Abstract
The high-throughput detection of organelle composition and proteomic mapping of protein environment directly from primary tissue as well as the identification of interactors of insoluble proteins that form higher-order structures have remained challenges in biological research. We report a proximity-based labeling approach that uses an antibody to a target antigen to guide biotin deposition onto adjacent proteins in fixed cells and primary tissues, which allows proteins in close proximity to the target antigen to be captured and identified by mass spectrometry. We demonstrated the specificity and sensitivity of our method by examining the well-studied mitochondrial matrix. We then used the method to profile the dynamic interactome of lamin A/C in multiple cell and tissue types under various treatment conditions. The ability to detect proximal proteins and putative interactors in intact tissues, and to quantify changes caused by different conditions or in the presence of disease mutations, can provide a window into cell biology and disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
27 July 2018
In the version of this article initially published, the introduction focused on methods for characterizing the nuclear envelope and did not include a comprehensive overview of proximity-based methods, some of which have similarly utilized antibody-conjugated peroxidase for proximity labeling by biotin deposition. The authors apologize for the omission of the following references: Kotani, N. et al., Proc. Natl. Acad. Sci. USA 105, 7405–7409 (2008); Hashimoto, N. et al., Proteomics 12, 3154–3163 (2012); Li, X. W. et al., J. Biol. Chem. 289, 14434–14447 (2014); and Rees, J. S. et al., Curr. Protoc. Protein Sci. 80, 19.27.1–19.27.18 (2015).
References
Roux, K.J., Kim, D.I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Kubben, N. et al. Identification of differential protein interactors of lamin A and progerin. Nucleus 1, 513–525 (2010).
Dittmer, T.A. et al. Systematic identification of pathological lamin A interactors. Mol. Biol. Cell 25, 1493–1510 (2014).
Depreux, F.F. et al. Disruption of the lamin A and matrin-3 interaction by myopathic LMNA mutations. Hum. Mol. Genet. 24, 4284–4295 (2015).
Fu, Y. et al. MacroH2A1 associates with nuclear lamina and maintains chromatin architecture in mouse liver cells. Sci. Rep. 5, 17186 (2015).
Engelke, R. et al. The quantitative nuclear matrix proteome as a biochemical snapshot of nuclear organization. J. Proteome Res. 13, 3940–3956 (2014).
Calvo, S.E., Clauser, K.R. & Mootha, V.K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016).
Pagliarini, D.J. et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 134, 112–123 (2008).
Smith, A.C. & Robinson, A.J. MitoMiner v3.1, an update on the mitochondrial proteomics database. Nucleic Acids Res. 44, D1258–D1261 (2016).
Rhee, H.W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Hung, V. et al. Proteomic mapping of the human mitochondrial intermembrane space in live cells via ratiometric APEX tagging. Mol. Cell 55, 332–341 (2014).
Forner, F., Foster, L.J., Campanaro, S., Valle, G. & Mann, M. Quantitative proteomic comparison of rat mitochondria from muscle, heart, and liver. Mol. Cell. Proteomics 5, 608–619 (2006).
Verstraeten, V.L. et al. Protein farnesylation inhibitors cause donut-shaped cell nuclei attributable to a centrosome separation defect. Proc. Natl. Acad. Sci. USA 108, 4997–5002 (2011).
Worman, H.J. & Schirmer, E.C. Nuclear membrane diversity: underlying tissue-specific pathologies in disease? Curr. Opin. Cell Biol. 34, 101–112 (2015).
Draviam, R.A., Shand, S.H. & Watkins, S.C. The beta-delta-core of sarcoglycan is essential for deposition at the plasma membrane. Muscle Nerve 34, 691–701 (2006).
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 28, 1248–1250 (2010).
Beck, B.D. & Dynlacht, J.R. Heat-induced aggregation of XRCC5 (Ku80) in nontolerant and thermotolerant cells. Radiat. Res. 156, 767–774 (2001).
Eriksson, M. et al. Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423, 293–298 (2003).
Liu, G.H. et al. Recapitulation of premature ageing with iPSCs from Hutchinson–Gilford progeria syndrome. Nature 472, 221–225 (2011).
de Haan, W. Lipodystrophy and muscular dystrophy caused by PTRF mutations. Clin. Genet. 77, 436–437 (2010).
Taranum, S. et al. LINC complex alterations in DMD and EDMD/CMT fibroblasts. Eur. J. Cell Biol. 91, 614–628 (2012).
Heydemann, A., Demonbreun, A., Hadhazy, M., Earley, J.U. & McNally, E.M. Nuclear sequestration of delta-sarcoglycan disrupts the nuclear localization of lamin A/C and emerin in cardiomyocytes. Hum. Mol. Genet. 16, 355–363 (2007).
Kandert, S. et al. Nesprin-2 giant safeguards nuclear envelope architecture in LMNA S143F progeria cells. Hum. Mol. Genet. 16, 2944–2959 (2007).
Lei, K. et al. Inner nuclear envelope proteins SUN1 and SUN2 play a prominent role in the DNA damage response. Curr. Biol. 22, 1609–1615 (2012).
Starr, D.A. Laminopathies: too much SUN is a bad thing. Curr. Biol. 22, R678–R680 (2012).
Datta, S., Snow, C.J. & Paschal, B.M. A pathway linking oxidative stress and the Ran GTPase system in progeria. Mol. Biol. Cell 25, 1202–1215 (2014).
Nikolova-Krstevski, V. et al. Nesprin-1 and actin contribute to nuclear and cytoskeletal defects in lamin A/C-deficient cardiomyopathy. J. Mol. Cell. Cardiol. 50, 479–486 (2011).
Swift, J. et al. Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341, 1240104 (2013).
Volonte, D. et al. Inhibition of nuclear factor-erythroid 2-related factor (Nrf2) by caveolin-1 promotes stress-induced premature senescence. Mol. Biol. Cell 24, 1852–1862 (2013).
Strambio-De-Castillia, C., Niepel, M. & Rout, M.P. The nuclear pore complex: bridging nuclear transport and gene regulation. Nat. Rev. Mol. Cell Biol. 11, 490–501 (2010).
Mi, H., Muruganujan, A., Casagrande, J.T. & Thomas, P.D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 8, 1551–1566 (2013).
Zwerger, M. et al. Altering lamina assembly reveals lamina-dependent and -independent functions for A-type lamins. J. Cell Sci. 128, 3607–3620 (2015).
Britton, S., Coates, J. & Jackson, S.P. A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J. Cell Biol. 202, 579–595 (2013).
Cao, K., Capell, B.C., Erdos, M.R., Djabali, K. & Collins, F.S. A lamin A protein isoform overexpressed in Hutchinson–Gilford progeria syndrome interferes with mitosis in progeria and normal cells. Proc. Natl. Acad. Sci. USA 104, 4949–4954 (2007).
Bar, D.Z. & Collins, F.S. Biotinylation by antibody recognition. Protocol Exchange http://dx.doi.org/10.1038/protex.2017.153 (2017).
Vizcaíno, J.A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We thank S. Patel and M. Gucek of the NHLBI Proteomics Core for operating the mass spectrometer and for technical assistance in sample preparation; S. Wincovitch for microscopy assistance; and J. Fekecs for preparing Figure 1a. We thank L. Bonnycastle and A. Dubose for critical reading of the manuscript, useful suggestions and comments.
Author information
Authors and Affiliations
Contributions
D.Z.B. and F.S.C. designed the experiments with input from Y.G. D.Z.B. and K.A. performed the experiments with assistance from M.R.E. U.T. performed the mouse work, and D.Z.B. analyzed the data. D.Z.B. and F.S.C. wrote the manuscript, and K.A and Y.G. provided comments and edits.
Corresponding author
Ethics declarations
Competing interests
The National Institutes of Health has filed for a patent application, number 62211160, covering some parts of the information contained in this article.
Integrated supplementary information
Supplementary Figure 1 BAR identifies known nuclear envelope proteins by Western blot and mass-spectrometry
A. Aligned signal distribution of lamin A/C (MAB3211; Cy3) and biotin (FITC-Avidin), derived from Fig 1B.
B. Western blot using Streptavidin-HRP showing antibody guided efficient biotin labeling and recovery of proteins. Beads – post-elution beads were boiled in Laemmli sample buffer to validate effective elution from the beads.
C. LAP2, a known lamin A/C binding protein, is pulled down by BAR, as evident from a Western blot.
D. A scaffold (Proteome Software, Portland, Oregon) screenshot showing multiple known NE and lamin A/C interacting proteins, and their peptide counts in the antibody sample and control.
Supplementary Figure 2 Effect of labeling time and antibody target on BAR
A. Protein level signal intensity (i.e. area) of 1 minute in-gel digestion vs 7 minutes on bead digestion. Proteins ordered (x-axis) in descending 7 minute intensity.
B. Heat-map showing the fraction of each dataset covered by any other dataset.
C. Heavy (lamin A/C MAB3211 antibody) to light (lamin A/C N-18 antibody) ratios of lamins, LAP2β and nuclear pore complex proteins.
D. Heavy (lamin A/C) to light (lamin B1) ratios of lamins, LAP2β and nuclear pore complex proteins.
Supplementary Figure 3 Additional protein staining in mouse and human tissues reproduce original findings
A. Primary mouse skeletal muscle stained for lamin A/C. BF - brightfield.
B. Primary human adipose tissue showing two doughnut shaped nuclei.
C. Primary human skeletal muscle tissue showing SGCA staining at the nuclear periphery.
Scale bar - 10 μm.
Supplementary Figure 4 SILAC controls for differential proteomics show minimal variations.
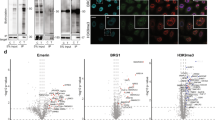
A. Scatter plot showing signal intensity of BAR extracted proteins from heavy vs. light labeled untreated HeLa cells.
B. Heavy/Light ratio of HeLa NE proteins in untreated cells. Proteins are ordered by peptide count. Proteins passing the filtering criteria in the LMNA-Unbound dataset were considered NE proteins.
Supplementary Figure 5 Ku70 and Ku80 interact with lamin A/C, as evident by FRET
Förster resonance energy transfer (FRET) in HeLa cells show that for Ku70 and Ku80, but not DNA-PKcs, fluorescence intensity increases after bleaching of lamin A/C adjacent fluorophores. FRET efficiency was 0 for DNA-PKcs, 0.34 for Ku70 and 0.15 for Ku80. We note that negative FRET results, as is the case for DNA-PKcs, cannot be interpreted as a lack of interaction.
Supplementary Figure 6 Heat shock affects the localization but not abundance of Ku70, Ku80 and HSPA8
A. Immunofluorescence of Ku80 and HSPA8 before and after heat shock shows nuclear envelope enrichment of Ku80 following heat shock, nuclear envelope localization of HSPA8 prior to heat shock and cytoplasmic depletion and nuclear envelope aggregation of HSPA8 following heat shock. Scale bar - 10 μm.
B. Western blot of lamin A/C, Ku70 and Ku80 showing no changes in amount of protein following heat shock. Cells were counted and equal amount of cells was loaded to gel.
Supplementary Figure 7 CAV1 localization in primary fibroblasts and skeletal muscle
A. HGPS fibroblasts stained for CAV1 and merged with brightfield.
B. Human skeletal muscle stained with CAV1 and DAPI.
Scale bar - 10 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 908 kb)
Supplementary Protocol
Supplementary Protocol (PDF 101 kb)
Supplementary Table 1
Antibodies, plasmids and cell lines used. (XLSX 48 kb)
Supplementary Data
Mass spectrometry data and analysis. (XLSX 3679 kb)
Rights and permissions
About this article
Cite this article
Bar, D., Atkatsh, K., Tavarez, U. et al. Biotinylation by antibody recognition—a method for proximity labeling. Nat Methods 15, 127–133 (2018). https://doi.org/10.1038/nmeth.4533
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4533