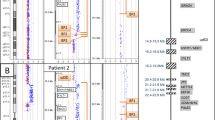
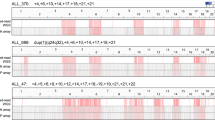
Abstract
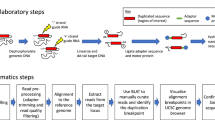
The most common recurrent copy-number variants associated with autism, developmental delay and epilepsy are flanked by segmental duplications. Complete genetic characterization of these events is challenging because their breakpoints often occur within high-identity, copy-number polymorphic paralogous sequences that cannot be specifically assayed using hybridization-based methods. Here we provide a protocol for breakpoint resolution with sequence-level precision. Massively parallel sequencing is performed on libraries generated from haplotype-resolved chromosomes, genomic DNA or molecular inversion probe (MIP)-captured breakpoint-informative regions harboring paralog-distinguishing variants. Quantification of sequencing depth over informative sites enables breakpoint localization, typically within several kilobases to tens of kilobases. Depending on the approach used, the sequencing platform, and the accuracy and completeness of the reference genome sequence, this protocol takes from a few days to several months to complete. Once established for a specific genomic disorder, it is possible to process thousands of DNA samples within as little as 3–4 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Inoue, K. & Lupski, J.R. Molecular mechanisms for genomic disorders. Annu. Rev. Genomics Hum. Genet. 3, 199–242 (2002).
Stankiewicz, P. & Lupski, J.R. Genome architecture, rearrangements and genomic disorders. Trends Genet. 18, 74–82 (2002).
Bailey, J.A., Yavor, A.M., Massa, H.F., Trask, B.J. & Eichler, E.E. Segmental duplications: organization and impact within the current human genome project assembly. Genome Res. 11, 1005–1017 (2001).
Stankiewicz, P. & Lupski, J.R. Structural variation in the human genome and its role in disease. Annu. Rev. Med. 61, 437–455 (2010).
Lupski, J.R. et al. DNA duplication associated with Charcot-Marie-Tooth disease type 1A. Cell 66, 219–232 (1991).
Pinkel, D. et al. GH (CGH). US Patent 5,976,790 (1999).
Wang, N.J. et al. High-resolution molecular characterization of 15q11-q13 rearrangements by array CGH (array CGH) with detection of gene dosage. Am. J. Hum. Genet. 75, 267–281 (2004).
Sahoo, T. et al. Prader-Willi phenotype caused by paternal deficiency for the HBII-85 C/D box small nucleolar RNA cluster. Nat. Genet. 40, 719–721 (2008).
Korbel, J.O. et al. Paired-end mapping reveals extensive structural variation in the human genome. Science 318, 420–426 (2007).
Mills, R.E. et al. Mapping copy number variation by population-scale genome sequencing. Nature 470, 59–65 (2011).
Sharp, A.J. et al. Segmental duplications and copy number variation in the human genome. Am. J. Hum. Genet. 77, 78–88 (2005).
Zody, M.C. et al. Evolutionary toggling of the MAPT 17q21.31 inversion region. Nat. Genet. 40, 1076–1083 (2008).
Girirajan, S. & Eichler, E.E. Phenotypic variability and genetic susceptibility to genomic disorders. Hum. Mol. Genet. 19, R176–R187 (2010).
Itsara, A. et al. Resolving the breakpoints of the 17q21.31 microdeletion syndrome with next-generation sequencing. Am. J. Hum. Genet. 90, 599–613 (2012).
Highsmith, W.E., Meyer, K.J., Marley, V.M. & Jenkins, R.B. Conversion technology for the separation of maternal and paternal copies of any autosomal chromosome in somatic cell hybrids. Curr. Prot. Hum. Genet. 3, 3.6 (2007).
van den Ijssel, P. & Ylstra, B. Oligonucleotide array CGH. in Methods in Molecular Biology vol. 396: Comparative Genomics, Volume 2 (ed. Bergman N.H.) 207-221 (Humana Press, 2007).
Horvath, J.E., Schwartz, S. & Eichler, E.E. The mosaic structure of human pericentromeric DNA: a strategy for characterizing complex regions of the human genome. Genome Res. 10, 839–852 (2000).
Saxena, R. et al. Four DAZ genes in two clusters found in the AZFc region of the human Y chromosome. Genomics 67, 256–267 (2000).
Tilford, C.A. et al. A physical map of the human Y chromosome. Nature 409, 943–945 (2001).
Estivill, X. et al. Chromosomal regions containing high-density and ambiguously mapped putative single-nucleotide polymorphisms (SNPs) correlate with segmental duplications in the human genome. Hum. Mol. Genet. 11, 1987–1995 (2002).
Fredman, D. et al. Complex SNP-related sequence variation in segmental genome duplications. Nat Genet. 36, 861–866 (2004).
Sudmant, P.H. et al. Diversity of human copy number variation and multicopy genes. Science 330, 641–646 (2010).
Nuttle, X. et al. Rapid and accurate large-scale genotyping of duplicated genes and discovery of interlocus gene conversions. Nat. Methods 10, 903–909 (2013).
Dennis, M.Y. et al. Evolution of human-specific neural SRGAP2 genes by incomplete segmental duplication. Cell 149, 912–922 (2012).
Bovee, D. et al. Closing gaps in the human genome with fosmid resources generated from multiple individuals. Nat. Genet. 40, 96–101 (2008).
Genovese, G. et al. Using population admixture to help complete maps of the human genome. Nat. Genet. 45, 406–414 (2013).
Carlson, C. et al. Molecular definition of 22q11 deletions in 151 velo-cardio-facial syndrome patients. Am. J. Hum. Genet. 61, 620–629 (1997).
Kitzman, J.O. et al. Haplotype-resolved genome sequencing of a Gujarati Indian individual. Nat. Biotechnol. 29, 59–63 (2011).
Peters, B.A. et al. Accurate whole-genome sequencing and haplotyping from 10 to 20 human cells. Nature 487, 190–195 (2012).
Porreca, G.J. et al. Multiplex amplification of large sets of human exons. Nat. Methods 4, 931–936 (2007).
Turner, E.H. et al. Massively parallel exon capture and library-free resequencing across 16 genomes. Nat. Methods 6, 315–316 (2009).
O'Roak, B.J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
Renshaw, M.A., Giresi, M. & Adams, J.O. Microsatellite fragment analysis using the ABI PRISM 377 DNA sequencer. in Methods in Molecular Biology, vol. 1006: Microsatellites (ed. Kantartzi, S.K.) 181–196 (Humana Press, 2013).
Broman, K.W., Murray, J.C., Sheffield, V.C., White, R.L. & Weber, J.L. Comprehensive human genetic maps: individual and sex-specific variation in recombination. Am. J. Hum. Genet. 63, 861–869 (1998).
Alkan, C. et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 41, 1061–1067 (2009).
Tuzun, E., Bailey, J.A. & Eichler, E.E. Recent segmental duplications in the working draft assembly of the brown Norway rat. Genome Res. 14, 493–506 (2004).
Kidd, J.M. et al. Characterization of missing human genome sequences and copy number polymorphic insertions. Nat. Methods 7, 365–371 (2010).
Chin, C.S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Acknowledgements
We thank J. Huddleston for assistance in testing analysis software and preparing it for public access; P. Sudmant for analyzing the number of SUNKs in windows across the reference genome; and T. Brown for assistance with manuscript preparation. X.N. is supported by a National Science Foundation Graduate Research Fellowship under grant no. DGE-1256082. This work was supported by US National Institutes of Health grants HG004120 and HG002385 to E.E.E. E.E.E. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
X.N., A.I., J.S. and E.E.E. developed the protocol. X.N. and E.E.E. wrote the paper, with input and approval from all coauthors.
Corresponding author
Ethics declarations
Competing interests
E.E.E. is on the scientific advisory board (SAB) of DNAnexus and was an SAB member of Pacific Biosciences (2009–2013) and SynapDx (2011–2013). The remaining authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Primers used for MIP experiments. (XLSX 33 kb)
Rights and permissions
About this article
Cite this article
Nuttle, X., Itsara, A., Shendure, J. et al. Resolving genomic disorder–associated breakpoints within segmental DNA duplications using massively parallel sequencing. Nat Protoc 9, 1496–1513 (2014). https://doi.org/10.1038/nprot.2014.096
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.096
This article is cited by
-
16p13.11p11.2 triplication syndrome: a new recognizable genomic disorder characterized by optical genome mapping and whole genome sequencing
European Journal of Human Genetics (2022)
-
A Combined Proteomics and Metabolomics Profiling to Investigate the Genetic Heterogeneity of Autistic Children
Molecular Neurobiology (2022)
-
Possible association of 16p11.2 copy number variation with altered lymphocyte and neutrophil counts
npj Genomic Medicine (2022)
-
Copy number variation of the REXO1L1 gene cluster; euchromatic deletion variant or susceptibility factor?
European Journal of Human Genetics (2017)
-
An exploratory study of predisposing genetic factors for DiGeorge/velocardiofacial syndrome
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.