Abstract
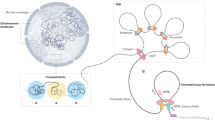
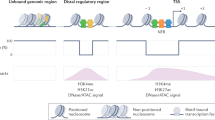
Development, differentiation and response to environmental stimuli are characterized by sequential changes in cellular state initiated by the de novo binding of regulated transcriptional factors to their cognate genomic sites1,2,3. The mechanism whereby a given regulatory factor selects a limited number of in vivo targets from a myriad of potential genomic binding sites is undetermined. Here we show that up to 95% of de novo genomic binding by the glucocorticoid receptor4, a paradigmatic ligand-activated transcription factor, is targeted to preexisting foci of accessible chromatin. Factor binding invariably potentiates chromatin accessibility. Cell-selective glucocorticoid receptor occupancy patterns appear to be comprehensively predetermined by cell-specific differences in baseline chromatin accessibility patterns, with secondary contributions from local sequence features. The results define a framework for understanding regulatory factor–genome interactions and provide a molecular basis for the tissue selectivity of steroid pharmaceuticals and other agents that intersect the living genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
Gene Expression Omnibus
Sequence Read Archive
References
Britten, R.J. & Davidson, E.H. Gene regulation for higher cells: a theory. Science 165, 349–357 (1969).
McKenna, N.J. & O'Malley, B.W. Combinatorial control of gene expression by nuclear receptors and coregulators. Cell 108, 465–474 (2002).
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007).
Evans, R.M. The steroid and thyroid hormone receptor superfamily. Science 240, 889–895 (1988).
Felsenfeld, G. & Groudine, M. Controlling the double helix. Nature 421, 448–453 (2003).
Wu, C. The 5′ ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature 286, 854–860 (1980).
Gross, D.S. & Garrard, W.T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988).
Htun, H., Barsony, J., Renyi, I., Gould, D.L. & Hager, G.L. Visualization of glucocorticoid receptor translocation and intranuclear organization in living cells with a green fluorescent protein chimera. Proc. Natl. Acad. Sci. USA 93, 4845–4850 (1996).
So, A.Y.-L., Chaivorapol, C., Bolton, E.C., Li, H. & Yamamoto, K.R. Determinants of cell- and gene-specific transcriptional regulation by the glucocorticoid receptor. PLoS Genet. 3, e94 (2007).
Reddy, T.E. et al. Genomic determination of the glucocorticoid response reveals unexpected mechanisms of gene regulation. Genome Res. 19, 2163–2171 (2009).
Richard-Foy, H. & Hager, G.L. Sequence-specific positioning of nucleosomes over the steroid-inducible MMTV promoter. EMBO J. 6, 2321–2328 (1987).
Becker, P., Renkawitz, R. & Schütz, G. Tissue-specific DNaseI hypersensitive sites in the 5′-flanking sequences of the tryptophan oxygenase and the tyrosine aminotransferase genes. EMBO J. 3, 2015–2020 (1984).
Hager, G.L. et al. Influence of chromatin structure on the binding of transcription factors to DNA. Cold Spring Harb. Symp. Quant. Biol. 58, 63–71 (1993).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Sekimata, M. et al. CCCTC-binding factor and the transcription factor T-bet orchestrate T helper 1 cell-specific structure and function at the interferon-gamma locus. Immunity 31, 551–564 (2009).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Johnson, D.S., Mortazavi, A., Myers, R.M. & Wold, B. Genome-wide mapping of in vivo protein-DNA interactions. Science 316, 1497–1502 (2007).
Stalder, J. et al. Tissue-specific DNA cleavages in the globin chromatin domain introduced by DNAase I. Cell 20, 451–460 (1980).
von der Ahe, D. et al. Glucocorticoid and progesterone receptors bind to the same sites in two hormonally regulated promoters. Nature 313, 706–709 (1985).
Diamond, M.I., Miner, J.N., Yoshinaga, S.K. & Yamamoto, K.R. Transcription factor interactions: selectors of positive or negative regulation from a single DNA element. Science 249, 1266–1272 (1990).
Bailey, T.L. & Gribskov, M. Concerning the accuracy of MAST E-values. Bioinformatics 16, 488–489 (2000).
Beck, I.M.E. et al. Crosstalk in inflammation: the interplay of glucocorticoid receptor-based mechanisms and kinases and phosphatases. Endocr. Rev. 30, 830–882 (2009).
Rigaud, G., Roux, J., Pictet, R. & Grange, T. In vivo footprinting of rat TAT gene: dynamic interplay between the glucocorticoid receptor and a liver-specific factor. Cell 67, 977–986 (1991).
Cordingley, M.G. & Hager, G.L. Binding of multiple factors to the MMTV promoter in crude and fractionated nuclear extracts. Nucleic Acids Res. 16, 609–628 (1988).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
John, S. et al. Kinetic complexity of the global response to glucocorticoid receptor action. Endocrinology 150, 1766–1774 (2009).
John, S. et al. Interaction of the glucocorticoid receptor with the chromatin landscape. Mol. Cell 29, 611–624 (2008).
Sekimata, M. et al. CCCTC-binding factor and the transcription factor T-bet orchestrate T helper 1 cell-specific structure and function at the interferon-gamma locus. Immunity 31, 551–564 (2009).
Sabo, P.J. et al. Genome-scale mapping of DNase I sensitivity in vivo using tiling DNA microarrays. Nat. Methods 3, 511–518 (2006).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Sabo, P.J. et al. Discovery of functional noncoding elements by digital analysis of chromatin structure. Proc. Natl. Acad. Sci. USA 101, 16837–16842 (2004).
Bailey, T.L., Williams, N., Misleh, C. & Li, W.W. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006).
Acknowledgements
We would like to thank T. Miranda, S. Morris, K. Nalley and L. Grontved for critical reading of the manuscript. We also thank M. Weaver, K. Lee, F. Neri, D. Bates and M. Diegel for technical assistance with the DNase I library preparation and sequencing. This research was supported in part by the Intramural Research Program of the US NIH, National Cancer Institute, Center for Cancer Research and funding from US NIH grant 1RC2HG005654 to J.A.S.
Author information
Authors and Affiliations
Contributions
S.J., P.J.S., G.L.H. and J.A.S. designed the experiments. S.J., P.J.S., S.C.B. and T.A.J. conducted the DNase-seq, ChIP-seq and expression array experiments. S.J., P.J.S., R.E.T., M.-H.S. and J.A.S. analyzed the data. S.J., P.J.S., R.E.T., M.-H.S., G.L.H. and J.A.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–10 (PDF 6921 kb)
Supplementary Table 1
DNaseI sensitive regions in the baseline (pre-hormone) state in the murine mammary adenocarcinoma cell line, 3134; DNase I sensitive regions in post dexamethasone-treated 3134 cells (XLS 3781 kb)
Supplementary Table 2
DNaseI hypersensitive sites (DHSs) in the baseline (pre-hormone) state in the murine mammary adenocarcinoma cell line, 3134 (XLS 3781 kb)
Supplementary Table 3
DNaseI hypersensitive sites (DHSs) in post dexamethasone-treated 3134 cells (XLS 3781 kb)
Supplementary Table 4
GR occupancy sites in the murine mammary adenocarcinoma cell line, 3134 (FDR 0%) (XLS 489 kb)
Supplementary Table 5
Expression analysis of mammary (3134) and pituitary (AtT-20) cells (XLS 132 kb)
Supplementary Table 6
GRBE sequence classes with greater than 50 instances in the genome. Chromatin Context Coefficient (CCC) classes in the murine genome (XLS 302 kb)
Supplementary Table 7
DNaseI sensitive regions in the baseline (pre-hormone) state in the murine pituitary cell line, AtT-20; DNase I sensitive regions in the post-hormone state in the murine pituitary cell line, AtT-20 (XLS 3781 kb)
Supplementary Table 8
DNaseI hypersensitive sites (DHSs) in the baseline (pre-hormone) state in the murine pituitary cell line, AtT-20 (XLS 3781 kb)
Supplementary Table 9
DNaseI hypersensitive sites (DHSs) post-hormone in AtT-20 cells (XLS 3781 kb)
Supplementary Table 10
GR occupancy sites in the murine pituitary cell line, AtT-20 (FDR 0%) (XLS 202 kb)
Rights and permissions
About this article
Cite this article
John, S., Sabo, P., Thurman, R. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat Genet 43, 264–268 (2011). https://doi.org/10.1038/ng.759
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.759
This article is cited by
-
Epigenetic profiling reveals key genes and cis-regulatory networks specific to human parathyroids
Nature Communications (2024)
-
Genome-wide chromatin accessibility landscape and dynamics of transcription factor networks during ovule and fiber development in cotton
BMC Biology (2023)
-
Remembering through the genome: the role of chromatin states in brain functions and diseases
Translational Psychiatry (2023)
-
H3K36 methylation maintains cell identity by regulating opposing lineage programmes
Nature Cell Biology (2023)
-
Thyroid hormone-regulated chromatin landscape and transcriptional sensitivity of the pituitary gland
Communications Biology (2023)