
Imamovic, L. & Sommer, M. O. A. Use of collateral sensitivity networks to design drug cycling protocols that avoid resistance development. Sci. Transl Med. 5, 204ra132 (2013).
Kim, S., Lieberman, T. D. & Kishony, R. Alternating antibiotic treatments constrain evolutionary paths to multidrug resistance. Proc. Natl Acad. Sci. USA 111, 14494–14499 (2014).
Lázár, V. et al. Genome-wide analysis captures the determinants of the antibiotic cross-resistance interaction network. Nat. Commun. 5, 4352 (2014).
Pena-Miller, R. et al. When the most potent combination of antibiotics selects for the greatest bacterial load: the smile–frown transition. PLoS Biol. 11, e1001540 (2013).
Périchon, B. & Courvalin, P. Synergism between β-lactams and glycopeptides against VanA-type methicillin-resistant Staphylococcus aureus and heterologous expression of the vanA operon. Antimicrob. Agents Chemother. 50, 3622–3630 (2006).
Brolund, A. & Sandegren, L. Characterization of ESBL disseminating plasmids. Infect. Dis. (Lond.) 48, 18–25 (2016).
Mathers, A. J., Peirano, G. & Pitout, J. D. D. The role of epidemic resistance plasmids and international high-risk clones in the spread of multidrug-resistant Enterobacteriaceae. Clin. Microbiol. Rev. 28, 565–591 (2015).
Molton, J. S., Tambyah, P. A., Ang, B. S. P., Ling, M. L. & Fisher, D. A. The global spread of healthcare-associated multidrug-resistant bacteria: a perspective from Asia. Clin. Infect. Dis. 56, 1310–1318 (2013).
Bean, D. C., Livermore, D. M., Papa, I. & Hall, L. M. C. Resistance among Escherichia coli to sulphonamides and other antimicrobials now little used in man. J. Antimicrob. Chemother. 56, 962–964 (2005).
Enne, V. I., Livermore, D. M., Stephens, P. & Hall, L. M. Persistence of sulphonamide resistance in Escherichia coli in the UK despite national prescribing restriction. Lancet 357, 1325–1328 (2001).
Sundqvist, M. et al. Little evidence for reversibility of trimethoprim resistance after a drastic reduction in trimethoprim use. J. Antimicrob. Chemother. 65, 350–360 (2010).
Locke, J. B., Hilgers, M. & Shaw, K. J. Novel ribosomal mutations in Staphylococcus aureus strains identified through selection with the oxazolidinones linezolid and torezolid (TR-700). Antimicrob. Agents Chemother. 53, 5265–5274 (2009).
Gordon, D. M. & Riley, M. A. A theoretical and experimental analysis of bacterial growth in the bladder. Mol. Microbiol. 6, 555–562 (1992).
Sandegren, L., Lindqvist, A., Kahlmeter, G. & Andersson, D. I. Nitrofurantoin resistance mechanism and fitness cost in Escherichia coli. J. Antimicrob. Chemother. 62, 495–503 (2008).
Thomas, C. M. & Nielsen, K. M. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat. Rev. Microbiol. 3, 711–721 (2005).
Naseer, U. & Sundsfjord, A. The CTX-M conundrum: dissemination of plasmids and Escherichia coli clones. Microb. Drug Resist. 17, 83–97 (2011).
Allen, H. K., Moe, L. A., Rodbumrer, J., Gaarder, A. & Handelsman, J. Functional metagenomics reveals diverse β-lactamases in a remote Alaskan soil. ISME J. 3, 243–251 (2008).
D'Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461 (2011).
Sommer, M. O. A., Dantas, G. & Church, G. M. Functional characterization of the antibiotic resistance reservoir in the human microflora. Science 325, 1128–1131 (2009).
Munck, C. et al. Limited dissemination of the wastewater treatment plant core resistome. Nat. Commun. 6, 8452 (2015).
Forsberg, K. J. et al. Bacterial phylogeny structures soil resistomes across habitats. Nature 509, 612–616 (2014).
Kahlmeter, G. & Poulsen, H. O. Antimicrobial susceptibility of Escherichia coli from community-acquired urinary tract infections in Europe: the ECO·SENS study revisited. Int. J. Antimicrob. Agents 39, 45–51 (2012).
Huseby, D. L. et al. Mutation supply and relative fitness shape the genotypes of ciprofloxacin-resistant Escherichia coli. Mol. Biol. Evol. 34, 1029–1039 (2017).
Moore, A. M., Munck, C., Sommer, M. O. A. & Dantas, G. Functional metagenomic investigations of the human intestinal microbiota. Front. Microbiol. 2, 188 (2011).
Dantas, G. & Sommer, M. O. Context matters — the complex interplay between resistome genotypes and resistance phenotypes. Curr. Opin. Microbiol. 15, 577–582 (2012).
Yoon, E.-J. et al. Origin in Acinetobacter gyllenbergii and dissemination of aminoglycoside-modifying enzyme AAC(6′)-Ih. J. Antimicrob. Chemother. 71, 601–606 (2016).
Martínez, J. L., Coque, T. M. & Baquero, F. Prioritizing risks of antibiotic resistance genes in all metagenomes. Nat. Rev. Microbiol. 13, 396–396 (2015).
Jaffé, A., Chabbert, Y. A. & Derlot, E. Selection and characterization of β-lactam-resistant Escherichia coli K-12 mutants. Antimicrob. Agents Chemother. 23, 622–625 (1983).
George, A. M. & Levy, S. B. Amplifiable resistance to tetracycline, chloramphenicol, and other antibiotics in Escherichia coli: involvement of a non-plasmid-determined efflux of tetracycline. J. Bacteriol. 155, 531–540 (1983).
Heisig, P. & Tschorny, R. Characterization of fluoroquinolone-resistant mutants of escherichia coli selected in vitro. Antimicrob. Agents Chemother. 38, 1284–1291 (1994).
Buckel, P., Buchberger, A., Böck, A. & Wittmann, H. G. Alteration of ribosomal protein L6 in mutants of Escherichia coli resistant to gentamicin. Mol. Gen. Genet. 158, 47–54 (1977).
Adler, M., Anjum, M., Andersson, D. I. & Sandegren, L. Influence of acquired β-lactamases on the evolution of spontaneous carbapenem resistance in Escherichia coli. J. Antimicrob. Chemother. 68, 51–59 (2013).
Oakberg, E. F. & Luria, S. W. Mutations to sulfonamide resistance in Staphylococcus aureus. Genetics 32, 249–261 (1947).
Zurenko, G. E. et al. In vitro activities of U-100592 and U-100766, novel oxazolidinone antibacterial agents. Antimicrob. Agents Chemother. 40, 839–845 (1996).
Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug. Discov. 12, 371–387 (2013).
Acknowledgements
Work in the authors' laboratories was supported by grants from the Swedish Research Council (to D.I.A.), and the Novo Nordisk Foundation, the Lundbeck Foundation and the Danish Free Research Council (to M.O.A.S. and C.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: M.O.A.S. and R.T.K. are shareholders in AntibioTx; C.M. declares no competing interests; D.I.A. is a consultant for Prebona and Bactiguard.
Rights and permissions
About this article
Cite this article
Sommer, M., Munck, C., Toft-Kehler, R. et al. Prediction of antibiotic resistance: time for a new preclinical paradigm?. Nat Rev Microbiol 15, 689–696 (2017). https://doi.org/10.1038/nrmicro.2017.75
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2017.75
This article is cited by
-
Pangenome analysis of Corynebacterium striatum: insights into a neglected multidrug-resistant pathogen
BMC Microbiology (2023)
-
Intermittent antibiotic treatment of bacterial biofilms favors the rapid evolution of resistance
Communications Biology (2023)
-
CanB is a metabolic mediator of antibiotic resistance in Neisseria gonorrhoeae
Nature Microbiology (2023)
-
A novel pyrazolone complex P-FAH-Cu-bpy induces death of Escherichia coli and Staphylococcus aureus by disrupting cell structure and blocking energy
Archives of Microbiology (2023)
-
Population size mediates the contribution of high-rate and large-benefit mutations to parallel evolution
Nature Ecology & Evolution (2022)