Abstract
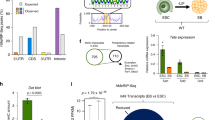
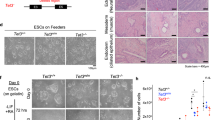
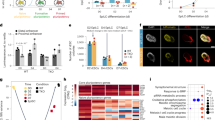
DNA methylation is one of the best-characterized epigenetic modifications1,2,3,4. Although the enzymes that catalyse DNA methylation have been characterized, enzymes responsible for demethylation have been elusive5. A recent study indicates that the human TET1 protein could catalyse the conversion of 5-methylcytosine (5mC) of DNA to 5-hydroxymethylcytosine (5hmC), raising the possibility that DNA demethylation may be a Tet1-mediated process6. Here we extend this study by demonstrating that all three mouse Tet proteins (Tet1, Tet2 and Tet3) can also catalyse a similar reaction. Tet1 has an important role in mouse embryonic stem (ES) cell maintenance through maintaining the expression of Nanog in ES cells. Downregulation of Nanog via Tet1 knockdown correlates with methylation of the Nanog promoter, supporting a role for Tet1 in regulating DNA methylation status. Furthermore, knockdown of Tet1 in pre-implantation embryos results in a bias towards trophectoderm differentiation. Thus, our studies not only uncover the enzymatic activity of the Tet proteins, but also demonstrate a role for Tet1 in ES cell maintenance and inner cell mass cell specification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
26 August 2010
A change was made to Acknowledgments
References
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002)
Cedar, H. & Bergman, Y. Linking DNA methylation and histone modification: patterns and paradigms. Nature Rev. Genet. 10, 295–304 (2009)
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nature Rev. Genet. 2008, 129–140 (2008)
Surani, M. A., Hayashi, K. & Hajkova, P. Genetic and epigenetic regulators of pluripotency. Cell 128, 747–762 (2007)
Ooi, S. K. & Bestor, T. H. The colorful history of active DNA demethylation. Cell 133, 1145–1148 (2008)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Tardy-Planechaud, S., Fujimoto, J., Lin, S. S. & Sowers, L. C. Solid phase synthesis and restriction endonuclease cleavage of oligodeoxynucleotides containing 5-(hydroxymethyl)-cytosine. Nucleic Acids Res. 25, 553–559 (1997)
Hattori, N. et al. Epigenetic regulation of Nanog gene in embryonic stem and trophoblast stem cells. Genes Cells 12, 387–396 (2007)
Tsumura, A. et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11, 805–814 (2006)
Jackson, M. et al. Severe global DNA hypomethylation blocks differentiation and induces histone hyperacetylation in embryonic stem cells. Mol. Cell. Biol. 24, 8862–8871 (2004)
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000)
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 10, 475–478 (2000)
Bruniquel, D. & Schwartz, R. H. Selective, stable demethylation of the interleukin-2 gene enhances transcription by an active process. Nature Immunol. 4, 235–240 (2003)
Popp, C. et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463, 1101–1105 (2010)
Okada, Y., Yamagata, K., Hong, K., Wakayama, T. & Zhang, Y. A role for the elongator complex in zygotic paternal genome demethylation. Nature 463, 554–558 (2010)
Cannon, S. V., Cummings, A. & Teebor, G. W. 5-Hydroxymethylcytosine DNA glycosylase activity in mammalian tissue. Biochem. Biophys. Res. Commun. 151, 1173–1179 (1988)
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007)
Ivanova, N. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006)
He, J., Kallin, E. M., Tsukada, Y. & Zhang, Y. The H3K36 demethylase Jhdm1b/Kdm2b regulates cell proliferation and senescence through p15Ink4b. Nature Struct. Mol. Biol. 15, 1169–1175 (2008)
Chen, Z. J. & Pikaard, C. S. Epigenetic silencing of RNA polymerase I transcription: a role for DNA methylation and histone modification in nucleolar dominance. Genes Dev. 11, 2124–2136 (1997)
Hasegawa, K., Cowan, A. B., Nakatsuji, N. & Suemori, H. Efficient multicistronic expression of a transgene in human embryonic stem cells. Stem Cells 25, 1707–1712 (2007)
Clark, S. J., Harrison, J., Paul, C. L. & Frommer, M. High sensitivity mapping of methylated cytosines. Nucleic Acids Res. 22, 2990–2997 (1994)
Acknowledgements
We thank M. Okano for the J1 and DNMT triple knockout ES cells; J. He and A. Nguyen for help in FACS sorting; and S. Wu for critical reading of the manuscript. This work was supported by NIH grants GM68804 (to Y.Z.) and CA084487 (to L.C.S.). S.I. is a research fellow of the Japan Society for the Promotion of Science. O.T. is a postdoctoral fellow of Juvenile Diabetes Research Foundation International. Y.Z. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.Z. conceived the project and wrote the manuscript. S.I., A.C.D., O.V.T. and K.H. designed and performed the experiments. L.C.S. provided the oligonucleotide substrates.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S13 with legends and Supplementary Tables 1-5. (PDF 12345 kb)
Rights and permissions
About this article
Cite this article
Ito, S., D’Alessio, A., Taranova, O. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010). https://doi.org/10.1038/nature09303
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09303
This article is cited by
-
Emerging role of RNA modification and long noncoding RNA interaction in cancer
Cancer Gene Therapy (2024)
-
Dynamic methylation pattern of H19DMR and KvDMR1 in bovine oocytes and preimplantation embryos
Journal of Assisted Reproduction and Genetics (2024)
-
Epigenetics and stroke: role of DNA methylation and effect of aging on blood–brain barrier recovery
Fluids and Barriers of the CNS (2023)
-
Epigenetic markers in the embryonal germ cell development and spermatogenesis
Basic and Clinical Andrology (2023)
-
Obesity and dyslipidemia are associated with partially reversible modifications to DNA hydroxymethylation of apoptosis- and senescence-related genes in swine adipose-derived mesenchymal stem/stromal cells
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.