Key Points
-
Systems-biology approaches, such as genetic perturbations combined with kinetic luminescence imaging, synthetic-biology approaches and mathematical modelling, are being used to address the complexity of clock function.
-
The circadian clock community has used mouse genetics to establish non-redundant roles for many clock factors that contribute to circadian oscillators that function within cells and in circadian behaviour.
-
Proportionality and paralogue compensation are key principles that convey robustness to the circadian clock.
-
Researchers have exploited the cyanobacterial system to establish that there are oscillations in phosphorylation status in experiments using three proteins (KaiA, KaiB and KaiC) and ATP. The cyanobacterial clock community has leveraged this model to understand the molecular events that establish periodicity as well as concepts such as molecular synchronization. Quantitative and conceptual models have been important in driving this research.
-
'Traditional' techniques such as genetics can be used to tackle complex issues in clock biology, such as identifying the components and establishing the mechanism for temperature compensation.
Abstract
After several decades dominated by reductionist approaches in biology, researchers are returning to the study of complex biology with a litany of new and old techniques — this paradigm has been termed systems biology. Here we detail how systems biology is being used to uncover complex systems-level properties of the circadian clock. These properties include robustness, periodicity and temperature compensation. We describe how clock researchers are using systems-biology techniques, such as genetic perturbations, kinetic luminescence imaging, synthetic biology and mathematical modelling, to untangle these complex properties in mammals, fungi and bacteria. The strategies developed in the context of circadian clocks may prove useful for tackling similar problems in other systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Green, C. B., Takahashi, J. S. & Bass, J. The meter of metabolism. Cell 134, 728–742 (2008).
Kovac, J., Husse, J. & Oster, H. A time to fast, a time to feast: the crosstalk between metabolism and the circadian clock. Mol. Cell 28, 75–80 (2009).
Froy, O. Metabolism and circadian rhythms — implications for obesity. Endocr. Rev. 31, 1–24 (2010).
Eckel-Mahan, K. L. & Storm, D. R. Circadian rhythms and memory: not so simple as cogs and gears. EMBO Rep. 10, 584–591 (2009).
Kyriacou, C. P. & Hastings, M. H. Circadian clocks: genes, sleep, and cognition. Trends Cogn. Sci. 14, 259–267 (2010).
Franken, P. & Dijk, D. J. Circadian clock genes and sleep homeostasis. Eur. J. Neurosci. 29, 1820–1829 (2009).
McIntosh, B. E., Hogenesch, J. B. & Bradfield, C. A. Mammalian Per-Arnt-Sim proteins in environmental adaptation. Annu. Rev. Physiol. 72, 625–645 (2010).
Rosato, E., Peixoto, A. A., Costa, R. & Kyriacou, C. P. Linkage disequilibrium, mutational analysis and natural selection in the repetitive region of the clock gene, period, in Drosophila melanogaster. Genet. Res. 69, 89–99 (1997).
Sawyer, L. A. et al. Natural variation in a Drosophila clock gene and temperature compensation. Science 278, 2117–2120 (1997).
Tauber, E. et al. Natural selection favors a newly derived timeless allele in Drosophila melanogaster. Science 316, 1895–1898 (2007).
Debruyne, J. P. et al. A clock shock: mouse CLOCK is not required for circadian oscillator function. Neuron 50, 465–477 (2006).
Dudley, C. A. et al. Altered patterns of sleep and behavioral adaptability in NPAS2-deficient mice. Science 301, 379–383 (2003).
van der Horst, G. T. et al. Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630 (1999).
Cermakian, N., Monaco, L., Pando, M. P., Dierich, A. & Sassone-Corsi, P. Altered behavioral rhythms and clock gene expression in mice with a targeted mutation in the Period1 gene. EMBO J. 20, 3967–3974 (2001).
Zheng, B. et al. The mPer2 gene encodes a functional component of the mammalian circadian clock. Nature 400, 169–173 (1999).
Shearman, L. P., Jin, X., Lee, C., Reppert, S. M. & Weaver, D. R. Targeted disruption of the mPer3 gene: subtle effects on circadian clock function. Mol. Cell. Biol. 20, 6269–6275 (2000).
Preitner, N. et al. The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110, 251–260 (2002).
Sato, T. K. et al. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 43, 527–537 (2004).
Andre, E. et al. Disruption of retinoid-related orphan receptor beta changes circadian behavior, causes retinal degeneration and leads to vacillans phenotype in mice. EMBO J. 17, 3867–3877 (1998).
Bae, K. et al. Differential functions of mPer1, mPer2, and mPer3 in the SCN circadian clock. Neuron 30, 525–536 (2001).
DeBruyne, J. P., Weaver, D. R. & Reppert, S. M. CLOCK and NPAS2 have overlapping roles in the suprachiasmatic circadian clock. Nature Neurosci. 10, 543–545 (2007).
Bunger, M. K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
Harmar, A. J. et al. The VPAC(2) receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 109, 497–508 (2002).
Baggs, J. E et al. Network features of the mammalian circadian clock. PLoS Biol. 7, e52 (2009). This study established the key principles of proportionality and paralogue compensation as common themes in the circadian clock network.
Liu, A. C et al. Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 129, 605–616 (2007). This paper demonstrated that knockout of clock factors resulted in tissue-specific circadian phenotypes and that the SCN was particularly robust to perturbation because of intercellular coupling.
DeBruyne, J. P., Weaver, D. R. & Reppert, S. M. Peripheral circadian oscillators require CLOCK. Curr. Biol. 17, R538–R539 (2007).
Kafri, R., Bar-Even, A. & Pilpel, Y. Transcription control reprogramming in genetic backup circuits. Nature Genet. 37, 295–299 (2005).
Kafri, R., Levy, M. & Pilpel, Y. The regulatory utilization of genetic redundancy through responsive backup circuits. Proc. Natl Acad. Sci. USA 103, 11653–11658 (2006).
Nakahata, Y., Akashi, M., Trcka, D., Yasuda, A. & Takumi, T. The in vitro real-time oscillation monitoring system identifies potential entrainment factors for circadian clocks. BMC Mol. Biol. 7, 5 (2006).
Hirota, T. et al. A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3β. Proc. Natl Acad. Sci. USA 105, 20746–20751 (2008).
Johnson, C. H., Egli, M. & Stewart, P. L. Structural insights into a circadian oscillator. Science 322, 697–701 (2008).
Dong, G. & Golden, S. S. How a cyanobacterium tells time. Curr. Opin. Microbiol. 11, 541–546 (2008).
Markson, J. S. & O'Shea, E. K. The molecular clockwork of a protein-based circadian oscillator. FEBS Lett. 583, 3938–3947 (2009).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005). This work demonstrated that it was possible to reconstitute circadian clock function in vitro with three purified proteins and ATP.
O'Neill, J. S. & Reddy, A. B. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011). This paper shows that circadian oscillations are present in red blood cells, in which transcriptional rhythms do not exist.
Tomita, J., Nakajima, M., Kondo, T. & Iwasaki, H. No transcription-translation feedback in circadian rhythm of KaiC phosphorylation. Science 307, 251–254 (2005).
Xu, Y., Mori, T. & Johnson, C. H. Cyanobacterial circadian clockwork: roles of KaiA, KaiB and the kaiBC promoter in regulating KaiC. EMBO J. 22, 2117–2126 (2003).
Nishiwaki, T., Iwasaki, H., Ishiura, M. & Kondo, T. Nucleotide binding and autophosphorylation of the clock protein KaiC as a circadian timing process of cyanobacteria. Proc. Natl Acad. Sci. USA 97, 495–499 (2000).
Kitayama, Y., Iwasaki, H., Nishiwaki, T. & Kondo, T. KaiB functions as an attenuator of KaiC phosphorylation in the cyanobacterial circadian clock system. EMBO J. 22, 2127–2134 (2003).
Kageyama, H. et al. Cyanobacterial circadian pacemaker: Kai protein complex dynamics in the KaiC phosphorylation cycle in vitro. Mol. Cell 23, 161–171 (2006).
Nishiwaki, T. et al. Role of KaiC phosphorylation in the circadian clock system of Synechococcus elongatus PCC 7942. Proc. Natl Acad. Sci. USA 101, 13927–13932 (2004).
Xu, Y. et al. Identification of key phosphorylation sites in the circadian clock protein KaiC by crystallographic and mutagenetic analyses. Proc. Natl Acad. Sci. USA 101, 13933–13938 (2004).
Rust, M. J., Markson, J. S., Lane, W. S., Fisher, D. S. & O'Shea, E. K. Ordered phosphorylation governs oscillation of a three-protein circadian clock. Science 318, 809–812 (2007). This study showed that ordered phosphorylation events govern oscillations in cyanobacteria clock function in vitro.
Nishiwaki, T. et al. A sequential program of dual phosphorylation of KaiC as a basis for circadian rhythm in cyanobacteria. EMBO J. 26, 4029–4037 (2007).
Ito, H. et al. Autonomous synchronization of the circadian KaiC phosphorylation rhythm. Nature Struct. Mol. Biol. 14, 1084–1088 (2007). Using the purified proteins model of the cyanobacterial clock, this study showed that mixing cultures of different phases results in adoption of one of the two phases. Further, it showed that monomer exchange between KaiC hexamers in the early dephosphorylation phase is linked with synchronization.
Nagai, T., Terada, T. P. & Sasai, M. Synchronization of circadian oscillation of phosphorylation level of KaiC in vitro. Biophys. J. 98, 2469–2477 (2010).
Yang, Q., Pando, B. F., Dong, G., Golden S.S. & van Oudenaarden, A. Circadian gating of the cell cycle revealed in single cyanobacterial cells. Science 327, 1522–1526 (2010).
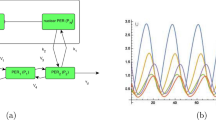
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000). In this study, a synthetic network, termed a repressilator, was developed that can produce synthetic oscillations in transcription.
Müller, S. et al. A generalized model of the repressilator. J. Math. Biol. 53, 905–937 (2006).
Zhu, R., Ribeiro, A. S., Salahub, D. & Kauffman, S. A. Studying genetic regulatory networks at the molecular level: delayed reaction stochastic models. J. Theor. Biol. 246, 725–745 (2007).
Mileyko, Y., Joh, R. I. & Weitz, J. S. Small-scale copy number variation and large-scale changes in gene expression. Proc. Natl Acad. Sci. USA 105, 16659–16664 (2008).
Ukai-Tadenuma, M. et al. Delay in feedback repression by cryptochrome 1 is required for circadian clock function. Cell 144, 268–281 (2011).
Izumo, M., Johnson, C. H. & Yamazaki, S. Circadian gene expression in mammalian fibroblasts revealed by real-time luminescence reporting: temperature compensation and damping. Proc. Natl Acad. Sci. USA 100, 16089–16094 (2003).
Lenhardt, R. & Sessler, D. I. Estimation of mean body temperature from mean skin and core temperature. Anesthesiology 105, 1117–1121 (2006).
Buhr, E. D., Yoo, S. H. & Takahashi, J. S. Temperature as a universal resetting cue for mammalian circadian oscillators. Science 330, 379–385 (2010).
Dunlap, J. C. et al. A circadian clock in Neurospora: how genes and proteins cooperate to produce a sustained, entrainable, and compensated biological oscillator with a period of about a day. Cold Spring Harb. Symp. Quant. Biol. 72, 57–68 (2007).
de Paula, R. M., Vitalini, M. W., Gomer, R. H. & Bell-Pedersen, D. Complexity of the Neurospora crassa circadian clock system: multiple loops and oscillators. Cold Spring Harb. Symp. Quant. Biol. 72, 345–351 (2007).
Hastings, J. W. & Sweeney, B. M. On the mechanism of temperature dependence in a biological clock. Proc. Natl Acad. Sci. USA 43, 804–811 (1957).
Liu, Y., Garceau, N. Y., Loros, J. J. & Dunlap, J. C. Thermally regulated translational control of FRQ mediates aspects of temperature responses in the neurospora circadian clock. Cell 89, 477–486 (1997).
Liu, Y., Merrow, M., Loros, J. J. & Dunlap, J. C. How temperature changes reset a circadian oscillator. Science 281, 825–829 (1998).
Diernfellner, A. C., Schafmeier, T., Merrow, M. W. & Brunner, M. Molecular mechanism of temperature sensing by the circadian clock of Neurospora crassa. Genes Dev. 19, 1968–1973 (2005).
Diernfellner, A. et al. Long and short isoforms of Neurospora clock protein FRQ support temperature-compensated circadian rhythms. FEBS Lett. 581, 5759–5764 (2007).
Mehra, A. et al. A role for casein kinase 2 in the mechanism underlying circadian temperature compensation. Cell 137, 749–760 (2009). This study established the role of CK-2 in the temperature-compensation mechanism in N. crassa.
Gardner, G. F. & Feldman, J. F. Temperature compensation of circadian period length in clock mutants of Neurospora crassa. Plant Physiol. 68, 1244–1248 (1981).
Terauchi, K. et al. ATPase activity of KaiC determines the basic timing for circadian clock of cyanobacteria. Proc. Natl Acad. Sci. USA 104, 16377–16381 (2007).
Isojima, Y. et al. CKIɛ/δ-dependent phosphorylation is a temperature-insensitive, period-determining process in the mammalian circadian clock. Proc. Natl Acad. Sci. USA 106, 15744–15749 (2009). This paper shows that temperature-insensitive enzyme reactions exist in eukaryotic circadian clocks.
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Winzeler, E. A. et al. Functional characterization of the, S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Tucker, C. L. & Fields, S. Quantitative genome-wide analysis of yeast deletion strain sensitivities to oxidative and chemical stress. Comp. Funct. Genomics 5, 216–224 (2004).
Gong, Y. & Zhang, Z. Alternative pathway approach for automating analysis and validation of cell perturbation networks and design of perturbation experiments. Ann. N.Y. Acad. Sci. 1115, 267–285 (2007).
Kundaje, A. et al. A predictive model of the oxygen and heme regulatory network in yeast. PLoS Comput. Biol. 4, e1000224 (2008).
Peleg, T., Yosef, N., Ruppin, E. & Sharan, R. Network-free inference of knockout effects in yeast. PLoS Comput. Biol. 6, e1000635 (2010).
Forger, D. B. Signal processing in cellular clocks. Proc. Natl Acad. Sci. USA 108, 4281–4285 (2011).
Troyanskaya, O. G., Dolinski, K., Owen, A. B., Altman, R. B. & Botstein, D. A Bayesian framework for combining heterogeneous data sources for gene function prediction (in Saccharomyces cerevisiae). Proc. Natl Acad. Sci. USA 100, 8348–8353 (2003).
Savage, R. S., Ghahramani, Z., Griffin J. E., de la Cruz, B. J. & Wild, D. L. Discovering transcriptional modules by Bayesian data integration. Bioinformatics 26, i158–i167 (2010).
Azuaje, F. Advancing post-genome data and system integration through machine learning. Comp. Funct. Genomics 3, 28–31 (2002).
Bidaut, G. & Stoeckert, C. J. Jr. Characterization of unknown adult stem cell samples by large scale data integration and artificial neural networks. Pac. Symp. Biocomput. 2009, 356–367 (2009).
Winfree, A. T. The Geometry of Biological Time 2nd edn (Springer, New York, USA, 2001).
Ruoff, P. Introducing temperature-compensation in any reaction kinetic oscillator model. J. Interdiscipl. Cycle Res. 23, 92–99 (1992).
Forger, D. B., Jewett, M. E. & Kronauer, R. E. A simpler model of the human circadian pacemaker. J. Biol. Rhythms 14, 532–537 (1999).
Goodwin, B.C. Temperature Organization in Cells. A Dynamic Theory of Cellular Control Processes (Academic, New York, USA, 1963).
Kuramoto, Y. Chemical Oscillators, Waves and Turbulence (Springer, Berlin, Germany, 1984).
Goldbeter, A. A model for circadian oscillations in the Drosophila period protein (PER). Proc. Biol. Sci. 261, 319–324 (1995).
Forger, D. B. & Peskin, C. S. A detailed predictive model of the mammalian circadian clock. Proc. Natl Acad. Sci. USA 100, 14806–14811 (2003).
Acknowledgements
The authors would like to thank members of the Hogenesch and Ueda laboratories for their comments on this Review. J.B.H. is supported by the National Heart, Lung and Blood Institute (NHBLI, 1R01HL097800), the National Institute for Neurological Disorders and Stroke (NINDS, 5R01NS054794) and the Pennsylvania Commonwealth Health Research Formula Funds. This work was partly supported by the RIKEN research grant and the Cell Innovation Program from MEXT, Japan.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
FURTHER INFORMATION
John B. Hogenesch's laboratory homepage
Glossary
- Temperature compensation
-
In biological clocks, this is the property by which an increase or decrease in temperature fails to change the period length of the circadian rhythm.
- Robustness
-
The resistance to perturbation by, for example, genetic or environmental factors.
- Poikilotherms
-
Organisms that do not regulate their body temperatures.
- Homeotherms
-
Organisms that regulate their body temperatures.
- Bayesian integration
-
The use of the Bayesian inference — a statistical inference where experimental data is used to infer new or update prior hypotheses — to integrate large-scale, genomic data sets.
- Diapause
-
The suspension of insect development after subjection to adverse environmental conditions.
- Rhythmicity
-
In biological clocks, this is the property by which a molecular, cellular or physiological response recurs with regularity.
- RNA interference
-
(RNAi). A biochemical system in cells that governs how dsRNA can interact with mRNAs to activate or inhibit their message levels or translation.
- Autonomous oscillator
-
In biological clocks, this is a system in which cellular oscillations — rhythmic fluctuations with a definable period length — in gene transcription, reporter gene activity or physiology persist in isolated cells.
- Repressilator
-
This is a synthetic network that generates stable oscillations in GFP.
Rights and permissions
About this article
Cite this article
Hogenesch, J., Ueda, H. Understanding systems-level properties: timely stories from the study of clocks. Nat Rev Genet 12, 407–416 (2011). https://doi.org/10.1038/nrg2972
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2972
This article is cited by
-
The Circadian Clocks, Oscillations of Pain-Related Mediators, and Pain
Cellular and Molecular Neurobiology (2023)
-
Circadian rhythms in septic shock patients
Annals of Intensive Care (2021)
-
Explaining features of fine-grained phenomena using abstract analyses of phenomena and mechanisms: two examples from chronobiology
Synthese (2021)
-
A mobile ELF4 delivers circadian temperature information from shoots to roots
Nature Plants (2020)
-
Synchronization of the Normal Human Peripheral Immune System: A Comprehensive Circadian Systems Immunology Analysis
Scientific Reports (2020)