Abstract
The thalamus connects the cortex with other brain regions and supports sensory perception, movement, and cognitive function via numerous distinct nuclei. However, the mechanisms underlying the development and organization of diverse thalamic nuclei remain largely unknown. Here we report an intricate ontogenetic logic of mouse thalamic structures. Individual radial glial progenitors in the developing thalamus actively divide and produce a cohort of neuronal progeny that shows striking spatial configuration and nuclear occupation related to functionality. Whereas the anterior clonal cluster displays relatively more tangential dispersion and contributes predominantly to nuclei with cognitive functions, the medial ventral posterior clonal cluster forms prominent radial arrays and contributes mostly to nuclei with sensory- or motor-related activities. Moreover, the first-order and higher-order sensory and motor nuclei across different modalities are largely segregated clonally. Notably, sonic hedgehog signaling activity influences clonal spatial distribution. Our study reveals lineage relationship to be a critical regulator of nonlaminated thalamus development and organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jones, E.G. The Thalamus (Cambridge University Press, 2007).
Steriade, M. & Llinás, R.R. The functional states of the thalamus and the associated neuronal interplay. Physiol. Rev. 68, 649–742 (1988).
Sherman, S.M. & Guillery, R.W. Exploring the Thalamus and its Role in Cortical Function (MIT Press, 2006).
Sherman, S.M. Thalamus plays a central role in ongoing cortical functioning. Nat. Neurosci. 19, 533–541 (2016).
O'Leary, D.D., Schlaggar, B.L. & Tuttle, R. Specification of neocortical areas and thalamocortical connections. Annu. Rev. Neurosci. 17, 419–439 (1994).
Hunnicutt, B.J. et al. A comprehensive thalamocortical projection map at the mesoscopic level. Nat. Neurosci. 17, 1276–1285 (2014).
Oh, S.W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Sherman, S.M. & Guillery, R.W. Functional Connetions of Cortical Areas (MIT Press, 2013).
Scholpp, S. & Lumsden, A. Building a bridal chamber: development of the thalamus. Trends Neurosci. 33, 373–380 (2010).
Nakagawa, Y. & Shimogori, T. Diversity of thalamic progenitor cells and postmitotic neurons. Eur. J. Neurosci. 35, 1554–1562 (2012).
Arcelli, P., Frassoni, C., Regondi, M.C., De Biasi, S. & Spreafico, R. GABAergic neurons in mammalian thalamus: a marker of thalamic complexity? Brain Res. Bull. 42, 27–37 (1997).
Vue, T.Y. et al. Characterization of progenitor domains in the developing mouse thalamus. J. Comp. Neurol. 505, 73–91 (2007).
Inamura, N., Ono, K., Takebayashi, H., Zalc, B. & Ikenaka, K. Olig2 lineage cells generate GABAergic neurons in the prethalamic nuclei, including the zona incerta, ventral lateral geniculate nucleus and reticular thalamic nucleus. Dev. Neurosci. 33, 118–129 (2011).
Rakic, P. Specification of cerebral cortical areas. Science 241, 170–176 (1988).
Yu, Y.C., Bultje, R.S., Wang, X. & Shi, S.H. Specific synapses develop preferentially among sister excitatory neurons in the neocortex. Nature 458, 501–504 (2009).
Yu, Y.C. et al. Preferential electrical coupling regulates neocortical lineage-dependent microcircuit assembly. Nature 486, 113–117 (2012).
Li, Y. et al. Clonally related visual cortical neurons show similar stimulus feature selectivity. Nature 486, 118–121 (2012).
He, S., Li, Z., Ge, S., Yu, Y.C. & Shi, S.H. Inside-out radial migration facilitates lineage-dependent neocortical microcircuit assembly. Neuron 86, 1159–1166 (2015).
McCormick, D.A., McGinley, M.J. & Salkoff, D.B. Brain state dependent activity in the cortex and thalamus. Curr. Opin. Neurobiol. 31, 133–140 (2015).
Lien, A.D. & Scanziani, M. Tuned thalamic excitation is amplified by visual cortical circuits. Nat. Neurosci. 16, 1315–1323 (2013).
Golden, J.A. & Cepko, C.L. Clones in the chick diencephalon contain multiple cell types and siblings are widely dispersed. Development 122, 65–78 (1996).
Golden, J.A., Zitz, J.C., McFadden, K. & Cepko, C.L. Cell migration in the developing chick diencephalon. Development 124, 3525–3533 (1997).
Zong, H., Espinosa, J.S., Su, H.H., Muzumdar, M.D. & Luo, L. Mosaic analysis with double markers in mice. Cell 121, 479–492 (2005).
Gao, P. et al. Deterministic progenitor behavior and unitary production of neurons in the neocortex. Cell 159, 775–788 (2014).
Ciceri, G. et al. Lineage-specific laminar organization of cortical GABAergic interneurons. Nat. Neurosci. 16, 1199–1210 (2013).
Imayoshi, I., Ohtsuka, T., Metzger, D., Chambon, P. & Kageyama, R. Temporal regulation of Cre recombinase activity in neural stem cells. Genesis 44, 233–238 (2006).
Hippenmeyer, S. et al. Genetic mosaic dissection of Lis1 and Ndel1 in neuronal migration. Neuron 68, 695–709 (2010).
Vue, T.Y. et al. Sonic hedgehog signaling controls thalamic progenitor identity and nuclei specification in mice. J. Neurosci. 29, 4484–4497 (2009).
Wang, L., Bluske, K.K., Dickel, L.K. & Nakagawa, Y. Basal progenitor cells in the embryonic mouse thalamus - their molecular characterization and the role of neurogenins and Pax6. Neural Dev. 6, 35 (2011).
Clascá, F., Rubio-Garrido, P. & Jabaudon, D. Unveiling the diversity of thalamocortical neuron subtypes. Eur. J. Neurosci. 35, 1524–1532 (2012).
Diggle, P.J. Statistical Analysis of Spatial Point Patterns (Oxford University Press, 2003).
Kuramoto, E. et al. Ventral medial nucleus neurons send thalamocortical afferents more widely and more preferentially to layer 1 than neurons of the ventral anterior-ventral lateral nuclear complex in the rat. Cereb. Cortex 25, 221–235 (2015).
Van der Werf, Y.D., Witter, M.P. & Groenewegen, H.J. The intralaminar and midline nuclei of the thalamus. Anatomical and functional evidence for participation in processes of arousal and awareness. Brain Res. Brain Res. Rev. 39, 107–140 (2002).
Szabó, N.E., Zhao, T., Zhou, X. & Alvarez-Bolado, G. The role of Sonic hedgehog of neural origin in thalamic differentiation in the mouse. J. Neurosci. 29, 2453–2466 (2009).
Jeong, J., Mao, J., Tenzen, T., Kottmann, A.H. & McMahon, A.P. Hedgehog signaling in the neural crest cells regulates the patterning and growth of facial primordia. Genes Dev. 18, 937–951 (2004).
Puelles, L. & Rubenstein, J.L. Expression patterns of homeobox and other putative regulatory genes in the embryonic mouse forebrain suggest a neuromeric organization. Trends Neurosci. 16, 472–479 (1993).
Nakagawa, Y. & O'Leary, D.D. Dynamic patterned expression of orphan nuclear receptor genes RORα and RORβ in developing mouse forebrain. Dev. Neurosci. 25, 234–244 (2003).
Frassoni, C., Amadeo, A., Ortino, B., Jaranowska, A. & Spreafico, R. Organization of radial and non-radial glia in the developing rat thalamus. J. Comp. Neurol. 428, 527–542 (2000).
Nakagawa, Y. & O'Leary, D.D. Combinatorial expression patterns of LIM-homeodomain and other regulatory genes parcellate developing thalamus. J. Neurosci. 21, 2711–2725 (2001).
Suzuki-Hirano, A. et al. Dynamic spatiotemporal gene expression in embryonic mouse thalamus. J. Comp. Neurol. 519, 528–543 (2011).
Kataoka, A. & Shimogori, T. Fgf8 controls regional identity in the developing thalamus. Development 135, 2873–2881 (2008).
Chatterjee, M. & Li, J.Y. Patterning and compartment formation in the diencephalon. Front. Neurosci. 6, 66 (2012).
Saalmann, Y.B. & Kastner, S. Cognitive and perceptual functions of the visual thalamus. Neuron 71, 209–223 (2011).
Roth, M.M. et al. Thalamic nuclei convey diverse contextual information to layer 1 of visual cortex. Nat. Neurosci. 19, 299–307 (2016).
Crabtree, J.W., Collingridge, G.L. & Isaac, J.T. A new intrathalamic pathway linking modality-related nuclei in the dorsal thalamus. Nat. Neurosci. 1, 389–394 (1998).
Shu, Y. & McCormick, D.A. Inhibitory interactions between ferret thalamic reticular neurons. J. Neurophysiol. 87, 2571–2576 (2002).
Lee, S.C., Patrick, S.L., Richardson, K.A. & Connors, B.W. Two functionally distinct networks of gap junction-coupled inhibitory neurons in the thalamic reticular nucleus. J. Neurosci. 34, 13170–13182 (2014).
Molnár, Z. et al. Normal development of embryonic thalamocortical connectivity in the absence of evoked synaptic activity. J. Neurosci. 22, 10313–10323 (2002).
Frangeul, L. et al. A cross-modal genetic framework for the development and plasticity of sensory pathways. Nature 538, 96–98 (2016).
Grant, E., Hoerder-Suabedissen, A. & Molnár, Z. The regulation of corticofugal fiber targeting by retinal inputs. Cereb. Cortex 26, 1336–1348 (2016).
Xu, H.T. et al. Distinct lineage-dependent structural and functional organization of the hippocampus. Cell 157, 1552–1564 (2014).
Brown, K.N. et al. Clonal production and organization of inhibitory interneurons in the neocortex. Science 334, 480–486 (2011).
Ha, G.E. et al. The Ca2+-activated chloride channel anoctamin-2 mediates spike-frequency adaptation and regulates sensory transmission in thalamocortical neurons. Nat. Commun. 7, 13791 (2016).
Bombeiro, A.L. et al. MHC-I and PirB upregulation in the central and peripheral nervous system following sciatic nerve injury. PLoS One 11, e0161463 (2016).
Tan, X. et al. vascular influence on ventral telencephalic progenitors and neocortical interneuron production. Dev. Cell 36, 624–638 (2016).
Blaess, S. et al. Temporal-spatial changes in Sonic Hedgehog expression and signaling reveal different potentials of ventral mesencephalic progenitors to populate distinct ventral midbrain nuclei. Neural Dev. 6, 29 (2011).
Acknowledgements
We thank A.L. Joyner (Memorial Sloan Kettering Cancer Center), R. Kageyama (Kyoto University), S. Hippenmeyer (The Institute of Science and Technology), and Y. Nakagawa (University of Minnesota) for providing the R26LSL-SmoM2-EYFP, Nes-CreERT2, MADM11, and Olig3-Cre mouse lines, respectively; J.N. Betley (University of Pennsylvania) for guinea pig anti-RFP antibody, and members of S.-H.S.'s laboratory for insightful discussion and input. This work was supported by the US National Institutes of Health (R01DA024681 and R01MH101382 to S.-H.S. and P30CA008748 to Memorial Sloan Kettering Cancer Center Core Facilities), the Human Frontier Science Program (RGP0053 to S.-H.S. and K.H.), and the National Natural Science Foundation of China (61572265 to Z.H. and 759881 to S.-H.S.).
Author information
Authors and Affiliations
Contributions
W.S. and S.-H.S. conceived the project; W.S. collected, reconstructed and analyzed MADM and retroviral labeling data with help from A.X.; Z.H., X.T., and K.H. performed NND analysis; Z.L. generated retrovirus; A.X., Z.H., and K.H. performed alignment and nuclear identity inference and clustering analysis with input from H.Z. and T.M.; H.Z. and T.M. provided thalamocortical axonal projection data; W.S. and S.-H.S wrote the paper with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 MADM labeling of thalamic clones.
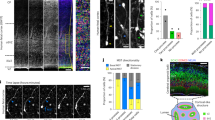
(a) Confocal images of an E12 Nestin-CreERT2;Ai9-tdTomato brain treated with TM at E10 and stained for BLBP (green), a radial glial progenitor-specific marker, tdTomato (red), and OLIG3 (white), and with DAPI (blue). A schematic of brain structures is shown at the top. Note that BLBP-expressing radial glial progenitors in the developing thalamus marked by OLIG3 expression are labeled by tdTomato upon TM treatment, indicating that CreERT2 is expressed in thalamic radial glial progenitors in the Nestin-CreERT2 mouse line. Scale bar: 100 μm. (b) Schematic of MADM labeling. (c) Experimental paradigm of MADM-based clonal analysis. (d) Confocal images of an E12 brain treated with TM at E10 and stained for EGFP (green) and tdTomato (red), and with DAPI (blue). High magnification image of a G2-Z yellow clone in the thalamus (boxed area) is shown to the right. Arrows indicate bipolar radial glial progenitors, arrowheads indicate the progeny, open arrowheads indicate the radial glial fibers, and the asterisk indicates the ventricular endfoot. Scale bars: 100 μm and 20 μm.
Supplementary Figure 2 Cre recombinase–dependent retroviral labeling of thalamic clones.
(a) Confocal image of a P21 Olig3-Cre;Ai9-tdTomato brain stained for tdTomato (red) and with DAPI (blue). Note the specific expression of tdTomato in the thalamus and thalamocortical projections. Scale bar: 500 μm. (b) Confocal image of an E15 Olig3-Cre brain injected with low titer Cre recombinase-dependent retrovirus expressing EGFP at E11 and stained for EGFP (green) and with DAPI (blue). Note two radial clonal clusters in the developing thalamus (broken line). High magnification image of a clone (boxed area) is shown at the bottom. The arrow indicates the bipolar radial glial progenitor and arrowheads indicate the progeny that are radially arrayed along the radial glial fiber (open arrowheads). Another labeled clonal cluster is located ventrally. Scale bars: 200 μm and 50 μm. TC, thalamocortical.
Supplementary Figure 3 Correlated localization of MADM-labeled RGPs at the embryonic stage and clones at the more mature stage.
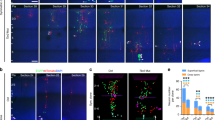
(a) Quantification of the average number of neurons in clones labeled at E10 and examined at E15 or P21 (E10-15, n=23; E10-P21, n=23). Data are presented as median with interquartile range and whiskers are the minimum and maximum. *, p=0.03 (Mann- Whitney test). (b) Confocal images of an E14 MADM-labeled thalamus treated with TM at E12 and stained for EGFP (green) and tdTomato (red), and with DAPI (blue). A two-cell local cluster without any radial glial progenitors (boxed area) is labeled in the developing thalamus. High magnification image is shown to the right. M, medial; L, lateral. Scale bars: 100 μm and 10 μm. (c) Quantification of the localizations of MADM-labeled RGPs at the embryonic stage and clones at the more mature stage. Note a clear correlation between the localizations of labeled progenitor cells and labeled clones at different embryonic stages. pTH-R: rostral progenitor domain of the thalamus; PTh: prethalamus. (d) NND analysis of clones labeled at different embryonic stages. Note that the cumulative frequency of NND of clones labeled at E9 (red, n=18), E10 (green, n=28), E11 (orange, n=19), and E12 (blue, n=11) is similar and significantly left-shifted than that of random simulated dataset (gray). Data are presented as mean±s.e.m. n.s., not significant; ****, p=1x10–15 (unpaired t test with Welch's correction). (e) Quantification of the fraction of clones containing neurons only (N) or neurons and glia (N+G) at different embryonic stages.
Supplementary Figure 4 Distinct morphological features of excitatory neurons, astrocytes, and oligodendrocytes in the thalamus.
(a, b) Representative 3D projection (a) and confocal (b) images of a G2-X clone (green and red) in the thalamus labeled at E9, and immunostained for neuronal marker NEUN (white) and counterstained with DAPI (blue) at P21. High magnification images of the somas of four cells are shown to the right in b. Broken line circles indicate the nuclei. Scale bars: 20 μm and 20 μm. (c, d) Representative 3D projection (c) and confocal (d) images of a G2-X thalamic clone (green and red) occupying TRN (contoured in yellow lines) and ZI labeled at E9, and immunostained for astrocyte marker S100 (white), and counterstained with DAPI (blue) at P21. High magnification images of the soma of the astrocyte are shown to the right in d. The broken line circle indicates the nucleus. Scale bars: 20 μm and 20 μm. (e, f) Representative 3D projection (e) and confocal (f) images of a G2-Z thalamic clone (yellow) occupying TRN (contoured in yellow lines) and ZI labeled at E9, and immunostained for oligodendrocyte marker OLIG2 (white), and counterstained with DAPI (blue) at P21. High magnification images of the somas of two oligodendrocytes (arrows) are shown to the right in e. Broken line circles indicate the nuclei. Note the labeled oligodendrocytes possess characteristic short parallel neurites. Scale bars: 20 μm and 20 μm. (g) Quantification of the fraction of labeled cells with a large soma/nucleus and long neurites or a small soma/nucleus and short bushy or parallel neurites that are positive for NEUN, S100, or OLIG2, respectively.
Supplementary Figure 5 Alignment of the experimental thalamus with the ABA reference thalamus.
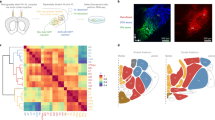
(a) Aligned single coronal sections from 34 experimental brains (gray outlines) in the position corresponding to ABA section 69. Red lines indicate 6 line profiles used to calculate thalamus edge (i.e. border) variability. D, dorsal; V, ventral. (b) Thalamus border variability for the section in a (n=45). (c) Average border variability for 5 sections along the anterior to posterior axis (n=45). (d) Aligned 3D reference ABA thalamus after normalization (red) with the average experimental thalamus (blue). (e) Percentage of volume overlap between reference ABA thalamus and individual experimental thalami (n=45). Data in b, c and r are presented as median with interquartile range, and whiskers are the minimum and maximum.
Supplementary Figure 6 Inhibitory interneuron clones in TRN.
(a) 3D reconstructed images of a G2-X clone containing 7 labeled neurons in the TRN contoured by yellow lines. (b) High magnification cross-section confocal images of the 7 labeled neurons stained for parvalbumin (PV, white), a GABAergic interneuron-specific marker. Note that all 7 neurons are positive for PV, indicating their interneuron identity. Scale bar: 10 μm. (c) Quantification of the average number of neurons in the four clonal clusters ('mvp', n=33; 'a', n=9; 'md', n=18; 'GABAergic', n=17). Data are presented as median with interquartile range, and whiskers are the minimum and maximum. **, p=0.005 (‘mvp’ versus ‘a’, MannWhitney test); **, p=0.001 (‘mvp’ versus ‘GABAergic’, Mann-Whitney test). (d) Quantification of the average number of nuclei that the four clonal clusters occupy. Data are presented as median with interquartile range, and whiskers are the minimum and maximum. n.s., not significant; ****, p<0.0001 (Mann Whitney test. Details are available in the Supplementary Methods Checklist).
Supplementary Figure 7 Validation of the number of excitatory neuron clonal subclusters based on nuclear occupation in the thalamus.
Quantification of the Silhouette coefficients with regard to the number of clusters. The red arrow indicates the highest Silhouette coefficient at three clusters.
Supplementary Figure 8 Overlap between the nuclei occupied by the ‘mvp’ clonal cluster and the nuclei harboring excitatory neuron progeny of OLIG2-positive progenitors in the developing thalamus.
(a) Representative 3D projection images of tdTomato-expressing excitatory neuron progeny (red filled circles) generated by OLIG2-positive progenitors in the thalamus. Note that the inhibitory interneurons located in the vLG and TRN, likely derived from OLIG2-positive progenitors in the prethalamus, were not traced. Scale bar: 500 μm. (b) Representative confocal images of tdTomato-expressing excitatory neuron (left), astrocyte (middle), and oligodendrocyte (right) progeny generated by OLIG2-positive thalamic progenitors. High magnification images of the somas (arrows) are shown at the bottom. Broken line circles indicate the nuclei. Scale bars: 10 μm and 10 μm. (c) Representative nuclear occupation of tdTomato-expressing excitatory neuron progeny (red filled circles) generated by OLIG2-positive progenitors.
Supplementary Figure 9 Distinct localization and functionality of labeled neurons in VM, RE, and RH between the ‘a’ and ‘mvp’ clonal clusters.
(a) Percentage of labeled neurons in the ‘a’ (left) and ‘mvp’ (right) clonal clusters that are located in the anterior or posterior half of VM, RE and RH. Note that labeled neurons in the ‘a’ clonal cluster are predominantly located in the anterior part, whereas those in the ‘mvp’ cluster are mostly located in the posterior part. (b) Distinct functionality of labeled neurons in the ‘a’ (top) and ‘mvp’ (bottom) clonal clusters located in VM, RE, and RH. Representative ‘a’ and ‘mvp’ cluster clones with corresponding ABA sections with nuclear boundaries are shown to the left. Asterisks indicate labeled neurons located in VM (black area), RE (dark gray area) and RH (light gray area). Gray lines indicate the contours of the thalamic sections where the labeled clones are located and black lines indicate the sections where labeled VM/RE/RH neurons are located. The corresponding cortical projection-based voxel maps of the same regions are shown to the right. Note that the anterior part of VM, RE, and RH predominantly contribute to higher order cognitive functions, whereas the posterior part of VM, RE, and RH mostly contribute to sensory/motor-related activities. TC, thalamocortical.
Supplementary Figure 10 Expression of SmoM2-EYFP in MADM-labeled thalamic clones.
Representative confocal images of a G2-X clone labeled at E10, and immunostained for EYFP (green) and tdTomato (red), and counterstained with DAPI (blue) at E14. High magnification images of a tdTomato-expressing RGP in the VZ (broken lines) are shown to the right. Note the membrane expression of SmoM2-EYFP in the tdTomato-expressing RGP (arrows) with an apical endfoot (asterisks), suggesting the successful recombination of both SmoM2-EYFP and MADM11 alleles in the clone. Arrowheads indicate the expression of SmoM2-EYFP in a nearby non-MADM-labeled cell, indicating a higher frequency of intra-chromosomal (i.e. SmoM2-EYFP allele) recombination than that of inter-chromosomal (i.e. MADM11 allele) recombination. Broken line circles indicate the somas. Scale bars: 20 μm and 10 μm.
Supplementary Figure 11 Shh signaling does not affect the spatial distribution of thalamic clones in the medial ventral posterior region.
(a) Representative 3D reconstructed images of the thalamic hemispheres containing a MADM-labeled clone in the medial ventral/posterior region of the wild type control (top) and SmoM2 (bottom) mice. Note that the clone in the SmoM2 mouse is similarly dispersed radially to that in the control mouse. Scale bar: 500 μm. (b) Quantification of the average number of neurons in green/red G2-X clones labeled at E10 (control, n=23; SmoM2, n=11), E11 (control, n=13; SmoM2, n=10), and E12 (control, n=9; SmoM2, n=3) between the control and SmoM2 mice. Data are presented as median with interquartile range, and whiskers are the minimum and maximum. n.s., not significant. (c) Quantification of the average number of neurons in clones located in the anterior (control, n=9; SmoM2, n=5), medial dorsal (control, n=18; SmoM2, n=6), and medial ventral/posterior (control, n=33; SmoM2, n=21) regions of the P4 control and SmoM2 mice. Data are presented as median with interquartile range, and whiskers are the minimum and maximum. n.s., not significant. (d) Representative in situ hybridization images of the anterior (top) and medial (bottom) section of the control (left) and SmoM2 (right) thalami for RORα. Note the expansion of RORα expression to the anterior region, as well as dorsally, in the SmoM2 thalamus compared to the control. Scale bar: 200 μm. (e) NND analysis of MADM-labeled neuronal clones in the SmoM2 thalamus. Note that compared to random datasets simulated 100 times (gray), the cumulative frequency of NND of thalamic neuronal clones (red, n=31) is significantly left-shifted towards shorter distances, indicating a spatial clustering. Data are presented as mean±s.e.m. ****, p p=10–15 (unpaired t test with Welch’s correction).
Supplementary Figure 12 Shh signaling affects gliogenesis in the thalamus.
(a) Quantification of the average number of glial cells in individual clones in the control (left) and SmoM2 (right) thalami (control, n=34; SmoM2, n=24). Data are presented as median with interquartile range, and whiskers are the minimum and maximum. n.s., not significant (Mann Whitney test). (b) Quantification of the number of clones containing neurons only (N) or neurons and glia (N+G) in the wild type control (left) and SmoM2 (right) thalami. Note the increase in the fraction of clones containing glial cells in the SmoM2 thalamus compared to the control. **, p=0.003 (χ2 test). (c) NND analysis of MADM-labeled glial clones in the SmoM2 thalamus. Note that compared to random datasets simulated 100 times (gray), the cumulative frequency of NND of thalamic glial clones (blue, n=14) is significantly left-shifted towards shorter distances, indicating a spatial clustering. Data are presented as mean±s.e.m. ****, p=1x10–15 (unpaired t test with Welch’s correction). (d) Schematic representation of lineage-related nuclear formation and functional organization in the thalamus.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 (PDF 2554 kb)
Supplementary Table 1
Spatial distribution (X/Y/Z coordinates) and nuclear occupation of labeled neurons and glia within individual RGP-derived clones. (XLSX 152 kb)
3D rendered image of a P21 thalamic hemisphere containing an ‘mvp’ cluster clone.
The thalamus is colored in light gray, the non-sensory/motor-related nuclei in light magenta, and the sensory/motor-related nuclei in cyan. Labeled clonally related neurons in the thalamus are represented by colored dots. A, anterior; D, dorsal; M, medial. Similar displays are used in Supplementary Movie 2. (MP4 2220 kb)
3D rendered image of a P21 thalamic hemisphere containing an ‘mvp’ cluster clone that predominantly occupies the FO sensory/motor-related nuclei.
The thalamus is colored in light gray, the HO nuclei in green, the FO nuclei in yellow, and the pRE/pRH/SMT nuclei in orange. Labeled clonally related neurons in the thalamus are represented by colored dots. Similar displays are used in Supplementary Movie 4. (MP4 4896 kb)
41593_2017_BFnn4519_MOESM41_ESM.mp4
3D rendered image of a P21 thalamic hemisphere containing an ‘md’ cluster clone that predominantly occupies the HO sensory/motor-related nuclei. (MP4 4585 kb)
Rights and permissions
About this article
Cite this article
Shi, W., Xianyu, A., Han, Z. et al. Ontogenetic establishment of order-specific nuclear organization in the mammalian thalamus. Nat Neurosci 20, 516–528 (2017). https://doi.org/10.1038/nn.4519
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4519
This article is cited by
-
Developmental timing-dependent organization of synaptic connections between mossy fibers and granule cells in the cerebellum
Communications Biology (2023)
-
Multicolor strategies for investigating clonal expansion and tissue plasticity
Cellular and Molecular Life Sciences (2022)
-
A repeated molecular architecture across thalamic pathways
Nature Neuroscience (2019)
-
Linking neuronal lineage and wiring specificity
Neural Development (2018)