Key Points
-
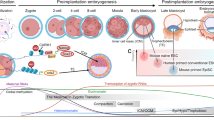
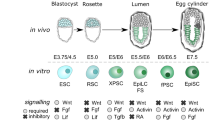
Pluripotency exists transiently in the early embryo and can be recapitulated in vitro.
-
Pluripotency is stabilized by an interconnected network of pluripotency-associated genes.
-
The pluripotency gene regulatory network integrates external signals, and exerts control over the decision between self-renewal and differentiation at the transcriptional, post-transcriptional and epigenetic levels.
-
Diverse pathways — including chromatin-mediated mechanisms, RNA-based regulation and 3D genome organization — work together to maintain pluripotency.
-
Recent evidence of alternative pluripotency states indicates the regulatory flexibility of this network.
Abstract
Pluripotency is a state that exists transiently in the early embryo and, remarkably, can be recapitulated in vitro by deriving embryonic stem cells or by reprogramming somatic cells to become induced pluripotent stem cells. The state of pluripotency, which is stabilized by an interconnected network of pluripotency-associated genes, integrates external signals and exerts control over the decision between self-renewal and differentiation at the transcriptional, post-transcriptional and epigenetic levels. Recent evidence of alternative pluripotency states indicates the regulatory flexibility of this network. Insights into the underlying principles of the pluripotency network may provide unprecedented opportunities for studying development and for regenerative medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ramalho-Santos, M. & Willenbring, H. On the origin of the term “stem cell”. Cell Stem Cell 1, 35–38 (2007).
Pappenheim, A. Über Entwicklung und Ausbildung der Erythroblasten. Virchows Arch. 145, 587–643 (in German) (1896).
Haeckel, E. Anthropogenie; oder, Entwicklungsgeschichte des Menschen 3rd edn (in German) (Wilhelm Engelmann, 1877).
Evans, M. J. The isolation and properties of a clonal tissue culture strain of pluripotent mouse teratoma cells. J. Embryol. Exp. Morphol. 28, 163–176 (1972).
Rosenthal, M. D., Wishnow, R. M. & Sato, G. H. In vitro growth and differentiation of clonal populations of multipotential mouse cells derived from a transplantable testicular teratocarcinoma. J. Natl Cancer Inst. 44, 1001–1014 (1970).
Kahan, B. W. & Ephrussi, B. Developmental potentialities of clonal in vitro cultures of mouse testicular teratoma. J. Natl Cancer Inst. 44, 1015–1036 (1970).
Finch, B. W. & Ephrussi, B. Retention of multiple developmental potentialities by cells of a mouse testicular teratocarcinoma during prolonged culture in vitro and their extinction upon hybridization with cells of permanent lines. Proc. Natl Acad. Sci. USA 57, 615–621 (1967).
Solter, D. From teratocarcinomas to embryonic stem cells and beyond: a history of embryonic stem cell research. Nat. Rev. Genet. 7, 319–327 (2006).
Wu, J., Yamauchi, T. & Izpisua Belmonte, J. C. An overview of mammalian pluripotency. Development 143, 1644–1648 (2016).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl Acad. Sci. USA 78, 7634–7638 (1981). References 10 and 11 are seminal papers that describe the successful derivation and stable culture of ESCs from mouse blastocysts, thereby 'setting the stage' for the study of the PGRN.
Matsui, Y., Zsebo, K. & Hogan, B. L. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell 70, 841–847 (1992).
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA 90, 8424–8428 (1993).
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007). References 14 and 15 describe for the first time the derivation of epiSCs, thereby 'setting the stage' for further study of the PGRN.
Huang, Y., Osorno, R., Tsakiridis, A. & Wilson, V. In vivo differentiation potential of epiblast stem cells revealed by chimeric embryo formation. Cell Rep. 2, 1571–1578 (2012).
Kojima, Y. et al. The transcriptional and functional properties of mouse epiblast stem cells resemble the anterior primitive streak. Cell Stem Cell 14, 107–120 (2014).
Wu, J. et al. An alternative pluripotent state confers interspecies chimaeric competency. Nature 521, 316–321 (2015). This paper describes a new type of PSC that selectively integrates into the posterior proximal region of the post-implantation epiblast and can contribute to interspecific chimaeras when injected into the posterior epiblast of post-implantation mouse embryos.
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006). This paper describes the seminal discovery that PSCs can be induced by reprogramming somatic cells with pluripotency-associated transcription factors.
Gonzalez, F., Boue, S. & Izpisua Belmonte, J. C. Methods for making induced pluripotent stem cells: reprogramming à la carte. Nat. Rev. Genet. 12, 231–242 (2011).
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Ng, H. H. & Surani, M. A. The transcriptional and signalling networks of pluripotency. Nat. Cell Biol. 13, 490–496 (2011).
Scholer, H. R., Hatzopoulos, A. K., Balling, R., Suzuki, N. & Gruss, P. A family of octamer-specific proteins present during mouse embryogenesis: evidence for germline-specific expression of an Oct factor. EMBO J. 8, 2543–2550 (1989).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003).
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell Biol. 9, 625–635 (2007).
Thomson, M. et al. Pluripotency factors in embryonic stem cells regulate differentiation into germ layers. Cell 145, 875–889 (2011).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009).
Suzuki, A. et al. Nanog binds to Smad1 and blocks bone morphogenetic protein-induced differentiation of embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 10294–10299 (2006).
Macarthur, B. D., Ma'ayan, A. & Lemischka, I. R. Systems biology of stem cell fate and cellular reprogramming. Nat. Rev. Mol. Cell Biol. 10, 672–681 (2009).
Orkin, S. H. et al. The transcriptional network controlling pluripotency in ES cells. Cold Spring Harb. Symp. Quant. Biol. 73, 195–202 (2008).
Hackett, J. A. & Surani, M. A. Regulatory principles of pluripotency: from the ground state up. Cell Stem Cell 15, 416–430 (2014).
Adamo, A. et al. LSD1 regulates the balance between self-renewal and differentiation in human embryonic stem cells. Nat. Cell Biol. 13, 652–659 (2011).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005). This seminal paper describes the discovery of the genome-wide co-occupancy of OSN, and suggests that the PGRN consists of autoregulatory and feedforward loops.
Chew, J. L. et al. Reciprocal transcriptional regulation of Pou5f1 and Sox2 via the Oct4/Sox2 complex in embryonic stem cells. Mol. Cell. Biol. 25, 6031–6046 (2005).
Rodda, D. J. et al. Transcriptional regulation of Nanog by OCT4 and SOX2. J. Biol. Chem. 280, 24731–24737 (2005).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Festuccia, N. et al. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 11, 477–490 (2012).
Silva, J. & Smith, A. Capturing pluripotency. Cell 132, 532–536 (2008).
Chen, J. et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell 156, 1274–1285 (2014).
Pan, X. et al. Site-specific disruption of the Oct4–Sox2 interaction reveals coordinated mesendodermal differentiation and the epithelial–mesenchymal transition. J. Biol. Chem. 291, 18353–18369 (2016).
Jerabek, S., Merino, F., Scholer, H. R. & Cojocaru, V. OCT4: dynamic DNA binding pioneers stem cell pluripotency. Biochim. Biophys. Acta 1839, 138–154 (2014).
Jauch, R. et al. Conversion of Sox17 into a pluripotency reprogramming factor by reengineering its association with Oct4 on DNA. Stem Cells 29, 940–951 (2011).
Tapia, N. et al. Dissecting the role of distinct OCT4–SOX2 heterodimer configurations in pluripotency. Sci. Rep. 5, 13533 (2015).
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 38, 431–440 (2006).
Galonska, C., Ziller, M. J., Karnik, R. & Meissner, A. Ground state conditions induce rapid reorganization of core pluripotency factor binding before global epigenetic reprogramming. Cell Stem Cell 17, 462–470 (2015). This paper demonstrates global retargeting of OSN binding and remodelling of the enhancer landscape during the transition between pluripotency states.
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
Fort, A. et al. Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance. Nat. Genet. 46, 558–566 (2014).
Schmidt, D. et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell 148, 335–348 (2012).
Kunath, T. et al. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 134, 2895–2902 (2007).
Yuan, H., Corbi, N., Basilico, C. & Dailey, L. Developmental-specific activity of the FGF-4 enhancer requires the synergistic action of Sox2 and Oct-3. Genes Dev. 9, 2635–2645 (1995).
Tee, W. W., Shen, S. S., Oksuz, O., Narendra, V. & Reinberg, D. Erk1/2 activity promotes chromatin features and RNAPII phosphorylation at developmental promoters in mouse ESCs. Cell 156, 678–690 (2014).
Niwa, H., Burdon, T., Chambers, I. & Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 12, 2048–2060 (1998).
Niwa, H., Ogawa, K., Shimosato, D. & Adachi, K. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 460, 118–122 (2009).
Martello, G., Bertone, P. & Smith, A. Identification of the missing pluripotency mediator downstream of leukaemia inhibitory factor. EMBO J. 32, 2561–2574 (2013).
Ye, S., Li, P., Tong, C. & Ying, Q. L. Embryonic stem cell self-renewal pathways converge on the transcription factor Tfcp2l1. EMBO J. 32, 2548–2560 (2013).
Chen, C. Y. et al. Bcl3 bridges LIF–STAT3 to Oct4 signaling in the maintenance of naive pluripotency. Stem Cells 33, 3468–3480 (2015).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008). This is the original report of the 2i condition and ground-state pluripotency.
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604 (2012).
Boroviak, T., Loos, R., Bertone, P., Smith, A. & Nichols, J. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 16, 516–528 (2014).
Yeo, J. C. et al. Klf2 is an essential factor that sustains ground state pluripotency. Cell Stem Cell 14, 864–872 (2014).
Qiu, D. et al. Klf2 and Tfcp2l1, two Wnt/β-catenin targets, act synergistically to induce and maintain naive pluripotency. Stem Cell Rep. 5, 314–322 (2015).
Kim, S. H. et al. ERK1 phosphorylates Nanog to regulate protein stability and stem cell self-renewal. Stem Cell Res. 13, 1–11 (2014).
Chen, H. et al. Erk signaling is indispensable for genomic stability and self-renewal of mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 112, E5936–E5943 (2015). This study challenges the idea that ERK signalling is dispensable for naive pluripotency by showing that ESCs cannot be maintained following double knockout of Erk1 and Erk2.
Cole, M. F., Johnstone, S. E., Newman, J. J., Kagey, M. H. & Young, R. A. Tcf3 is an integral component of the core regulatory circuitry of embryonic stem cells. Genes Dev. 22, 746–755 (2008).
Martello, G. et al. Esrrb is a pivotal target of the Gsk3/Tcf3 axis regulating embryonic stem cell self-renewal. Cell Stem Cell 11, 491–504 (2012).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Fazzio, T. G., Huff, J. T. & Panning, B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell 134, 162–174 (2008).
Ding, L. et al. Systems analyses reveal shared and diverse attributes of Oct4 regulation in pluripotent cells. Cell Syst. 1, 141–151 (2015).
Hu, G. et al. A genome-wide RNAi screen identifies a new transcriptional module required for self-renewal. Genes Dev. 23, 837–848 (2009). This paper describes a screen that discovered a large number of new genes in the PGRN and a new PGRN module involving CNOT3, TRIM28 and c-MYC.
Li, M., Liu, G. H. & Izpisua Belmonte, J. C. Navigating the epigenetic landscape of pluripotent stem cells. Nat. Rev. Mol. Cell Biol. 13, 524–535 (2012).
Gorkin, D. U., Leung, D. & Ren, B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14, 762–775 (2014).
Wei, Z. et al. Klf4 organizes long-range chromosomal interactions with the Oct4 locus in reprogramming and pluripotency. Cell Stem Cell 13, 36–47 (2013).
Zhang, H. et al. Intrachromosomal looping is required for activation of endogenous pluripotency genes during reprogramming. Cell Stem Cell 13, 30–35 (2013).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013). References 78–80 show that pluripotency-associated genes regulate 3D genome organization, which in turn influences their own expression and cell fate determination.
Rowe, H. M. et al. TRIM28 repression of retrotransposon-based enhancers is necessary to preserve transcriptional dynamics in embryonic stem cells. Genome Res. 23, 452–461 (2013).
Schultz, D. C., Ayyanathan, K., Negorev, D., Maul, G. G. & Rauscher, F. J. 3rd. SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 16, 919–932 (2002).
Cheng, B., Ren, X. & Kerppola, T. K. KAP1 represses differentiation-inducible genes in embryonic stem cells through cooperative binding with PRC1 and derepresses pluripotency-associated genes. Mol. Cell. Biol. 34, 2075–2091 (2014).
Sun, C. et al. Dax1 binds to Oct3/4 and inhibits its transcriptional activity in embryonic stem cells. Mol. Cell. Biol. 29, 4574–4583 (2009).
Fujii, S. et al. Nr0b1 is a negative regulator of Zscan4c in mouse embryonic stem cells. Sci. Rep. 5, 9146 (2015).
Zhang, J. et al. Dax1 and Nanog act in parallel to stabilize mouse embryonic stem cells and induced pluripotency. Nat. Commun. 5, 5042 (2014).
Ding, J. et al. Tex10 coordinates epigenetic control of super-enhancer activity in pluripotency and reprogramming. Cell Stem Cell 16, 653–668 (2015).
Leitch, H. G. et al. Naive pluripotency is associated with global DNA hypomethylation. Nat. Struct. Mol. Biol. 20, 311–316 (2013).
Yamaji, M. et al. PRDM14 ensures naive pluripotency through dual regulation of signaling and epigenetic pathways in mouse embryonic stem cells. Cell Stem Cell 12, 368–382 (2013).
Tu, S. et al. Co-repressor CBFA2T2 regulates pluripotency and germline development. Nature 534, 387–390 (2016).
Tiscornia, G. & Izpisua Belmonte, J. C. MicroRNAs in embryonic stem cell function and fate. Genes Dev. 24, 2732–2741 (2010).
Gangaraju, V. K. & Lin, H. MicroRNAs: key regulators of stem cells. Nat. Rev. Mol. Cell Biol. 10, 116–125 (2009).
Houbaviy, H. B., Murray, M. F. & Sharp, P. A. Embryonic stem cell-specific microRNAs. Dev. Cell 5, 351–358 (2003).
Melton, C., Judson, R. L. & Blelloch, R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 463, 621–626 (2010).
Li, M. & Izpisua Belmonte, J. C. Roles for noncoding RNAs in cell-fate determination and regeneration. Nat. Struct. Mol. Biol. 22, 2–4 (2015).
Gu, K. L. et al. Pluripotency-associated miR-290/302 family of microRNAs promote the dismantling of naive pluripotency. Cell Res. 26, 350–366 (2016).
Sheik Mohamed, J., Gaughwin, P. M., Lim, B., Robson, P. & Lipovich, L. Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. RNA 16, 324–337 (2010).
Loewer, S. et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat. Genet. 42, 1113–1117 (2010). References 97 and 98 highlight the roles of lncRNAs in modulating the PGRN.
Wang, Y. et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 25, 69–80 (2013).
Xu, N., Papagiannakopoulos, T., Pan, G., Thomson, J. A. & Kosik, K. S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137, 647–658 (2009).
Santoni, F. A., Guerra, J. & Luban, J. HERV-H RNA is abundant in human embryonic stem cells and a precise marker for pluripotency. Retrovirology 9, 111 (2012).
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 21, 423–425 (2014).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Salomonis, N. et al. Alternative splicing regulates mouse embryonic stem cell pluripotency and differentiation. Proc. Natl Acad. Sci. USA 107, 10514–10519 (2010).
Gabut, M. et al. An alternative splicing switch regulates embryonic stem cell pluripotency and reprogramming. Cell 147, 132–146 (2011).
Das, S., Jena, S. & Levasseur, D. N. Alternative splicing produces Nanog protein variants with different capacities for self-renewal and pluripotency in embryonic stem cells. J. Biol. Chem. 286, 42690–42703 (2011).
Lu, Y. et al. Alternative splicing of MBD2 supports self-renewal in human pluripotent stem cells. Cell Stem Cell 15, 92–101 (2014).
Han, H. et al. MBNL proteins repress ES-cell-specific alternative splicing and reprogramming. Nature 498, 241–245 (2013). References 107 and 108 show the roles of alternative splicing in regulating pluripotency and self-renewal.
Jia, G., Fu, Y. & He, C. Reversible RNA adenosine methylation in biological regulation. Trends Genet. 29, 108–115 (2013).
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Wang, Y. et al. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat. Cell Biol. 16, 191–198 (2014).
Morgani, S. M. et al. Totipotent embryonic stem cells arise in ground-state culture conditions. Cell Rep. 3, 1945–1957 (2013).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Wu, J. & Izpisua Belmonte, J. C. Dynamic pluripotent stem cell states and their applications. Cell Stem Cell 17, 509–525 (2015).
Tonge, P. D. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014).
Weinberger, L., Ayyash, M., Novershtern, N. & Hanna, J. H. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 17, 155–169 (2016).
Ficz, G. et al. FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 13, 351–359 (2013).
von Meyenn, F. et al. Impairment of DNA methylation maintenance is the main cause of global demethylation in naive embryonic stem cells. Mol. Cell 62, 848–861 (2016).
Murakami, K. et al. NANOG alone induces germ cells in primed epiblast in vitro by activation of enhancers. Nature 529, 403–407 (2016).
Factor, D. C. et al. Epigenomic comparison reveals activation of “seed” enhancers during transition from naive to primed pluripotency. Cell Stem Cell 14, 854–863 (2014).
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014). References 120 and 121 demonstrate the global retargeting of OSN binding and remodelling of the enhancer landscape during the transition between pluripotency states.
Zhang, H. et al. MLL1 inhibition reprograms epiblast stem cells to naive pluripotency. Cell Stem Cell 18, 481–494 (2016).
Li, M., Suzuki, K., Kim, N. Y., Liu, G. H. & Izpisua Belmonte, J. C. A cut above the rest: targeted genome editing technologies in human pluripotent stem cells. J. Biol. Chem. 289, 4594–4599 (2014).
Suzuki, K. et al. Targeted gene correction minimally impacts whole-genome mutational load in human-disease-specific induced pluripotent stem cell clones. Cell Stem Cell 15, 31–36 (2014).
Li, M. et al. Efficient correction of hemoglobinopathy-causing mutations by homologous recombination in integration-free patient iPSCs. Cell Res. 21, 1740–1744 (2011).
Li, M. & Izpisua Belmonte, J. C. Looking to the future following 10 years of induced pluripotent stem cell technologies. Nat. Protoc. 11, 1579–1585 (2016).
Goolam, M. et al. Heterogeneity in Oct4 and Sox2 targets biases cell fate in 4-cell mouse embryos. Cell 165, 61–74 (2016).
Kolodziejczyk, A. A. et al. Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17, 471–485 (2015).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Guo, G. et al. Serum-based culture conditions provoke gene expression variability in mouse embryonic stem cells as revealed by single-cell analysis. Cell Rep. 14, 956–965 (2016).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Apostolou, E. et al. Genome-wide chromatin interactions of the Nanog locus in pluripotency, differentiation, and reprogramming. Cell Stem Cell 12, 699–712 (2013).
Acknowledgements
The authors apologize to the colleagues whose work is not covered in this article because of space constraints. The authors would like to thank D. O'Keefe and M. Schwarz for critical reading and generous help during the preparation of the manuscript. Work in the laboratory of J.C.I.B. is supported by the G. Harold and Leila Y. Mathers Charitable Foundation, The Leona M. and Harry B. Helmsley Charitable Trust (grant 2012-PG-MED002), the Moxie Foundation, Fundacion Dr. Pedro Guillen and the Universidad Católica San Antonio de Murcia (UCAM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Inner cell mass
-
(ICM). A small cluster of cells inside the early embryo (that is, the blastocyst). These cells give rise to all the tissues of the future embryo but not to extra-embryonic tissues (for example, the placenta). The ICM may be isolated to generate embryonic stem cells.
- Trophectoderm
-
The outer cell layer of a blastocyst. It is formed from the first specialized lineage of cells and gives rise to extra-embryonic tissues.
- Mesendoderm
-
A layer of cells that are formed during early gastrulation, and are destined to become mesoderm and endoderm.
- Ectoderm
-
The outermost of the three germ layers that are formed during gastrulation of the early embryo. Ectoderm-derived tissues include the nervous system, sensory organs and the skin.
- Stemness genes
-
Genes that constitute the stem cell-specific gene expression programme.
- Epithelial–mesenchymal transition
-
(EMT). A process in which cells of an epithelial layer lose their polarity and cell–cell adhesion, and become disorganized migratory mesenchymal cells. EMT is an integral part of normal developmental, wound healing and cancer development.
Rights and permissions
About this article
Cite this article
Li, M., Belmonte, J. Ground rules of the pluripotency gene regulatory network. Nat Rev Genet 18, 180–191 (2017). https://doi.org/10.1038/nrg.2016.156
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.156
This article is cited by
-
Boldine promotes stemness of human urine-derived stem cells by activating the Wnt/β-catenin signaling pathway
Molecular and Cellular Biochemistry (2024)
-
Ubiquitination-specific protease 7 enhances stemness of hepatocellular carcinoma by stabilizing basic transcription factor 3
Functional & Integrative Genomics (2024)
-
Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Nature Communications (2024)
-
Ascl1 and Ngn2 convert mouse embryonic stem cells to neurons via functionally distinct paths
Nature Communications (2023)
-
A multi-omics integrative analysis based on CRISPR screens re-defines the pluripotency regulatory network in ESCs
Communications Biology (2023)