Abstract
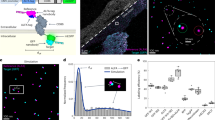
Studies that rely on fluorescence imaging of nonadherent cells that are cultured in suspension, such as Escherichia coli, are often hampered by trade-offs that must be made between data throughput and imaging resolution. We developed a platform for microfluidics-assisted cell screening (MACS) that overcomes this trade-off by temporarily immobilizing suspension cells within a microfluidics chip. This enables high-throughput and automated single-cell microscopy for a wide range of cell types and sizes. As cells can be rapidly sampled directly from a suspension culture, MACS bypasses the need for sample preparation, and therefore allows measurements without perturbing the native cell physiology. The setup can also be integrated with complex growth chambers, and can be used to enrich or sort the imaged cells. Furthermore, MACS facilitates the visualization of individual cytoplasmic fluorescent proteins (FPs) in E. coli, allowing low-abundance proteins to be counted using standard total internal reflection fluorescence (TIRF) microscopy. Finally, MACS can be used to impart mechanical pressure for assessing the structural integrity of individual cells and their response to mechanical perturbations, or to make cells take up chemicals that otherwise would not pass through the membrane. This protocol describes the assembly of electronic control circuitry, the construction of liquid-handling components and the creation of the MACS microfluidics chip. The operation of MACS is described, and automation software is provided to integrate MACS control with image acquisition. Finally, we provide instructions for extending MACS using an external growth chamber (1 d) and for how to sort rare cells of interest.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Tracy, B.P., Gaida, S.M. & Papoutsakis, E.T. Flow cytometry for bacteria: enabling metabolic engineering, synthetic biology and the elucidation of complex phenotypes. Curr. Opin. Biotechnol. 21, 85–99 (2010).
Bongaerts, R.J., Hautefort, I., Sidebotham, J.M. & Hinton, J.C. Green fluorescent protein as a marker for conditional gene expression in bacterial cells. Methods Enzymol. 358, 43–66 (2002).
Felip, M., Andreatta, S., Sommaruga, R., Straskrábová, V. & Catalan, J. Suitability of flow cytometry for estimating bacterial biovolume in natural plankton samples: comparison with microscopy data. Appl. Environ. Microbiol. 73, 4508–4514 (2007).
Cisse, I.I. et al. Real-time dynamics of RNA polymerase II clustering in live human cells. Science 341, 664–667 (2013).
Okumus, B., Yildiz, S. & Toprak, E. Fluidic and microfluidic tools for quantitative systems biology. Curr. Opin. Biotechnol. 25, 30–38 (2014).
Wang, P. et al. Robust growth of Escherichia coli. Curr. Biol. 20, 1099–1103 (2010).
Sheats, J., Sclavi, B., Cosentino Lagomarsino, M., Cicuta, P. & Dorfman, K.D. Role of growth rate on the orientational alignment of Escherichia coli in a slit. R. Soc. Open Sci. 4, 170463 (2017).
Xie, X.S., Choi, P.J., Li, G.-W., Lee, N.K. & Lia, G. Single-molecule approach to molecular biology in living bacterial cells. Annu. Rev. Biophys. 37, 417–444 (2008).
Li, G.-W., Burkhardt, D., Gross, C. & Weissman, J.S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Uphoff, S. et al. Stochastic activation of a DNA damage response causes cell-to-cell mutation rate variation. Science 351, 1094–1097 (2016).
Okumus, B. et al. Mechanical slowing-down of cytoplasmic diffusion allows in vivo counting of proteins in individual cells. Nat. Commun. 7, 11641 (2016).
Unger, M.A. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000).
Studer, V. Scaling properties of a low-actuation pressure microfluidic valve. J. Appl. Phys. 95, 393 (2004).
Mazutis, L. et al. Single-cell analysis and sorting using droplet-based microfluidics. Nat. Protoc. 8, 870–891 (2013).
Parry, B.R. et al. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 156, 183–194 (2013).
Elf, J., Li, G.W. & Xie, X.S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Frost, L.S. & Koraimann, G. Regulation of bacterial conjugation: balancing opportunity with adversity. Future Microbiol. 5, 1057–1071 (2010).
Lindqvist, R.C. & Nordström, K. Resistance of Escherichia coli to penicillins VII. Purification and characterization of a penicillinase mediated by the R factor R1. J. Bacteriol. 101, 232–239 (1970).
Veres, A. et al. Building a morbidostat: an automated continuous-culture device for studying bacterial drug resistance under dynamically sustained drug inhibition. Nat. Protoc. 8, 555–567 (2013).
Acknowledgements
We thank M. Cokol, R. Fernandez-Lopez, S. Yildiz, M. El Karoui and E. Toprak for their helpful discussions and useful suggestions. We are grateful to P. Gorelik and O. Mazor at the HMS Research Instrumentation Core Facility for their assistance in instrument design and fabrication. We thank C. Saenz and V. Lien at the Microfluidics Core Facility at Harvard Medical School for the fabrication of and testing process for the microfluidics chips. J.C.A.-C. thanks the School of Sciences at Universidad de los Andes for PhD support grant 2015-1. J.P. acknowledges support from NIH grants GM081563 and GM09578, and B.O. acknowledges support from a Novartis Fellowship in Systems Biology.
Author information
Authors and Affiliations
Contributions
B.O. and C.J.B., J.C.A.-C., G.C.L., S.L. and D.L. performed the experiments described in this protocol, C.J.B. provided the control and analysis codes and built the system, S.B. wrote the spot-detection algorithm, D.L. wrote the segmentation algorithm, E.L. designed the growth chamber and wrote the optical density measurement code. B.O., C.J.B., J.C.A.-C., E.L. and J.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
Patents were filed on behalf of B.O., D.L., C.J.B., G.C.L. and J.P. by the President and Fellows of Harvard College.
Integrated supplementary information
Supplementary Figure 1 Images showing the assembly of the electronic control base.
Images showing (a) the top of the MOSFET circuit board, (b) the bottom of the MOSFET circuit board and (c) the fully assembled Electronic Control Base.
Supplementary Figure 2 Exploded-view diagram of the electronic control base.
Fasteners required to connect the components to the acrylic base are listed in the table above. For clarity, the second pressure regulator is omitted from the view. It is attached in an identical fashion to the pressure regulator shown in the back of the diagram.
Supplementary Figure 3 Exploded-view diagram of the pinch valve base.
Fasteners required to assemble the acrylic components are listed in the table above. For clarity, the connection for only one of the pinch valves is shown. The others are attached in an identical manner to the one shown.
Supplementary Figure 4 Exploded-view diagram of the cleaning reservoir stand.
Fasteners required to assemble the acrylic components are listed in the table above.
Supplementary Figure 5 Images of the construction of the pressure tubes.
Images showing (a) the pressure tube cap surrounded with the tape cup, (b) the bottom of the pressure tube with a 9/64” hole drilled in the base and surrounded with the tape cup, (c) the cap of the pressure tube after being filled with epoxy and (d) the bottom of the pressure tube after being filled with epoxy.
Supplementary Figure 6 Images of the pinch valve base assembly.
Images of the (a) silicone pinch valve tubing assembled for the Pinch Valve Base and (b) the Pinch Valve Base with the pinch valves and pressure tubes attached.
Supplementary Figure 7 Images showing the assembly of the growth chamber.
Images showing (a) the LED holder attached to acrylic panel, (b) the assembled emitter/detector acrylic sides connected with middle shelf, (c) the fully assembled growth vial holder top with tape to aid with gluing, (d) the assembled growth vial holder base, i.e. stirrer, (e) the connection between the base and top of the vial holder, (f) the LED emitter and photodiode detector attached to the acrylic vial holder and (g) an electronic diagram of the connections required to wire up the stirrer and OD measurement, image created using Fritzing.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and the Supplementary Tutorial. (PDF 1766 kb)
Supplementary Data 1
AutoCAD design files for the transparency masks used to make the silicon masters. (ZIP 386 kb)
Supplementary Data 2
Gerber files for manufacturing the MOSFET printed circuit board. (ZIP 78 kb)
Supplementary Data 3
Code required to run MACS via Micro-Manager. (ZIP 538 kb)
Supplementary Data 4
Source code for the MACS Controller Micro-Manager plugin. (ZIP 1233 kb)
Supplementary Data 5
Laser cutter design file for various acrylic components needed to construct the standard MACS. (ZIP 1181 kb)
Supplementary Data 6
Laser cutter design file for acrylic components required to build the optional growth chamber. (ZIP 651 kb)
Supplementary Data 7
Laser cutter design file for the optional cover-slide stage-holder adaptor. (ZIP 1026 kb)
Supplementary Data 8
Code required to run OD measurement for the optional growth chamber. (ZIP 73 kb)
Supplementary Data 9
MATLAB code for analysis. (ZIP 13485 kb)
Rights and permissions
About this article
Cite this article
Okumus, B., Baker, C., Arias-Castro, J. et al. Single-cell microscopy of suspension cultures using a microfluidics-assisted cell screening platform. Nat Protoc 13, 170–194 (2018). https://doi.org/10.1038/nprot.2017.127
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.127
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.