Abstract
All commercial influenza vaccines elicit antibody responses that protect against seasonal infection, but this approach is limited by the need for annual vaccine reformulation that precludes efficient responses against epidemic and pandemic disease. In this study we describe a novel vaccination approach in which a nanoparticulate, liposome-based agent containing short, highly conserved influenza-derived peptides is delivered to the respiratory tract to elicit potent innate and selective T cell-based adaptive immune responses. Prepared without virus-specific peptides, mucosal immunostimulatory therapeutic (MIT) provided robust, but short-lived, protection against multiple, highly lethal strains of influenza in mice of diverse genetic backgrounds. MIT prepared with three highly conserved epitopes that elicited virus-specific memory T-cell responses but not neutralizing antibodies, termed MITpep, provided equivalent, but more durable, protection relative to MIT. Alveolar macrophages were more important than dendritic cells in determining the protective efficacy of MIT, which induced both canonical and non-canonical antiviral immune pathways. Through activation of airway mucosal innate and highly specific T-cell responses, MIT and MITpep represent novel approaches to antiviral protection that offer the possibility of universal protection against epidemic and pandemic influenza.
Similar content being viewed by others
Introduction
Viral infections of the lung are caused by some of the most important and lethal human pathogens, producing debilitating infections that can be lethal in the immunosuppressed and those at the extremes of age (http://www.cdc.gov/flu/about/disease/index.htm). Although many causes of respiratory infection represent regional health concerns, influenza is a global health threat producing symptomatic disease in up to 20% of children and 5% of adults annually.1 In addition to a continuous, low-level disease prevalence, the influenza virus induces seasonally enhanced disease incidence on local (epidemic) and global (pandemic) scales. Influenza infects and kills lung and airway cells, leading to potentially overwhelming inflammation accompanied by fever, cough, and difficulty in breathing. A majority of annual cases and pandemics are because of influenza A (70%) with less frequent cases caused by influenza B (30%).2
Although little has changed conceptually since their first scientific description >200 years ago, vaccines today represent a first line of defense against respiratory pathogens.3 Almost all influenza vaccines currently involve a single intramuscular injection of killed virus or a suspension of viral proteins containing aluminum-based adjuvants and are produced from infected chicken eggs.4 A second type of vaccine, Flumist, is applied to the nasal mucosa through aerosol delivery and elicits a limited infection.5 Both vaccines induce primarily antibody responses that inhibit viral infection of airway cells. Current influenza vaccines are extremely safe, except for excluded subject groups (egg-allergic subjects and, for Flumist, immunocompromised, pregnant, and elderly subjects) and are relatively effective.6, 7
Despite such benefits, annual influenza vaccination often fails to prevent serious infections in elderly and institutionalized patients.8 Furthermore, many find the requirement for intramuscular injection objectionable, suggesting an important barrier to more widespread and effective vaccination campaigns. An additional limitation of all current influenza vaccines is the requirement for annual reformulation because of mutations arising regularly within the influenza genome. The “just-in-time” strategy for influenza vaccine production is susceptible to a wide variety of misadventures ranging from selection of subdominant virus strains, as occurred in the 2007 and 2008 season,9 to difficulties in vaccine deployment that result in either ineffective or insufficient quantities of vaccine.10 Moreover, the slow pace of current vaccine manufacturing through embryonated chicken eggs precludes rapid, widespread vaccination against highly virulent strains with the potential to cause pandemic disease, as was seen for the 2009 and 2010 influenza season.11
In this study, we describe a novel vaccination platform designed to address the many limitations of current vaccine technology. We hypothesized that a vaccine that mimics closely the properties of a respiratory pathogen would stimulate optimal protective immunity in the lung, while simultaneously minimizing side effects. We focused on the following developmental criteria: nanoparticulate (10–1,000 nm in size); inclusion of adjuvants known to potently activate antimicrobial pathways;12 inclusion of short conserved influenza peptides sufficient to activate robust and potentially universal antipathogen memory T-cell responses;13 and formulation for delivery to the respiratory mucosa, i.e., by aerosol. We report herein the immunological properties and efficacy of our bimodal product developed against mouse-adapted strains of influenza.
Results
Designing a nanoparticulate airway mucosal vaccine
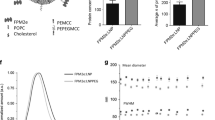
Our anti-influenza therapeutic was produced in two forms to elicit two modes of protection. Mucosal immunostimulatory therapeutic (MIT) consists of dilauroylphosphatidylcholine (DLPC) liposomes containing the adjuvants monophosphoryl lipid A (MPL) and trehalose 6,6′ dimycolate and, if prepared with short synthetic peptides derived from highly conserved regions of pathogen-derived proteins, is termed MITpep. Both preparations consist of nanoparticles (30–100 nm) in aqueous solution suitable for intranasal or aerosol delivery. A complete description of the development of MIT and MITpep is provided in Supplementary Appendix 1 online.
MIT nonspecifically confers protection against lethal influenza infection
To determine the protective efficacy of MIT in vivo, C57BL/6 mice were administered either MIT or DLPC vehicle 2 days before infection (day −2). On day 0, mice were exposed to aerosolized influenza A or influenza B (LD90) (Figure 1a). Despite the complete lack of viral immune epitopes, MIT conferred substantial protection against both influenza viruses, as assessed by survival, weight change, and symptom scores (Figure 1b–d). Similar protective efficacy was observed in influenza A-infected BALB/c mice (Figure 1e–g). Thus, MIT administered acutely to the respiratory tract of mice conferred protection against diverse influenza strains.
Mucosal immunostimulatory therapeutic (MIT) confers protection against lethal influenza infection. (a) Protocol for IN challenge 1 × with either MIT or dilauroylphosphatidylcholine (DLPC) vehicle followed by infection with an aerosolized LD90 dose of influenza A (H3N2) or B/Lee viruses. Changes in (b) survival, (c) body weight, and (d) symptom scores (see Methods) were recorded over 20 days in C57BL/6 mice (n=10 per group). (e–g) Identical experiments were performed in BALB/c mice using influenza A H3N2 (n=10 per group). IM, intramuscular; IN, intranasal. *P<0.05, MIT vs. vehicle; **IN MIT vs. IM H1N1 vaccine. Data are representative of three independent experiments.
Lung macrophages are a major cellular target of MIT
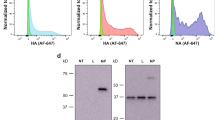
To gain insight into the critical lung cellular targets likely to confer MIT-dependent immunity against influenza, we determined changes in lung dendritic cell (DC) populations following intranasal administration of MIT. Although plasmacytoid DCs are powerful sources of type 1 interferons (IFNs) and are potent antiviral effector cells,14 proportional representation of lung plasmacytoid DCs decreased in response to MIT, whereas that of myeloid DCs increased (Figure 2a, b). These changes were specific to the lung because under the same conditions the relative abundance of myeloid and plasmacytoid DC populations in the spleen did not change (Supplementary Figure S1a online). Surprisingly, however, we failed to detect fluorescent lung DCs (CD11c+/mPDCA-1+ lung cells) after mice had been challenged with a fluorescent version of MIT, suggesting that lung DCs interacted with MIT only inefficiently (Figure 2c). Furthermore, lung DCs isolated from mice treated with MIT or vehicle when adoptively transferred to syngeneic mice failed to protect against influenza A infection (Figure 2d, e). These findings cannot alone rule out a significant contribution of lung DCs in this model, but suggest that other lung cells may be targeted by MIT and confer greater protection against influenza.
Lung dendritic cells (DCs) are not a major cellular target of mucosal immunostimulatory therapeutic (MIT). (a) Representative flow cytometry-derived two-color analyses of myeloid DCs (mDCs; CD11b+/CD11c+) and plasmacytoid DCs (pDCs; B220+/CD11b+) derived from lungs of mice challenge with either MIT or dilauroylphosphatidylcholine (DLPC) vehicle. Quadrant numbers indicate percentage of positive cells. (b) Cumulative flow cytometry data of the same mice showing DC data for lung and spleen (n=5 per group). (c) Representative mouse was challenged intranasally with fluorescein isothiocyanate (FITC)-labeled liposomes, after which total lung DCs (CD11c+/mPDCA-1+) were isolated and assessed for fluorochrome uptake. (d, e) Groups of mice received intratracheally (IT) lung-derived DC obtained from donor mice that were intranasally (IN) treated 2 days prior with either MIT or DLPC vehicle (d) and survival was determined (e) after challenge with an LD90 dose of influenza A (H3N2) (n=10 per group). *P<0.05. Data are representative of three independent experiments.
Therefore, we next focused on macrophages, the major phagocytic cell of the lung that is also capable of efficient T-cell activation.15 Fluorescence microscopy analysis confirmed that naive alveolar macrophages were able to internalize fluorescent liposomes in vitro (Figure 3a). To confirm that macrophages are functionally important cellular targets of MIT, we adoptively transferred MIT-treated bone marrow-derived macrophages to mice. In contrast to MIT-treated DCs, MIT-treated macrophages substantially protected mice against a lethal challenge with influenza A (Figure 3b). These studies indicate that lung macrophages are a major cellular target conferring protective immunity against influenza following activation by MIT.
Macrophages uptake mucosal immunostimulatory therapeutic (MIT) nanoparticles and confer protection against influenza. (a) Fluorescence microscopy image of alveolar macrophages after in vitro incubation with fluorescein isothiocyanate (FITC)-conjugated liposomes (n=5 per group). (b) Percent survival after influenza challenge (H3N2) of mice that received 1.5 × 106 MIT- or dilauroylphosphatidylcholine (DLPC) vehicle-stimulated bone marrow-derived macrophages intratracheally (n=15 per group). *P<0.05; **P<0.05 MIT vs. all other groups. Data are representative of three independent experiments.
MIT enhances lung secretion of antiviral cytokines and inhibits viral replication
To further elucidate the protective mechanisms of MIT in this model, we determined lung cytokine responses induced in response to MIT in influenza A-infected mice. C57BL/6 mice were treated intranasally with MIT or vehicle 2 days before influenza A infection, after which whole lungs were collected on days 0, 2, 4, 6, 8, and 30 after infection for quantification of cytokines and viral titers. Substantial increases in interleukin (IL)-10, IL-12, and tumor necrosis factor were detected after day 4 in both infected/untreated lungs and after treatment with DLPC vehicle before infection (Figure 4a, b). In contrast, MIT treatment elicited approximately 10- and 5-fold higher IL-17A and IL-12 responses on days 0 and 8 after infection, respectively (Figure 4c). However, whereas infection of naive and vehicle-challenged mice resulted in high-level secretion of IFN-γ especially at day 6 after infection, MIT-treated animals showed relatively increased IFN-γ responses at the earlier time points, including before infection (day 0) and relatively reduced responses at day 6 and thereafter (Figure 4d). Similar trends were observed with lung IL-6, IL-1α, and IL-1β levels (Supplementary Figures S2 and S3 online). These divergent cytokine responses corresponded to lung viral burdens that were much lower (up to 1,000-fold) in MIT-treated animals at all time points where virus was detectable (Figure 4e). Thus, MIT treatment induced the production of several cytokines with antiviral effects, including IL-12,16, 17 IL-6,18 and IL-1β.19
Mucosal immunostimulatory therapeutic (MIT) enhances lung IL-17 and IL-12 production and inhibits viral replication. At 2 days before infection with influenza, H3N2 mice were either untreated or treated with inhaled dilauroylphosphatidylcholine (DLPC) liposomes (vehicle) or MIT followed by infection. At the indicated days after infection, lung homogenates were prepared and the indicated (a–c) cytokines and (d) interferon-γ (IFN-γ) were measured. (e) Lung viral titers determined from the same samples at the indicated time points. *P<0.05 (n=3 mice/group/time point). Data are representative of three independent experiments.
MIT mediates protection against influenza through innate and adaptive immune cells and TRIF: TIR-domain-containing adapter-inducing interferon-b
We performed additional studies to determine if MIT protected mice through innate, adaptive, or both immune systems. Recombinase activating gene 1-deficient (Rag1−/−) mice that lack adaptive immune cells were challenged with MIT followed by influenza infection as in Figure 1a. Although most MIT-treated Rag1−/− mice ultimately succumbed to infection (Figure 5a), the time to death was significantly extended overall; one mouse survived to 30 days with no apparent residual disease (Figure 5a–c). Moreover, death in some MIT-treated Rag1−/− mice was not solely because of pneumonitis, as seen in wild-type mice (Supplementary Figure S4 and Supplementary Video 1 online), but also likely because of viral encephalitis, as assessed by pronounced ataxia manifested in moribund animals (3/10 MIT-treated mice) and a positive brain viral culture that was determined post mortem in one ataxic animal (Supplementary Figure S5 online). These studies demonstrate that adaptive immune cells, especially T cells, are necessary for full protection conferred by MIT, but that innate immune defenses are also requisite in this regard and, in some instances, may alone be sufficient for complete protection. Moreover, these findings emphasize that a principal function of adaptive immune cells is to confine replicating virus to the lung and prevent dissemination to the central nervous system.20
Mucosal immunostimulatory therapeutic (MIT) mediates protection against influenza through TRIF and innate immune cells. Rag1−/− mice were challenged with MIT or dilauroylphosphatidylcholine (DLPC) vehicle 2 days before infection with influenza H3N2, and (a) survival, (b) body weight, and (c) symptom scores were monitored (n=10 per group). The same end points were determined in identically treated wild-type (WT), MyD88−/−, and TRIF−/− mice (d–f) and interferon (IFN)α/βR−/− mice (g–i) (n=10 per group). *P<0.05 MIT vs. vehicle-challenged mice of identical genotype. Data are representative of two independent experiments.
Toll-like receptor (TLR) ligands such as lipopolysaccharide are known to activate more than one pathogen-associated molecular pattern receptor; thus, we considered the possibility that MPL and trehalose 6,6′ dimycolate activate both TLR-dependent and TLR-independent pathways.21 To determine this, the protective efficacy of MIT was assessed in mice with disruptions of major TLR signaling pathways (MyD88−/− and TRIF−/− mice; Figure 5d–f). Unexpectedly, MIT essentially fully protected MyD88−/− mice; moreover, weight loss and symptom score data revealed that a functional MyD88 gene was associated with enhanced weight loss and more severe symptoms (Figure 5e, f). In contrast, MIT-treated TRIF−/− mice showed marked susceptibility to infection, with only 1 of 10 animals surviving long term (Figure 5d–f). These findings are in agreement with the previous observation that MPL is a TLR4 agonist that signals predominantly through TRIF.22
Enhanced production of antiviral cytokines correlated with inhibition of viral replication at early time points, potentially accounting for the beneficial effect of MIT in vivo (Figure 4e). Although IL-1α and β were most strongly induced by MIT (Supplementary Figure S3a–c online), they could not account for the antiviral effect of MIT because these cytokines signal through MyD88.23 To confirm an important role for type I IFNs, we challenged mice deficient in the IFN α/β receptor with MIT and influenza (Figure 5g–i). As expected, MIT-treated IFNα/βR−/− mice that are unable to respond to all type 1 IFNs were more susceptible to influenza relative to control animals, but again we observed a substantial long-term survival (3/10 animals; Figure 5g–i). Together, these data demonstrate that innate and adaptive immune cells interact cooperatively to mediate MIT-based protection. Moreover, TRIF and type I IFNs are critical components of the protective efficacy of MIT, but other independent signaling pathways contribute significantly to MIT-based protection.
MITpep induces broad influenza protection and long-term memory T-cell responses
Although MIT conferred broad-spectrum protection against influenza, the duration of protective efficacy was limited to 5 days after a single challenge (data not shown). Moreover, MIT contains no peptide epitopes and is therefore not a pathogen-specific vaccine. To develop an influenza-specific mucosal vaccine based on MIT, short, highly conserved myristylated peptides derived from matrix 2 (M2)(2–24), hemagglutinin (HA)(307−329), and nucleoprotein (NP)(366–374) proteins of influenza A/PR8 (H1N1) were incorporated into MIT, thus creating MITpep (Supplementary Appendix, Table A1 online). Initial experiments with MITpep revealed that 100% protection against influenza required a minimum of three intranasal challenges with infection occurring 7 days after the final vaccination (data not shown). We then compared the protective efficacy of MIT against MITpep using this protocol (Figure 6a). These studies confirmed the much greater efficacy of MITpep relative to MIT (Figure 6b–d; 10/10 vs. 2/10 survivors; P<0.05).
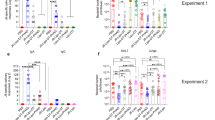
MITpep elicits durable protection against diverse influenza strains. Mice were challenged intranasally with vehicle, mucosal immunostimulatory therapeutic (MIT) or MIT to which was added synthetic peptides representing highly conserved regions of three influenza proteins (MITpep) according to the indicated schedule before infection with influenza (a). (b) Survival, (c) body weight, and (d) symptom scores were then monitored over the next 20 days (n=10 per group). Mice were immunized with MITpep and challenged with influenza strains A/H1N1, A/H3N2, and B/Lee as in Figure 5. (e) Survival, (f) body weight, and (g) symptom scores were similarly monitored (n=10 per group). (h) Lung histopathology 30 days following influenza infection of naive mice compared with dilauroylphosphatidylcholine (DLPC) vehicle and MITpep-challenged animals. Periodic acid-Schiff (original magnification × 100). a, arteriole; b, bronchiole; I, inflammatory cells; M, mucus impaction. (i) H2Db NP366–374-reactive CD8 T cells as determined by flow cytometry and (j) nucleoprotein (NP)-reactive interferon-γ (IFN-γ)-secreting cells as detected in lungs by enzyme-linked immunosorbent spot (ELISpot) following MITpep vaccination (n=5 per group). (k) Mice were challenged with influenza H3N2 at 2, 4, 6, and 8 weeks following the final vaccination with MITpep and survival was monitored over 20 days (n=10 per group). (l) Mice vaccinated with MITpep received a booster immunization 83 days after the final vaccination series and were infected with influenza 7 days later. Survival was monitored over 20 days (n=10 per group). Solid arrow indicates IN vaccination or booster, and dotted arrow, indicates influenza infection. (m) Nucleoprotein (NP)-reactive CD8+ T cells in lungs after booster immunization (n=5 per group). *P<0.05 MITpep vs. MIT or vehicle; **P<0.05, MITpep vs. vehicle for isogenic virus; ***P<0.05; ****P<0.05 vs. 8 weeks post IN. Data are representative of two independent experiments.
Relative to MIT, the increased time required to achieve maximal MITpep protection suggested that adaptive immune B and/or T cells were required. However, analysis of serum of vaccinated mice revealed no virus-neutralizing antibodies. Moreover, although a weak serum anti-M2 peptide IgG1 response was observed, no IgG2a or airway IgA responses to any peptides were detected (Supplementary Figure S6 online). Together with the observation that MIT failed to substantially protect Rag1−/− mice (Figure 5a–c), these findings indicate that the protective efficacy of both MIT and MITpep is mediated largely through T cells.
MITpep elicited complete protection against three diverse influenza strains in C57BL/6 mice (H-2b) (Figure 6e–g) and substantially protected two additional major histocompatibility complex class I-divergent strains of mice (CBA/J (H-2k) and BALB/c (H-2d)) from lethal influenza A/HK (H3N2) infection (Supplementary Figure S7a–c online). Comparison of naive mouse lungs with lungs from mice exposed to DLPC vehicle or MITpep and collected 30 days after influenza infection revealed unexpectedly extensive residual alveolar and interstitial inflammation in vehicle-treated animals. Bronchial dilatation, goblet cell metaplasia, and mucus impaction were observed widely in infected, vehicle-challenged animals, but were only rarely seen in infected, MITpep-treated animals, with the majority of lungs from these mice appearing similar to those from naive animals (Figure 6h).
The NP(366–374) peptide contained in MITpep has previously been shown to elicit robust CD8 T-cell responses,24 and indeed by tetramer staining, ∼17% of lung T cells consisted of NP+ CD8+ T cells after three immunizations, whereas no tetramer-positive cells were detected from spleens (Figure 6i and Supplementary Figure S8 online). MITpep further elicited exclusively IFN-γ and no IL-4 secretion from lung homogenate cells reactive against NP (Figure 6j and data not shown), HA, and M2 peptides (data not shown and Supplementary Figure S9 online), and again such cells could not be found in spleens or mediastinal lymph nodes (Figure 6j). These data indicate that MITpep elicits multiple peptide-specific T-cell responses that at least acutely are confined to the lung.
Finally, we explored the duration of MITpep efficacy. Whereas mice challenged three times with MIT exhibited poor (2/10) survival when infected 7 days after the final exposure (Figure 6b), 9/10 of mice immunized identically with MITpep survived when infected 42 days after the final vaccination (Figure 6k). However, efficacy was entirely lost by 60 days after completing the vaccine series (Figure 6k). This finding indicates either a loss of previously extant lung resident antiviral effector/memory cells or the failure entirely to generate effector/memory T-cell responses. To distinguish these possibilities, mice received a booster immunization with MITpep 90 days after the initial vaccination series and were infected with influenza 7 days later. A single MITpep boost improved survival from 0 to 40% (Figure 6l). Moreover, vaccine boosting restored NP tetramer-positive CD8 T cells to lung in a manner that was not seen with a single MITpep challenge of naive animals (Figure 6m and data not shown). Thus, MITpep induced broad-spectrum protection against influenza and complete long-term protection that was durable for a minimum of 6 weeks. Moreover, protective efficacy could be readily boosted concomitant with the rapid restoration of NP-specific CD8 T cells to the lung.
Discussion
The major limitation of current influenza vaccination practice is the requirement for annual reformulation because of antigenic shift and drift that are characteristic of this and other viruses.25 The complexity of current vaccine manufacturing procedures further impairs the ability of public health systems to respond efficiently to the added threats of epidemic and especially pandemic influenza.11 We have developed and tested a new approach to influenza vaccination that obviates in part these concerns. Rather than deriving immune epitopes from whole virus, we chose a largely synthetic approach involving short, highly conserved peptides. This vaccine design represents a marked departure from the current strategy of inducing predominantly antibody responses to prevent infection to one that emphasizes highly specific T-cell responses that sharply limit viral replication following infection, thereby limiting disease expression and the potential for lethal dissemination.
Our resulting nanoparticulate products, MIT and MITpep, are liposomal and delivered directly to the respiratory tract by intranasal application, or potentially by aerosol. Based on the criteria of survival, weight change, and symptom score, MIT elicited profound protection against influenza despite lacking canonical epitopes (i.e., full-length HA and NA proteins). MIT preferentially targeted lung macrophages, induced secretion of antiviral cytokines, and provided immediate protection against multiple influenza A and B strains in mice of diverse genetic backgrounds. MITpep further induced expansion and recruitment to lung of peptide-specific T cells in the absence of neutralizing viral antibodies and provided similar, but much greater, duration of protection relative to MIT. Together, these findings indicate that MITpep induces synergistic innate immune and adaptive T-cell memory responses through activation of macrophages with the potential to provide universal protection against influenza. We believe that this new approach to vaccination is applicable to influenza and other respiratory infections affecting diverse populations.
Although MIT and MITpep were designed with the objective of mimicking an infectious agent to most efficiently induce virus-neutralizing immunity, several lines of evidence indicate that MIT and MITpep induce atypical and possibly unique anti-influenza immune pathways. First, macrophages, and not DCs, were found to be essential cellular targets of MIT. This was unanticipated given the importance of DCs to influenza immunity as previously shown in the naive host.26, 27, 28, 29, 30, 31 Protective roles for macrophages in influenza have been identified, especially in previous studies of liposomal vaccines.32, 33 However, through TRAIL, macrophages mediate airway epithelial cell death, lung fluid exudation, and mortality in influenza, especially if unconstrained by CD200.34, 35 Thus, MIT and MITpep appear to induce protective immunity against influenza in part by inducing protective, and bypassing potentially lethal, macrophage activation pathways as seen with infection of naive animals.
Furthermore, although recognition of influenza requires TLR7 signaling through MyD88,36 adaptive immune responses against native influenza infection were found to be variably dependent on these molecules, with antibody responses requiring both TLR7 and MyD88, whereas T-cell responses required no TLR signaling.37 Our findings with MIT emphasize that immunity against influenza is inherently flexible and that MIT specifically harnesses protective immune pathways through TRIF and most likely TLR4, while avoiding potentially injurious pathways through MyD88 that are activated by virus in unvaccinated hosts.
Consistent with previous observations,38 our findings indicate that mucosal application of the vaccine to the target organ (airway) is superior to systemic administration. This is demonstrated by the rapid viral clearance elicited by topically applied MIT (Figure 4e) and the inability of MIT or MITpep to elicit any degree of protection if given intraperitoneally or subcutaneously (Supplementary Figure S10 online). Local induction by MIT of antiviral cytokines (Figure 4) is one such mechanism of local antiviral effect. Further study is required to demonstrate if MIT is capable of inducing complementary or alternate antimicrobial pathways involving IFITM (IFN-induced transmembrane) proteins,39 Nod 1 and 2 (nucleotide-binding oligomerization domain containing 1 and 2),40 IL-17 A,41 and retinoic acid inducible-gene I (RIG-I).42
MIT was alone capable of inducing protective immunity in the absence of virus-specific epitopes because of the MPL and trehalose 6,6′ dimycolate components. This nonspecific protective response is consistent with previous studies demonstrating the ability of compounds, including cholera toxin, polyprenols, flagellin, and bacterial lysates, to protect against unrelated infectious agents by activating airway innate immune defenses.43, 44, 45, 46 Thus, although the typical lung immune response to innocuous antigens is tolerogenic,47 our studies emphasize that the lungs perceive potentially dangerous infectious agents and rapidly activate effective countermeasures through diverse innate molecular pathways.
Many previous attempts to use M2,48, 49 HA,50 and NP proteins51 to achieve broad-spectrum protection against influenza have been made. However, these studies have relied on the generation of “universal” antibodies that cross-react against diverse influenza strains—an approach that has yet to be proven practical in diverse populations.52 Limited success using immunodominant cytotoxic T-cell epitopes derived from NP has also been demonstrated using liposomal influenza vaccines applied intranasally, but the requirement of anti-CD40 antibodies as adjuvants limits the practicality of this approach.53 The greatest conceptual advance revealed through our studies is that T cells are alone sufficient to provide complete protection against diverse influenza strains, and potentially extending to avian (H5N1) and swine (H1N1) influenza. The major potential drawback to this approach is that the duration of protective immunity was relatively short lived, despite generating durable (∼8 weeks) memory T-cell responses (Figure 6l, m). We hypothesize that susceptibility to infection re-emerges following vaccination because of the in situ loss of memory-effector T cells that are required at the onset of infection.54 Such cells are clearly recalled quickly to the lung following infection of vaccinated mice in which immunity has waned (Figure 6m), but the rapid pace of infection because of these mouse-adapted influenza strains appears to overwhelm even a brisk recall of memory T-cell response.
It should be noted, however, that the experimental infections used here are far more lethal than conventional or even pandemic influenza in humans and in this regard are not representative of naturally occurring disease. Consequently, the actual duration of efficacy of our vaccination if applied to humans cannot be determined from these studies alone. We predict that following full vaccination with MITpep in humans, annual or semiannual influenza infections will continue to occur, but are likely to be either asymptomatic or low grade. Such minor infections in a T-cell-vaccinated host represent an ideal response to influenza or, indeed, any infectious agent as they will activate the same protective T-cell subsets, yet elicit distinct antibody profiles with minimal host morbidity. The resulting combined immune response should reinforce protective T-cell memory responses, and the antibodies and T cells generated will likely prevent or reduce the severity of disease because of the same and heterosubtypic influenza strains.55, 56 The overall level of herd immunity should therefore improve with widespread vaccination, thereby reducing the severity of annual epidemics and reducing, if not eliminating, the possibility of pandemic disease.
In summary, we have shown that airway vaccination results in a highly effective and localized immune response that affords immediate and durable protection against a lethal viral pathogen through both innate- and T cell-based immune responses. By emphasizing T cell-dependent and not antibody-dependent immunity, the described method uniquely offers a practical means of achieving multistrain, if not universal influenza protection. The largely synthetic “plug-and-play” platform design further allows for the rapid development and testing of vaccines against virtually any infectious organism for which the sequence of target proteins is known.
Methods
Mice.
C57BL/6, BALB/c, and CBA/J females, 4–6 weeks old, were purchased from Jackson Laboratories (Bar Harbor, ME). Rag1−/− breeder mice were also purchased from Jackson Laboratories. MyD88−/− and TRIF−/− mice were the kind gifts of S. Akira (Osaka University, Japan). IFN α/β R−/− mice were originally generated by Dr Michel Aguet.57 All mice were bred in Baylor College of Medicine's transgenic animal facility, accredited by the Association for Assessment and Accreditation of Laboratory Animal Care. Animals were housed under pathogen-free conditions and mouse experiments were conducted in accordance with all institutional and federal guidelines.
Peptides and TLR ligands.
Conserved influenza A/PR8 (H1N1) peptides were synthesized with a myristic acid tag in the N-terminus by the Baylor College of Medicine Protein Chemistry Core. The peptide sequences are: M2(2–24): Mys-SLLTEVETPIRNEWGCRCNDSSD, HAFP(307−329): Mys-NIPSIQSRGLFGAIAGFIE, and NP(366–374): Mys-ASNENMETM. The TLR ligands MPL, and trehalose 6,6′ dimycolate were purchased from Invivogen (San Diego, CA) and Sigma Chemical (St Louis, MO), respectively.
Liposome preparation.
DLPC (Avanti Polar Lipid, Alabaster, AL) was initially dissolved and mixed in pre-warmed (50 °C) tert-butanol and dimethylsulfoxide to generate a homogeneous mixture of lipids and organic solvents. TLR ligands and peptides were then added to the mixture in a 1:5 molar ratio of epitopes to lipids. The lipid solution was then frozen at 0 °C for 24 h. The lipid cake was placed on a vacuum pump and lyophilized until dry for 24 h. Samples were rehydrated in phosphate-buffered saline, vortexed for 1 min, sonicated in a water bath for 1 min, and vortexed again for 1 min before administration. Fluorescent lipid (1-myristoyl-2-{12-[(7-nitro-2-1,3-benzoxadiazol-4-yl)amino]lauroyl}-sn-glycero-3-[phospho-rac-(1-glycerol)] (ammonium salt)) was also purchased from Avanti Polar Lipid.
Immunization and infection challenge.
MIT (50 μl) or DLPC vehicle were administered intranasally to animals once on day −2 before influenza infection (day 0). MITpep (50 μl) or DLPC vehicle were administered intranasally on days −21, −14, and −7. On day 0, mice were infected with either aerosolized LD90 of influenza A (A/PR/8/34 (H1N1) or A/HK/8/68 (H3N2) or influenza B/Lee/40. Animals were placed in an enclosed chamber with oxygen supplied through a tube connected to an air pump and virus was aerosolized for 20 min at 10 l min–1. Survival, weight change, and symptom scores (Supplementary Table S1 online) were assessed and monitored for 20 days post infection.
Adoptive transfer of lung DCs.
Mice were challenged with MIT or DLPC vehicle on day −2, and on day 0, lung DCs were enriched from lungs using pan-DC (CD11c and anti-mPDCA-1) paramagnetic microbeads (Miltenyi, Auburn, CA) and injected intratracheally (1.5 × 106 cells) into recipient mice.58 At 24 h after adoptive transfer, recipients were challenged intranasally with a LD90 of influenza A/HK (H3N2). Data were collected 24 h later.
Isolation of alveolar macrophages.
Lungs from naive C57BL/6 mice were harvested and single-cell suspension was made by manually pushing the organ through a 40 μm mesh. Alveolar macrophages were enriched using CD11b paramagnetic microbeads (Miltenyi). Sample purity was analyzed via flow cytometry using allophycocyanin-Alexa Fluor 750 F4/80 (eBioscience, San Diego, CA) and Phycoerythrin-CD11b (BD Biosciences, San Jose, CA) antibodies to identify macrophage populations. CD11b+/F4/80+ macrophages comprised 90% of the enriched cell population.
Adoptive transfer of bone marrow-derived macrophages.
Because insufficient alveolar macrophages could be obtained for in vivo adoptive transfer experiments, bone marrow-derived macrophages were used. Bone marrow cells were harvested from C57BL/6 mice and cultured in the presence of L929 cell supernatant (source of macrophage colony-stimulating factor), fetal bovine serum (Hyclone, Waltham, MA), and Dulbecco's modified Eagle's medium (Life Technologies, Carlsbad, CA) for 5 days for differentiation. On day 6, macrophages were harvested and incubated with either MIT or DLPC vehicle for 2 h before intratracheal injection (1.5 × 106 cells). For intratracheal injection, mice were anesthetized with etomidate (14 mg kg–1), and orally intubated with a sterile 1.25″ 20-gauge angiocatheter (Protect IV, Medic, Carlsbad, CA) using a tuberculin syringe. At 24 h after adoptive transfer, mice were challenged intranasally with LD90 of influenza A/HK (H3N2). Data were collected 24 h later.
Flow cytometry.
Single-cell suspensions were incubated with the following fluorochrome-conjugated antibodies: allophycocyanin-B220 (BD Biosciences); allophycocyanin-Alexa Fluor 750 F4/80 (eBioscience); Pacific Blue-CD11c (eBioscience); phycoerythrin-CD11b (BD Biosciences); and fluorescein isothiocyanate-CD8a (BD Biosciences). Major histocompatibility complex class I tetramer phycoerythrin-H2Db NP366–374 was synthesized by the Protein Chemistry Core MHC Tetramer Laboratory at Baylor.
Determination of lung viral titers.
On days 0, 2, 4, 6, 8, and 30 after immunization with MIT or DLPC vehicle, mouse lungs were collected and rinsed in sterile water to lyse excess red blood cells. Lungs were resuspended in Dulbecco's modified Eagle's medium and homogenized using a glass bead beater (Biospec Products, Bartlesville, OK). Samples were diluted in Dulbecco's modified Eagles’ medium containing 0.05% trypsin (Worthington Biochemical, Lakewood, NJ), centrifuged for 5 min, and supernatants were serially diluted in 96-well round-bottom plates (Fisher Scientific, Atlanta, GA). Samples were then transferred to 96-well round-bottom plates containing MDCK (Madin Darby canine kidney) cell monolayers. Lung dilutions and MDCK cells were allowed to incubate for 5 days, and then visualized for characteristic adherence of turkey red blood cells (Fitzgerald Industries, Concord, MA).
Quantification of cytokines from lung homogenates.
Cytokines (IL-1 α, IL-1 β, IL-4, IL-6, IL-10, IL-12, IL-13, IL-17A, IFN-γ, and tumor necrosis factor) were quantitated from samples collected for lung viral titers by luminex-based multiplex assay (Milliplex; Millipore, Billerica, MA) using a Bioplex analyzer (Bio-Rad, Hercules, CA).
Quantitation of cytokine-producing cells.
Total lung IFN-γ and IL-4-secreting cells were quantitated on day 0 after 3 × intranasal immunization with MITpep or vehicle on days −21, −14, and −7 by enzyme-linked immunosorbent spot assay as previously described.59 Total nucleoprotein peptide-reactive cells from lungs were determined by enzyme-linked immunosorbent spot assay after stimulation of whole lung leukocytes with 100 μl of nucleoprotein peptide (4.5 μg μl–1) for 48 h.
Statistics.
Data are presented as means±s.d. and are representative of at least three independent in vivo experiments, with 5 to >10 mice per group as indicated. Significant differences (P⩽0.05) were determined using the Mann–Whitney nonparametric t-test or log-rank test to determine statistical differences in survival studies.
References
Lewis, D.B. Avian flu to human influenza. Annu. Rev. Med. 57, 139–154 (2006).
Knipe, D.M. & Howley, P.M. Fundamental Virology (Lippincott Williams & Wilkins, 2001).
Rimmelzwaan, G.F. & Osterhaust, A.D. Influenza vaccines: new developments. Curr. Opin. Pharmacol. 1, 491–496 (2001).
Stephenson, I. & Nicholson, K.G. Influenza: vaccination and treatment. Eur. Respir. J. 17, 1282–1293 (2001).
Rhorer, J. et al. Efficacy of live attenuated influenza vaccine in children: a meta-analysis of nine randomized clinical trials. Vaccine 27, 1101–1110 (2009).
Govaert, T.M. et al. The efficacy of influenza vaccination in elderly individuals. A randomized double-blind placebo-controlled trial. JAMA 272, 1661–1665 (1994).
Wang, C.S., Wang, S.T., Lai, C.T., Lin, L.J. & Chou, P. Impact of influenza vaccination on major cause-specific mortality. Vaccine 25, 1196–1203 (2007).
Thompson, W.W. et al. Mortality associated with influenza and respiratory syncytial virus in the United States. JAMA 289, 179–186 (2003).
Centers for Disease Control and Prevention (CDC). Interim within-season estimate of the effectiveness of trivalent inactivated influenza vaccine—Marshfield, Wisconsin, 2007–08 influenza season. MMWR Morb. Mortal. Wkly. Rep. 57, 393–398 (2008).
Mossad, S.B. Coping with the influenza vaccine shortage. Cleve. Clin. J. Med. 71, 918, 920, 923, 927 (2004).
Itoh, Y. et al. In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature 460, 1021–1025 (2009).
Evans, J.T. et al. Enhancement of antigen-specific immunity via the TLR4 ligands MPL adjuvant and Ribi.529. Expert Rev. Vaccines 2, 219–229 (2003).
Taylor, P.M. & Askonas, B.A. Influenza nucleoprotein-specific cytotoxic T-cell clones are protective in vivo. Immunology 58, 417–420 (1986).
Liu, Y.J. IPC: professional type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annu. Rev. Immunol. 23, 275–306 (2005).
Peschke, T., Bender, A., Nain, M. & Gemsa, D. Role of macrophage cytokines in influenza A virus infections. Immunobiology 189, 340–355 (1993).
Mbawuike, I.N. et al. Human interleukin-12 enhances interferon-gamma-producing influenza-specific memory CD8+ cytotoxic T lymphocytes. J. Infect. Dis. 180, 1477–1486 (1999).
Monteiro, J.M., Harvey, C. & Trinchieri, G. Role of interleukin-12 in primary influenza virus infection. J. Virol. 72, 4825–4831 (1998).
Lee, S.W., Youn, J.W., Seong, B.L. & Sung, Y.C. IL-6 induces long-term protective immunity against a lethal challenge of influenza virus. Vaccine 17, 490–496 (1999).
Kozak, W. et al. Thermal and behavioral effects of lipopolysaccharide and influenza in interleukin-1 beta-deficient mice. Am. J. Physiol. 269, R969–R977 (1995).
Bergmann, C.C., Lane, T.E. & Stohlman, S.A. Coronavirus infection of the central nervous system: host-virus stand-off. Nat. Rev. Microbiol. 4, 121–132 (2006).
Shen, H., Tesar, B.M., Walker, W.E. & Goldstein, D.R. Dual signaling of MyD88 and TRIF is critical for maximal TLR4-induced dendritic cell maturation. J. Immunol. 181, 1849–1858 (2008).
Mata-Haro, V. et al. The vaccine adjuvant monophosphoryl lipid A as a TRIF-biased agonist of TLR4. Science 316, 1628–1632 (2007).
Burns, K. et al. MyD88, an adapter protein involved in interleukin-1 signaling. J. Biol. Chem. 273, 12203–12209 (1998).
Palmowski, M.J. et al. A single-chain H-2Db molecule presenting an influenza virus nucleoprotein epitope shows enhanced ability at stimulating CD8+ T cell responses in vivo. J. Immunol. 182, 4565–4571 (2009).
Kaiser, J. A one-size-fits-all flu vaccine? Science 312, 380–382 (2006).
GeurtsvanKessel, C.H. et al. Clearance of influenza virus from the lung depends on migratory langerin+CD11b- but not plasmacytoid dendritic cells. J. Exp. Med. 205, 1621–1634 (2008).
Krug, A. et al. CpG-A oligonucleotides induce a monocyte-derived dendritic cell-like phenotype that preferentially activates CD8 T cells. J. Immunol. 170, 3468–3477 (2003).
Jego, G. et al. Plasmacytoid dendritic cells induce plasma cell differentiation through type I interferon and interleukin 6. Immunity 19, 225–234 (2003).
Diebold, S.S. et al. Viral infection switches non-plasmacytoid dendritic cells into high interferon producers. Nature 424, 324–328 (2003).
Cella, M., Facchetti, F., Lanzavecchia, A. & Colonna, M. Plasmacytoid dendritic cells activated by influenza virus and CD40L drive a potent TH1 polarization. Nat. Immunol. 1, 305–310 (2000).
McGill, J., Van Rooijen, N. & Legge, K.L. IL-15 trans-presentation by pulmonary dendritic cells promotes effector CD8 T cell survival during influenza virus infection. J. Exp. Med. 207, 521–534 (2010).
Wijburg, O.L. et al. The role of macrophages in the induction and regulation of immunity elicited by exogenous antigens. Eur. J. Immunol. 28, 479–487 (1998).
Reading, P.C., Miller, J.L. & Anders, E.M. Involvement of the mannose receptor in infection of macrophages by influenza virus. J. Virol. 74, 5190–5197 (2000).
Snelgrove, R.J. et al. A critical function for CD200 in lung immune homeostasis and the severity of influenza infection. Nat. Immunol. 9, 1074–1083 (2008).
Herold, S. et al. Lung epithelial apoptosis in influenza virus pneumonia: the role of macrophage-expressed TNF-related apoptosis-inducing ligand. J. Exp. Med. 205, 3065–3077 (2008).
Lund, J.M. et al. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl. Acad. Sci. USA 101, 5598–5603 (2004).
Heer, A.K. et al. TLR signaling fine-tunes anti-influenza B cell responses without regulating effector T cell responses. J. Immunol. 178, 2182–2191 (2007).
Babai, I., Samira, S., Barenholz, Y., Zakay-Rones, Z. & Kedar, E. A novel influenza subunit vaccine composed of liposome-encapsulated haemagglutinin/neuraminidase and IL-2 or GM-CSF. II. Induction of TH1 and TH2 responses in mice. Vaccine 17, 1239–1250 (1999).
Brass, A.L. et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 139, 1243–1254 (2009).
Kim, Y.G. et al. The cytosolic sensors Nod1 and Nod2 are critical for bacterial recognition and host defense after exposure to Toll-like receptor ligands. Immunity 28, 246–257 (2008).
Liang, S.C. et al. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 203, 2271–2279 (2006).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Iida, J. et al. Prophylactic activity of dihydroheptaprenol, a synthetic polyprenol derivative, against Sendai virus infection in mice. Vaccine 8, 376–380 (1990).
Matsuo, K. et al. Induction of innate immunity by nasal influenza vaccine administered in combination with an adjuvant (cholera toxin). Vaccine 18, 2713–2722 (2000).
Vijay-Kumar, M. et al. Flagellin treatment protects against chemicals, bacteria, viruses, and radiation. J. Immunol. 180, 8280–8285 (2008).
Clement, C.G. et al. Stimulation of lung innate immunity protects against lethal pneumococcal pneumonia in mice. Am. J. Respir. Crit. Care Med. 177, 1322–1330 (2008).
Lloyd, C.M. & Hawrylowicz, C.M. Regulatory T cells in asthma. Immunity 31, 438–449 (2009).
De Filette, M. et al. Universal influenza A M2e-HBc vaccine protects against disease even in the presence of pre-existing anti-HBc antibodies. Vaccine 26, 6503–6507 (2008).
Ernst, W.A. et al. Protection against H1, H5, H6 and H9 influenza A infection with liposomal matrix 2 epitope vaccines. Vaccine 24, 5158–5168 (2006).
Bianchi, E. et al. Universal influenza B vaccine based on the maturational cleavage site of the hemagglutinin precursor. J. Virol. 79, 7380–7388 (2005).
Epstein, S.L. et al. Protection against multiple influenza A subtypes by vaccination with highly conserved nucleoprotein. Vaccine 23, 5404–5410 (2005).
Carrat, F. & Flahault, A. Influenza vaccine: the challenge of antigenic drift. Vaccine 25, 6852–6862 (2007).
Ninomiya, A., Ogasawara, K., Kajino, K., Takada, A. & Kida, H. Intranasal administration of a synthetic peptide vaccine encapsulated in liposome together with an anti-CD40 antibody induces protective immunity against influenza A virus in mice. Vaccine 20, 3123–3129 (2002).
Roman, E. et al. CD4 effector T cell subsets in the response to influenza: heterogeneity, migration, and function. J. Exp. Med. 196, 957–968 (2002).
Bodewes, R., Kreijtz, J.H. & Rimmelzwaan, G.F. Yearly influenza vaccinations: a double-edged sword? Lancet Infect. Dis. 9, 784–788 (2009).
Lee, L.Y. et al. Memory T cells established by seasonal human influenza A infection cross-react with avian influenza A (H5N1) in healthy individuals. J. Clin. Invest. 118, 3478–3490 (2008).
Muller, U. et al. Functional role of type I and type II interferons in antiviral defense. Science 264, 1918–1921 (1994).
De Heer, H.J. et al. Essential role of lung plasmacytoid dendritic cells in preventing asthmatic reactions to harmless inhaled antigen. J. Exp. Med. 200, 89–98 (2004).
Kheradmand, F. et al. A protease-activated pathway underlying Th cell type 2 activation and allergic lung disease. J. Immunol. 169, 5904–5911 (2002).
Acknowledgements
We thank members of the laboratories of D.B.C. and F.K. for helpful discussions and assistance. This work was supported by the United States National Institutes of Health grants AI057696 and HL69585 (to D.B.C.), HL095382 and AI070973 (to F.K. and D.B.C.), and AI007495 (to W.T.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Tai, W., Roberts, L., Seryshev, A. et al. Multistrain influenza protection induced by a nanoparticulate mucosal immunotherapeutic. Mucosal Immunol 4, 197–207 (2011). https://doi.org/10.1038/mi.2010.50
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2010.50
This article is cited by
-
Lipid-Based Delivery Systems in Development of Genetic and Subunit Vaccines
Molecular Biotechnology (2023)
-
Protective role of γδ T cells in cigarette smoke and influenza infection
Mucosal Immunology (2018)
-
Adsorption of a porcine reproductive and respiratory syndrome virus DNA vaccine candidate onto biodegradable nanoparticles improves immunogenicity in mice
Archives of Virology (2015)
-
Essential role for autophagy in the maintenance of immunological memory against influenza infection
Nature Medicine (2014)
-
A model for homeopathic remedy effects: low dose nanoparticles, allostatic cross-adaptation, and time-dependent sensitization in a complex adaptive system
BMC Complementary and Alternative Medicine (2012)