Abstract
The bacterium Wolbachia (order Rickettsiales), representing perhaps the most abundant vertically transmitted microbe worldwide, infects arthropods and filarial nematodes. In arthropods, Wolbachia can induce reproductive alterations and interfere with the transmission of several arthropod-borne pathogens. In addition, Wolbachia is an obligate mutualist of the filarial parasites that cause lymphatic filariasis and onchocerciasis in the tropics. Targeting Wolbachia with tetracycline antibiotics leads to sterilisation and ultimately death of adult filariae. However, several weeks of treatment are required, restricting the implementation of this control strategy. To date, the response of Wolbachia to stress has not been investigated, and almost nothing is known about global regulation of gene expression in this organism. We exposed an arthropod Wolbachia strain to doxycycline in vitro, and analysed differential expression by directional RNA-seq and label-free, quantitative proteomics. We found that Wolbachia responded not only by modulating expression of the translation machinery, but also by upregulating nucleotide synthesis and energy metabolism, while downregulating outer membrane proteins. Moreover, Wolbachia increased the expression of a key component of the twin-arginine translocase (tatA) and a phosphate ABC transporter ATPase (PstB); the latter is associated with decreased susceptibility to antimicrobials in free-living bacteria. Finally, the downregulation of 6S RNA during translational inhibition suggests that this small RNA is involved in growth rate control. Despite its highly reduced genome, Wolbachia shows a surprising ability to regulate gene expression during exposure to a potent stressor. Our findings have general relevance for the chemotherapy of obligate intracellular bacteria and the mechanistic basis of persistence in the Rickettsiales.
Similar content being viewed by others
Introduction
Wolbachia is an obligate intracellular α-proteobacterium (family Anaplasmataceae, order Rickettsiales) that infects an estimated 40% of terrestrial arthropods, suggesting that it is the most prevalent vertically transmitted symbiont worldwide (Zug and Hammerstein, 2012). In contrast with its pandemic distribution in the Arthropoda, Wolbachia has a limited range of infection in the Nematoda, being restricted to ∼40% of species in the superfamily Filarioidea (arthropod-transmitted parasites of vertebrates; the filariae) (Ferri et al., 2011) and a single genus of phytoparasitic Pratylenchidae (Haegeman et al., 2009). The nature of the symbiosis between Wolbachia and its host varies considerably both within and between arthropods and nematodes, although some general trends are apparent. First, arthropod Wolbachia tend to have relatively large genomes (1.3–1.7 Mb; Wu et al., 2004; Geniez et al., 2012) that frequently contain prophage regions (Kent and Bordenstein, 2010), insertion sequences (Cerveau et al., 2011) and large expansions of protein-coding genes containing ankyrin-like domains (ANKs) (Siozios et al., 2013), whereas their counterparts in the filariae have smaller genomes (0.9–1.1 Mb), in which these features are absent or greatly reduced (Foster et al., 2005; Godel et al., 2012). Second, in many instances of filaria–Wolbachia symbiosis, there is considerable evidence for host–symbiont coevolution over millions of years (Bandi et al., 1998), coupled with an obligate dependency of the nematode on Wolbachia for normal embryogenesis and viability (Hoerauf et al., 1999; Langworthy et al., 2000). In contrast, essential mutualisms between arthropods and Wolbachia appear to be rare (Dedeine et al., 2001; Hosokawa et al., 2010; Miller et al., 2010). Third, Wolbachia can induce a range of reproductive manipulations in arthropods that facilitate vertical transmission (Werren et al., 2008), but to date reproductive phenotypes have not been reported in nematode hosts.
Over the past two decades, Wolbachia has aroused intense interest not only because of its fascinating and complex biology but also because of its potential to contribute to disease control. Certain arthropod strains of Wolbachia have been demonstrated to reduce vector competence by truncating host life span (McMeniman et al., 2009) and/or by inhibiting the development or proliferation of various pathogens (Kambris et al., 2009; Moreira et al., 2009). In addition, the obligate dependency of important filarial parasites such as Wuchereria bancrofti (a cause of lymphatic filariasis) and Onchocerca volvulus (the aetiological agent of onchocerciasis) on their Wolbachia symbionts renders them susceptible to chemotherapy with tetracycline and rifamycin antibiotics (Hoerauf et al., 2000; Taylor et al., 2005; Specht et al., 2008), which deplete the bacteria from nematode tissues. Unfortunately, this promising approach to the control of two major neglected tropical diseases has been hampered by the treatment regimen required to irreversibly suppress Wolbachia in filariae. This comprises 4–6 weeks of daily therapy, which is considered too prolonged by the World Health Organisation for a mass drug administration programme akin to those currently based on conventional anthelminthics (Hoerauf et al., 2011). Nevertheless, antibiotics such as doxycycline remain the only safe drugs that exhibit potent activity against the long-lived adult filariae.
Here, we apply RNA-seq and label-free quantitative proteomics to dissect the phenotypic response of Wolbachia to short-term doxycycline exposure. As filarial Wolbachia cannot be cultured in vitro, we have utilised a mosquito cell line containing a Wolbachia strain (wMelPop-CLA) originally derived from Drosophila melanogaster (McMeniman et al., 2008), as the drug susceptibility of arthropod- and nematode-derived Wolbachia appears to be equivalent (Schiefer et al., 2012, 2013). Our data represent the first global gene expression study of an arthropod Wolbachia strain and reveal an unexpected capacity to regulate metabolic pathways and small RNAs during exposure to a potent stressor. Moreover, we identify shifts in the expression of outer membrane proteins and inner membrane transporters that may contribute to antibiotic tolerance in Wolbachia, including a phosphate ABC transporter ATPase associated with reduced susceptibility to antimicrobials in free-living bacteria.
Materials and methods
Cell culture, drug treatment and enrichment of Wolbachia
The Aedes albopictus cell line RML-12, stably transfected with Wolbachia strain wMelPop-CLA (McMeniman et al., 2008), was kindly provided by Scott O’Neill (Monash University, Victoria, Australia). The cells were maintained as previously described for the Wolbachia-infected mosquito cell line Aa23 (Makepeace et al., 2006). For antibiotic treatment, doxycycline hyclate (Fluka, Buchs, Switzerland) was added 4 days after subculture at a final concentration of 0.25 μg ml−1. Treated and control cells were harvested 3 days later. Wolbachia were enriched from the host cells using glass bead lysis and filtration, as previously described (Rasgon et al., 2006). The pellet containing the bacteria was stored in RNAlater (Sigma-Aldrich, Gillingham, UK) at 4 °C.
RNA extraction and RNA-seq
Total RNA was extracted from the pellet using TRI Reagent (Sigma-Aldrich) according to the manufacturer’s instructions, and quantified by RiboGreen fluorimetry (Invitrogen, Paisley, UK) on an Infinite F200 PRO multimode reader (Tecan, Männedorf, Switzerland). To deplete processed transcripts containing a 5′-monophosphate, a fraction of each RNA sample (2.5 μg) was incubated with 1 U of Terminator 5′-phosphate-dependent exonuclease (Epicentre, Madison, WI, USA) in Terminator reaction buffer B according to the manufacturer’s instructions. The RNA was purified by phenol–chloroform extraction, quantified as before and stored at −80 °C.
Contaminating genomic DNA was digested using the TURBO DNA-free Kit (Ambion, Austin, TX, USA) according to the manufacturer’s instructions, and sequencing libraries were prepared using the directional ScriptSeq v2 RNA-seq Library Preparation Kit (Epicentre) as directed by the manufacturer. Twelve libraries, comprising matched pre- and post-exonuclease-treated aliquots from three control and three doxycycline-exposed samples, were multiplexed and sequenced as 100-bp paired-ends on a HiSeq 2000 platform (Illumina, San Diego, CA, USA). Confirmatory runs were also performed using a similar protocol on a MiSeq personal sequencer (Illumina). The resulting reads were mapped to the wMel genome (NCBI RefSeq: NC_002978.6) in Burrows-Wheeler Aligner (Li and Durbin, 2009), and counts per gene were calculated by htseq-count. Differential expression (DE) analysis was performed in edgeR (Bioconductor), using a binomial distribution model (Robinson and Oshlack, 2010; Robinson et al., 2010). Genes were considered to exhibit DE where the fold change (FC) was ⩾2 and the P-value was <0.01. Data were deposited at the NCBI Sequence Read Archive under project ID SRA091852.
Protein extraction and nanoflow liquid chromatography electrospray ionisation tandem mass spectrometry
Before protein extraction, we further purified the Wolbachia-enriched material using additional filtration and a Percoll (Sigma-Aldrich) gradient, as used for sequencing of the wBol1 genome (Duplouy et al., 2013). Wolbachia pellets were washed twice in Hank’s balanced salt solution and stored at −80 °C. To solubilise protein, the pellets were sonicated in 25 mM ammonium bicarbonate (Sigma-Aldrich) and 0.1% (w/v) Rapigest (Waters, Elstree, UK) on ice. Proteomic-grade trypsin (Sigma-Aldrich) was added at a protein:trypsin ratio of 50:1, and samples were incubated at 37 °C overnight before removal of Rapigest by trifluoroacetic acid precipitation.
Peptide mixtures (2 μl) were analysed by online nanoflow liquid chromatography using the nanoACQUITY-nLC system (Waters) coupled to an LTQ-Orbitrap Velos (ThermoFisher Scientific, Bremen, Germany) mass spectrometer (MS) as previously described (Darby et al., 2012). Thermo RAW files were imported into Progenesis LC–MS (version 4.1, Nonlinear Dynamics, Newcastle, UK), and runs were time-aligned using default settings. Peptide intensities were normalised (using wMel features only) against an auto-selected reference run, and differences in protein expression and associated analysis of variance (ANOVA) P-values between seven control and nine doxycycline-treated biological replicates were calculated by Progenesis LC-MS. Spectral data were transformed to MGF files with Progenesis LC–MS, exported using the Mascot search engine (version 2.3.02, Matrix Science, London, UK) and searched against all protein sequences expected to be present in the sample (see Supplementary Materials and Methods). Search parameters were as reported previously (Darby et al., 2012), and the results from Mascot were further processed using the machine-learning algorithm Percolator. The false discovery rate was <1%, and individual ion scores >13 were considered to indicate identity or extensive homology (P <0.05). For a protein to be classified as undergoing DE, an FC of ⩾1.5 and a P-value of <0.01 were required, supported by ⩾2 unique peptides. Deeper proteome coverage was obtained by fractionation of a single bulk sample using strong anion exchange (Wiśniewski et al., 2009), and the resultant peptide data were used for linear regression analysis against the transcriptomic data set (see Supplementary Materials and Methods). The mass spectrometric data were deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository (Vizcaino et al., 2013) with the data set identifier PXD000345 and DOI 10.6019/PXD000345.
Results and Discussion
Features of the wMelPop-CLA transcriptome and proteome
We applied 5′-phosphate-dependent exonuclease treatment to total RNA extracted from Wolbachia-enriched cell culture material to enhance representation of unprocessed messenger RNA (mRNA) transcripts containing a 5′ triphosphate (Supplementary Figure S1). To maximise our ability to detect gene products from the wMel reference genome, we combined RNA-seq reads with quantitative peptide data obtained from a purified, fractionated isolate of wMelPop-CLA. We found transcript evidence (⩾2 reads in any one of three untreated biological replicates) and/or peptide evidence (⩾2 unique peptides from a single-bulk sample) for 798 (66.8%) of 1,195 protein-coding genes predicted from the wMel genome (Wu et al., 2004; Figure 1a). Notably, both the RNA-seq and proteomic data corresponded in detecting zones of limited or absent gene expression within regions of the genome identified as prophages (Wu et al., 2004) or a chromosomal inversion containing several transposase genes (Riegler et al., 2005; Figure 2, Supplementary Table S1). However, we did detect a small number of highly restricted peaks of gene expression in both the sense and antisense orientation within the prophages (Supplementary Figures S2 and S3, Supplementary Table S2).
Correlations between protein and mRNA expression. (a) Venn diagram of the number of proteins and transcripts quantified using quantitative proteomics and RNA-Seq, respectively. (b) Scatterplot of the relationship between genes quantified in both data sets. (c) Scatterplots for protein and transcript gene expression classified by six key Clusters of Orthologous Genes (COG) categories. In (b) and (c), mean non-normalised RNA reads (n=3) were log10-transformed and non-normalised protein abundances (n=1) were subjected to a log10(10−3) transformation. Scatterplots display the rectilinear equation and coefficient of determination (R2).
Differential expression of transcripts and peptides across the Wolbachia strain wMel chromosome. The wMel genome was used as a reference for strain wMelPop-CLA. Circles in both panels are numbered sequentially from the perimeter to the centre. Circles 1 and 2 represent protein-coding genes (blue) on the positive and negative strands, respectively, and the third circle shows structural RNA genes (rRNA (purple), sRNA (green) and tRNA (pink)), prophage regions (yellow) and the site of a chromosomal inversion (red). (a) Circles 4–9 are heat maps of transcript counts from control (4–6) and doxycycline-treated replicates (7–9), with high expression coloured red and low expression coloured blue; circle 10 is a plot of differential expression (fold-change) between control (outer profile) and treatment (inner profile). Note that expression from tRNA and rRNA genes was forced to baseline. (b) Circles 4–19 are heat maps of protein abundance (summed peptide ion intensity scores) from control (4–10) and doxycycline-treated replicates (11–19), with high abundance coloured red and low abundance coloured blue; circle 20 is a plot of differential abundance (fold-change) between control (outer profile) and treatment (inner profile).
The degree of global correlation between raw transcript reads and raw protein abundance scores was remarkably low (Figure 1b), as has been noted in other studies on bacteria (Waldbauer et al., 2012). However, within certain Clusters of Orthologous Groups (COGs), the degree of correlation was much higher, with R2 exceeding 0.8 for ‘cell wall, membrane, envelope biogenesis’ (Figure 1c). When expression was visualised across all COGs, the largest fold-differences between normalised means for RNA and protein data were observed for the categories ‘general function prediction only’, ‘coenzyme transport and metabolism’ and ‘carbohydrate transport and metabolism’ (where RNA dominated relative to protein in all cases) (Figure 3). Closer inspection of the data revealed that two proteins were vastly overrepresented by ion intensity score (Supplementary Table S3), molecular chaperone GroEL and Wolbachia surface protein, which are also highly abundant in the proteomes of the filarial Wolbachia strains wBm (Bennuru et al., 2011) and wOo (Darby et al., 2012). In addition, transcripts encoding these two proteins were among the most dominant in the wMelPop-CLA transcriptome (Supplementary Table S4).
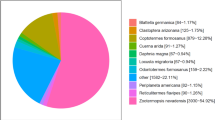
Mean abundance of transcripts and peptides across COG groups for Wolbachia strain wMel. Pie charts representing (a) the mean number of unadjusted transcript reads and (b) the mean protein abundance (summed peptide ion intensity score) for untreated Wolbachia cells, normalised to the number of genes per Cluster of Orthologous Groups (COG). The COGs are labelled clockwise from 12 o’clock and n per COG is shown in parentheses.
Identification of differentially expressed genes after doxycycline treatment
Wolbachia strain wMelPop-CLA was grown in mosquito cells with and without doxycycline to identify changes in RNA expression and protein abundance. For the analysis of transcriptional DE, 549 genes passed our filter (see Supplementary Materials and Methods), of which 36 were classified as upregulated and 32 as downregulated following doxycycline treatment (Supplementary Table S5, Figure 2a). To determine parallel changes in the proteome, 434 wMel proteins were identified by the presence of ⩾2 unique peptides. Of these, 10 proteins were found to be upregulated and 28 were downregulated (Supplementary Table S6, Figure 2b). The wMel genome contains an abundance of hypothetical proteins, and, accordingly, 37% of genes exhibiting DE fell into this category (Supplementary Tables S7 and S8). Putative functions could be assigned to several of these features, including an upregulated relE-like addiction module toxin (WD0124) and a downregulated protein with similarity to stringent starvation protein B (WD0128). These may have roles in mRNA degradation (Maisonneuve et al., 2011) and the regulation of proteolysis following ribosome stalling (Lessner et al., 2007), respectively. Importantly, despite this high proportion of uncharacterised genes among the regulated data set, 81.6% of all genes showing DE at the RNA and/or protein level had orthologues in at least one filarial Wolbachia genome, or represented conserved non-coding RNAs (Supplementary Tables S5–S8). In contrast, 61.6% of 258 orthologues found only in arthropod Wolbachia strains displayed insufficient expression to be included in the analysis of DE for both RNA and protein (data not shown).
Effects on nucleotide synthesis, energy metabolism and membrane composition
Analysis of DE identified changes in genes involved in a variety of cellular processes (Figure 4), although three broad categories contained the majority (∼60%) of the regulated genes with an annotation: translation and ribosome assembly; nucleotide, cofactor and energy metabolism; and DNA replication and transcription. Tetracyclines bind with high affinity to prokaryotic ribosomes, preventing docking of aminoacyl-tRNA and inhibiting translation (Griffin et al., 2010). Thus, the upregulation of ribosomal protein genes that we observed (Supplementary Figure S4) was fully anticipated, and has been reported previously from studies on free-living bacteria exposed to translational inhibitors (VanBogelen and Neidhardt, 1990; Evers et al., 2001; Ng et al., 2003). However, we also noted a surprisingly broad impact of doxycycline on interconnected metabolic pathways involved in energy generation and de novo nucleotide synthesis (Figure 5). The Wolbachia genome encodes several glycolytic enzymes, and three of these were upregulated after doxycycline treatment (Figure 5). As Wolbachia lack enzymes that catalyse the irreversible reactions of glycolysis, those remaining in the genome function predominantly to supply precursors for nucleotide and phospholipid synthesis via the gluconeogenesis pathway (Foster et al., 2005). Accordingly, three enzymes that participate in de novo nucleotide generation were also upregulated at the mRNA level, as was an enzyme involved in phospholipid synthesis (Figure 5). Thus, upregulation of glycolytic enzymes could reflect a regulatory mechanism to increase the size of the substrate pool for these pathways. Moreover, the upregulation of enzymes involved in phospholipid synthesis is compatible with a possible role for membrane remodelling in reducing doxycycline uptake. In support of this hypothesis, we observed concurrent downregulation of two Wolbachia surface protein paralogues and a further five predicted membrane proteins (Supplementary Tables S5–S7). A significant reduction in outer membrane protein abundance has also been demonstrated both in tetracycline-resistant strains of free-living bacteria, such as E. coli (Lin et al., 2010) and Acinetobacter baumannii (Yun et al., 2008), and in doxycycline-insensitive isolates of Orientia tsutsugamushi (Chao et al., 2009), another member of the Rickettsiales.
Annotated wMel genes differentially expressed following doxycycline treatment. Bar chart of fold changes for individual genes (proteins, red; RNA, blue; locus tags, right) exhibiting statistically significant (P<0.01) differential expression after doxycycline treatment, grouped into broad functional categories. Within each group, genes are ranked in ascending order by P-value. For those genes where a significant change was apparent at both the protein and mRNA level, the smaller P-value was used.
Metabolic pathways and membrane components affected by doxycycline stress in Wolbachia strain wMel. Summary of the major metabolic pathways, membrane transporters and respiratory chain components that are affected by doxycycline exposure. Differentially expressed gene transcripts are coloured blue and differentially expressed proteins are coloured red. Arrows indicate the direction of change (up represents a significant increase under doxycycline stress). Numbers in parentheses denote the fold change between doxycycline-treated and control samples.
Three inner membrane transporters were upregulated following exposure to doxycycline, including the ATPase subunit (PstB) of the high-affinity phosphate uptake system (Supplementary Tables S5 and S6; Figure 5). Mutations in the pstB gene lead to increased susceptibility to aminoglycosides in Pseudomonas aeruginosa (Krahn et al., 2012), whereas overexpression of this gene is associated with ciprofloxacin resistance in Mycobacterium smegmatis (Bhatt et al., 2000). Moreover, the antimicrobial preservative sodium benzoate has been shown to increase transcription of pst genes in E. coli O157:H7 (Critzer et al., 2010). Upregulation of PstB by doxycycline in Wolbachia could reflect an increased demand for nucleotide or phospholipid synthesis, or may provide a means of drug efflux. In addition, a sodium-alanine symporter (the most highly regulated annotated transcript in our study) and a tatA homologue exhibited significant upregulation (Supplementary Table S5). In E. coli, the twin-arginine translocation (Tat) system consists of three proteins (TatA, TatB and TatC) and is responsible for the export of fully folded proteins, including redox enzymes requiring cofactor insertion before translocation (Harrison et al., 2005; Lee et al., 2006). One predicted substrate of the Tat translocase in the Anaplasmataceae (Nuñez et al., 2012) is the fatty acid biosynthesis enzyme 3-oxoacyl-ACP reductase (fabG), which also displayed elevated gene expression after doxycycline exposure (Supplementary Table S5). Interestingly, an upregulated hypothetical protein (WD0222; Supplementary Table S8) showed some amino-acid sequence similarity with TatB from Anaplasma marginale (65% query coverage, 29% similarity), and the latter provides partial restoration of function to E. coli ΔtatB mutants (Nuñez et al., 2012).
The wMel genome encodes enzymes for the synthesis of riboflavin and flavin adenine dinucleotide, which are essential cofactors that are provisioned by strain wBm in its mutualistic relationship with Brugia malayi, a human filarial pathogen (Li and Carlow, 2012). Two of the enzymes in this pathway showed DE at the protein level in different directions (Figure 5). Although the net effect of these changes would require experimental validation, comparisons with other bacteria suggest that guanosine triphosphate (GTP) cyclohydrolase II constitutes the rate-limiting enzyme (Hümbelin et al., 1999), and thus doxycycline probably suppresses riboflavin synthesis. Indeed, it has been known since the early phase of the antibiotic era that tetracyclines inhibit riboflavin metabolism; conversely, riboflavin is a competitive inhibitor of tetracycline-induced bacteriostasis (Foster and Pittillo, 1953). Moreover, this interaction is potentially deleterious to bacterial cells, as the vitamin acts as a photosensitizer of the drug, producing reactive oxygen species in the presence of visible light (Castillo et al., 2007).
Effects on host–symbiont interactions and global regulators
As ANKs represent a protein domain that is widespread in eukaryotes but relatively scarce in prokaryotes (Sedgwick and Smerdon, 1999), the high numbers of ANK-encoding genes in arthropod-associated Wolbachia have led to the speculation that they may be important in symbiont–host interactions (Siozios et al., 2013). Despite this, substantive functional data for the role of ANKs in Wolbachia have yet to be obtained. In arthropod Wolbachia, ANK-encoding genes are often found in close proximity to phage clusters, and there is evidence for lateral gene transfer via bacteriophages both within and between Wolbachia supergroups (Ellegaard et al., 2013; Siozios et al., 2013). In our study, four ANKs, including three located in prophage regions, were downregulated after doxycycline treatment (Figure 4, Supplementary Tables S5 and S6). This suggests that doxycycline could have an impact on interactions between Wolbachia and host cells during a relatively short treatment period, even if symbiont densities are not substantially affected. Indeed, it is possible that this contributes to the mechanism of action of doxycycline against filarial Wolbachia such as wBm, in which an orthologue of ANK WD0754 is conserved (Foster et al., 2005; Supplementary Table S6). In addition, our data strongly suggest that doxycycline does not induce the lytic cycle of bacteriophage WO in wMelPop-CLA, despite the well-known ability of the tetracyclines to enhance phage production in free-living bacteria (Kaur et al., 2012).
The characterisation of global regulation in obligate intracellular bacteria is constrained by the paucity of available genetic tools, although gene expression studies in which external stressors are applied can provide valuable insights into potential regulatory networks. In this context, we observed upregulation of two transcripts involved in cell division in our study: the bacterial ‘tubulin’ ftsZ, and a gene encoding a Fic family protein (Supplementary Table S5). In E. coli, the adenylylase Fic is essential for cell division (Komano et al., 1991) and may have a role in intracellular signalling, although in some pathogenic bacteria it is secreted as a virulence factor that AMPylates host cell proteins (Roy and Mukherjee, 2009). In Wolbachia, it is probable that upregulation of these transcripts forms part of a regulatory effort to overcome cellular stasis following inhibition of translation. Two other transcripts that have global regulatory roles in other bacteria were downregulated following drug exposure: ctrA and ssrS (6S RNA) (Supplementary Table S5). The response regulator CtrA has been demonstrated in Ehrlichia chaffeensis (family Anaplasmataceae) to be upregulated during the differentiation of dense-cored cells and is associated with stress resistance (Cheng et al., 2011); hence, it is surprising that ctrA transcripts decreased during doxycycline treatment of Wolbachia. Furthermore, we failed to detect any significant changes in other components of the E. chaffeensis CtrA regulon (BolA, SurE and peptidoglycan-associated lipoprotein), despite their conservation in wMel. It is possible that although CtrA is present in all members of the Anaplasmataceae sequenced to date, its role has diverged between genera; indeed, significant differences exist in its C-terminal amino-acid residues between Anaplasma, Ehrlichia, Neorickettsia and Wolbachia (Cheng et al., 2011).
The non-coding RNA ssrS is an important factor controlling the rate of intracellular replication in Legionella pneumophila, as deletion of this gene reduces the ability of the pathogen to multiply in eukaryotic cells (Faucher et al., 2010). In a transcriptomic study of Wolbachia strain wOo from the filarial nematode Onchocerca ochengi, the expression of ssrS was significantly higher in symbionts located in the female worm gonad, where bacterial replication is relatively rapid than in somatic tissues in which Wolbachia divides slowly (Darby et al., 2012). Thus, ssrS is a key contender as a global regulator of growth rate in Wolbachia, and its reduced expression following exposure to a bacteriostat supports such a role.
Susceptibility to tetracyclines in the Rickettsiales
Although the tetracycline derivatives have been in clinical use since the late 1940s, they remain the treatment of choice for infections caused by obligate intracellular bacteria (McOrist, 2000). In marked contrast to recent global trends for free-living bacterial pathogens, reports of antibiotic resistance in obligate intracellular bacteria are very rare, and geographical spread of such isolates has not become clinically significant to date. For instance, a 2- to 5-day course of doxycycline is still effective for the treatment of spotted fever group rickettsiae (Botelho-Nevers et al., 2012), and although isolates of the scrub typhus agent O. tsutsugamushi with reduced doxycycline susceptibility were reported from northern Thailand in the mid-1990s (Watt et al., 1996) this drug has remained effective in other endemic areas (Rajapakse et al., 2011). However, despite this apparent lack of heritable resistance, many members of the Rickettsiales can persist following tetracycline treatment and may recrudesce months or even years after the cessation of therapy (Andrew and Norval, 1989; Stuen and Bergstrom, 2001; McClure et al., 2010). This is also the case for Wolbachia, as a 2-week course of oxytetracycline only depleted the bacteria transiently from O. ochengi worms, with full recovery of both the endosymbionts and their host 6 months later (Gilbert et al., 2005). Failure of tetracyclines to penetrate into worms can effectively be ruled out as an explanation for the necessity of prolonged treatment, as tetracycline accumulation in filariae has been visualised during human infection by autofluorescence (Tobie and Beye, 1960).
These reports suggest that Wolbachia and its relatives are capable of producing subpopulations that are phenotypically tolerant to tetracyclines. Indeed, as antibiotic tolerance or ‘metabolic resistance’ is a universal property of bacteria and yeasts (Lewis, 2012), its occurrence in the Rickettsiales is inevitable. However, the potential significance of the phenomenon apparently has not been recognised, perhaps because anti-Wolbachia therapy of filariae is a relatively recent development that accentuates this feature of the Rickettsiales. Two characteristics of filarial Wolbachia appear to facilitate tetracycline tolerance: the low level of bacterial growth in the adult worm soma (McGarry et al., 2004) and their location beneath a physical barrier (the nematode cuticle) that is normally impervious to the mammalian immune system (Hansen et al., 2011). These properties may promote tolerance because the proportion of ‘persister’ bacteria is 10−5 during exponential growth, but increases to 10-2 in stationary phase (Lewis, 2007).
Mechanisms of antibiotic tolerance
To date, most studies of antibiotic tolerance have focused on free-living bacteria and facultative intracellular pathogens exposed to bactericidal antibiotics. Functional analyses of the persister phenotype in tractable organisms have identified global regulators, toxin–antitoxin modules and antioxidant enzymes as having critical roles in maintaining antibiotic tolerance in slow-growing subpopulations (Kint et al., 2012). However, many of these candidate molecules and their associated pathways are either absent in Wolbachia genomes or have become severely limited. For instance, sigma factors are important for persistence in mycobacteria (Michele et al., 1999), but Wolbachia has only two of these regulators and the expression of neither was changed in our study. Similarly, decades of research in E. coli and more recently in M. tuberculosis have highlighted the central role of toxin–antitoxin modules in antibiotic tolerance (Gerdes and Maisonneuve, 2012). Although we did note the upregulation of a relE toxin-like transcript (Supplementary Table S8), this gene is not conserved among other Wolbachia genomes (including those from nematode strains) and levels of the Lon protease that mediates its effects (Maisonneuve et al., 2011) were unaffected in our study. Thus, it is unlikely that toxin–antitoxin modules are of general importance in the response of Wolbachia to doxycycline. Conversely, some persister gene candidates did exhibit parallels between studies in E. coli and our own experiment. Specifically, knockout of the dnaK gene (Supplementary Table S5) reduced the rate of persister formation in E. coli by 22-fold in one study, and deletion of a gene (ygfA) involved in the stabilisation of ssrS (although not knockout of ssrS itself) also had a significant, four-fold effect (Hansen et al., 2008). However, we could not identify a homologue of ygfA in any Wolbachia strain by BLAST analysis. Finally, in the context of the reduced levels of oxidative stress associated with antibiotic tolerance in P. aeruginosa (Nguyen et al., 2011), the upregulation of ppnK (Supplementary Table S5) in our study suggests that Wolbachia also induces an antioxidant response following antibiotic exposure. Nevertheless, the underlying mechanism must be quite different from that for P. aeruginosa, as Wolbachia apparently lacks the stringent response pathway.
Potential impacts on the symbiotic relationship
The global regulation of gene expression in Wolbachia during antibiotic stress is further complicated by the potential demands of the host cell in the symbiotic partnership. Clearly, in this context, caution is required when extrapolating data obtained from an arthropod symbiont cultured in vitro to congeneric endobacteria in filarial nematodes. For instance, uncharacterised phage-associated protein genes (including those with ANK domains) are almost entirely absent in filarial Wolbachia, and the putative deletion in the wMelPop-CLA chromosome (Supplementary Table S1) suggests a degree of genomic plasticity that is probably unlikely in filarial Wolbachia. However, it is noteworthy that the pathways containing the largest number of genes undergoing DE were involved in highly conserved aspects of core metabolism (e.g., de novo nucleotide synthesis and energy generation). In a previous gene expression study of Wolbachia (strain wOo), ATP provisioning was identified as the primary contender for the contribution of the symbiont to the mutualistic relationship (Darby et al., 2012). The observed upregulation of PstB may represent an additional compensatory process to maintain ATP production, although in this case coupling of phosphate uptake to drug efflux (Bhatt et al., 2000) could be critical to the survival of the microbial partner.
As natural and anthropogenic sources of antibiotics are widely dispersed in the environment, significant exposure of Wolbachia to these compounds in arthropod hosts is probably commonplace. Indeed, recent studies have examined the impact of antimicrobial compounds on arthropod-associated microbiota in both terrestrial (Adams et al., 2011) and aquatic (Edlund et al., 2012) habitats, including in a collembolan species (Folsomia candida) that has an obligate dependency on Wolbachia for egg hatching (Timmermans and Ellers, 2009; Giordano et al., 2010). Interestingly, oral dosing of F. candida with oxytetracycline failed to eliminate the Wolbachia infection, and the authors speculated that this was due to detoxification of the antibiotic by the host, its intestinal flora, or ‘resistant’ Wolbachia (Giordano et al., 2010). In addition, the potential complexity of interactions between symbionts and environmental pollutants was revealed by an expression microarray analysis in F. candida exposed to cadmium, which demonstrated an increase in gene expression across the penicillin and cephalosporin biosynthesis pathway (Nota et al., 2008). Although Wolbachia is not susceptible to β-lactams and the impact of cadmium on the collembolan microbiome was not quantified, this study suggests that the population dynamics of arthropod symbionts could be disrupted not only by antibiotics but also by indirect effects of abiotic stressors.
Conclusions
Our integrated transcriptomic and proteomic study has revealed that a 70-year-old drug with a well-defined target displays a plethora of effects on gene expression, even in a bacterium with a highly reduced genome. Thus, our data support the emerging paradigm of antibiotics as agents that act by exploiting and disrupting genome-wide bacterial regulatory networks (Kohanski et al., 2010), which, in the context of a mutualistic symbiosis, may be equally or more important than inhibiting bacterial growth. A fuller understanding of the ability of obligate intracellular bacteria to compensate for tetracycline-mediated inhibition of translation may allow the development of new strategies to overcome persistence in these remarkably resilient organisms.
References
Adams AS, Boone CK, Bohlmann J, Raffa KF . (2011). Responses of bark beetle-associated bacteria to host monoterpenes and their relationship to insect life histories. J Chem Ecol 37: 808–817.
Andrew HR, Norval RAI . (1989). The carrier status of sheep, cattle and african buffalo recovered from heartwater. Vet Parasitol 34: 261–266.
Bandi C, Anderson TJC, Genchi C, Blaxter ML . (1998). Phylogeny of Wolbachia in filarial nematodes. Proc R Soc B 265: 2407–2413.
Bennuru S, Meng Z, Ribeiro JMC, Semnani RT, Ghedin E, Chan K et al (2011). Stage-specific proteomic expression patterns of the human filarial parasite Brugia malayi and its endosymbiont Wolbachia. Proc Natl Acad Sci USA 108: 9649–9654.
Bhatt K, Banerjee SK, Chakraborti PK . (2000). Evidence that phosphate specific transporter is amplified in a fluoroquinolone resistant Mycobacterium smegmatis. Eur J Biochem 267: 4028–4032.
Botelho-Nevers E, Socolovschi C, Raoult D, Parola P . (2012). Treatment of Rickettsia spp. infections: a review. Expert Rev Anti Infect Ther 10: 1425–1437.
Castillo C, Criado S, Díaz M, García NA . (2007). Riboflavin as a sensitiser in the photodegradation of tetracyclines. Kinetics, mechanism and microbiological implications. Dyes Pigm 72: 178–184.
Cerveau N, Leclercq S, Leroy E, Bouchon D, Cordaux R . (2011). Short- and long-term evolutionary dynamics of bacterial insertion sequences: insights from Wolbachia endosymbionts. Genome Biol Evol 3: 1175–1186.
Chao CC, Garland DL, Dasch GA, Ching WM . (2009). Comparative proteomic analysis of antibiotic-sensitive and insensitive isolates of Orientia tsutsugamushi. Ann N Y Acad Sci 1166: 27–37.
Cheng Z, Miura K, Popov VL, Kumagai Y, Rikihisa Y . (2011). Insights into the CtrA regulon in development of stress resistance in obligatory intracellular pathogen Ehrlichia chaffeensis. Mol Microbiol 82: 1217–1234.
Critzer FJ, D'Souza DH, Saxton AM, Golden DA . (2010). Increased transcription of the phosphate-specific transport system of Escherichia coli O157: H7 after exposure to sodium benzoate. J Food Prot 73: 819–824.
Darby AC, Armstrong SD, Bah GS, Kaur G, Hughes MA, Kay SM et al (2012). Analysis of gene expression from the Wolbachia genome of a filarial nematode supports both metabolic and defensive roles within the symbiosis. Genome Res 22: 2467–2477.
Dedeine F, Vavre F, Fleury F, Loppin B, Hochberg ME, Bouletreau M . (2001). Removing symbiotic Wolbachia bacteria specifically inhibits oogenesis in a parasitic wasp. Proc Natl Acad Sci USA 98: 6247–6252.
Duplouy A, Iturbe-Ormaetxe I, Beatson SA, Szubert JM, Brownlie JC, McMeniman CJ et al (2013). Draft genome sequence of the male-killing Wolbachia strain wBol1 reveals recent horizontal gene transfer from diverse sources. BMC Genomics 14: 20.
Edlund A, Ek K, Breitholtz M, Gorokhova E . (2012). Antibiotic-induced change of bacterial communities associated with the copepod Nitocra spinipes. PLoS One 7: 9.
Ellegaard KM, Klasson L, Näslund K, Bourtzis K, Andersson SGE . (2013). Comparative genomics of Wolbachia and the bacterial species concept. PLoS Genet 9: e1003381.
Evers S, Di Padova K, Meyer M, Langen H, Fountoulakis M, Keck W et al (2001). Mechanism-related changes in the gene transcription and protein synthesis patterns of Haemophilus influenzae after treatment with transcriptional and translational inhibitors. Proteomics 1: 522–544.
Faucher SP, Friedlander G, Livny J, Margalit H, Shuman HA . (2010). Legionella pneumophila 6S RNA optimizes intracellular multiplication. Proc Natl Acad Sci USA 107: 7533–7538.
Ferri E, Bain O, Barbuto M, Martin C, Lo N, Uni S et al (2011). New insights into the evolution of Wolbachia infections in filarial nematodes inferred from a large range of screened species. PLoS One 6: e20843.
Foster J, Ganatra M, Kamal I, Ware J, Makarova K, Ivanova N et al (2005). The Wolbachia genome of Brugia malayi: endosymbiont evolution within a human pathogenic nematode. PLoS Biol 3: e121.
Foster J, Pittillo RF . (1953). Reversal by complex natural materials of growth inhibition caused by antibiotics. J Bacteriol 65: 361–367.
Geniez S, Foster JM, Kumar S, Moumen B, Leproust E, Hardy O et al (2012). Targeted genome enrichment for efficient purification of endosymbiont DNA from host DNA. Symbiosis 58: 201–207.
Gerdes K, Maisonneuve E . (2012). Bacterial persistence and toxin-antitoxin loci. Annu Rev Microbiol 66: 103–123.
Gilbert J, Nfon CK, Makepeace BL, Njongmeta LM, Hastings IM, Pfarr KM et al (2005). Antibiotic chemotherapy of onchocerciasis: in a bovine model, killing of adult parasites requires a sustained depletion of endosymbiotic bacteria (Wolbachia species). J Infect Dis 192: 1483–1493.
Giordano R, Weber E, Waite J, Bencivenga N, Krogh PH, Soto-Adames F . (2010). Effect of a high dose of three antibiotics on the reproduction of a parthenogenetic strain of Folsomia candida (Isotomidae: Collembola). Environ Entomol 39: 1170–1177.
Godel C, Kumar S, Koutsovoulos G, Ludin P, Nilsson D, Comandatore F et al (2012). The genome of the heartworm, Dirofilaria immitis, reveals drug and vaccine targets. FASEB J 26: 4650–4661.
Griffin MO, Fricovsky E, Ceballos G, Villarreal F . (2010). Tetracyclines: a pleitropic family of compounds with promising therapeutic properties. Review of the literature. Am J Physiol Cell Physiol 299: C539–C548.
Haegeman A, Vanholme B, Jacob J, Vandekerckhove TTM, Claeys M, Borgonie G et al (2009). An endosymbiotic bacterium in a plant-parasitic nematode: member of a new Wolbachia supergroup. Int J Parasitol 39: 1045–1054.
Hansen RDE, Trees AJ, Bah GS, Hetzel U, Martin C, Bain O et al (2011). A worm's best friend: recruitment of neutrophils by Wolbachia confounds eosinophil degranulation against the filarial nematode Onchocerca ochengi. Proc R Soc B 278: 2293–2302.
Hansen S, Lewis K, Vulić M . (2008). Role of global regulators and nucleotide metabolism in antibiotic tolerance in Escherichia coli. Antimicrob Agents Chemother 52: 2718–2726.
Harrison JJ, Ceri H, Badry EA, Roper NJ, Tomlin KL, Turner RJ . (2005). Effects of the twin-arginine translocase on the structure and antimicrobial susceptibility of Escherichia coli biofilms. Can J Microbiol 51: 671–683.
Hoerauf A, Nissen-Pähle K, Schmetz C, Henkle-Dührsen K, Blaxter ML, Büttner DW et al (1999). Tetracycline therapy targets intracellular bacteria in the filarial nematode Litomosoides sigmodontis and results in filarial infertility. J Clin Invest 103: 11–18.
Hoerauf A, Pfarr K, Mand S, Debrah AY, Specht S . (2011). Filariasis in Africa-treatment challenges and prospects. Clin Microbiol Infect 17: 977–985.
Hoerauf A, Volkmann L, Hamelmann C, Adjei O, Autenrieth IB, Fleischer B et al (2000). Endosymbiotic bacteria in worms as targets for a novel chemotherapy in filariasis. Lancet 355: 1242–1243.
Hosokawa T, Koga R, Kikuchi Y, Meng XY, Fukatsu T . (2010). Wolbachia as a bacteriocyte-associated nutritional mutualist. Proc Natl Acad Sci USA 107: 769–774.
Hümbelin M, Griesser V, Keller T, Schurter W, Haiker M, Hohmann HP et al (1999). GTP cyclohydrolase II and 3,4-dihydroxy-2-butanone 4-phosphate synthase are rate-limiting enzymes in riboflavin synthesis of an industrial Bacillus subtilis strain used for riboflavin production. J Ind Microbiol Biotechnol 22: 1–7.
Kambris Z, Cook PE, Phuc HK, Sinkins SP . (2009). Immune activation by life-shortening Wolbachia and reduced filarial competence in mosquitoes. Science 326: 134–136.
Kaur S, Harjai K, Chhibber S . (2012). Methicillin-resistant Staphylococcus aureus phage plaque size enhancement using sublethal concentrations of antibiotics. Appl Environ Microbiol 78: 8227–8233.
Kent BN, Bordenstein SR . (2010). Phage WO of Wolbachia: lambda of the endosymbiont world. Trends Microbiol 18: 173–181.
Kint CI, Verstraeten N, Fauvart M, Michiels J . (2012). New-found fundamentals of bacterial persistence. Trends Microbiol 20: 577–585.
Kohanski MA, Dwyer DJ, Collins JJ . (2010). How antibiotics kill bacteria: from targets to networks. Nat Rev Microbiol 8: 423–435.
Komano T, Utsumi R, Kawamukai M . (1991). Functional analysis of the fic gene involved in regulation of cell division. Res Microbiol 142: 269–277.
Krahn T, Gilmour C, Tilak J, Fraud S, Kerr N, Lau CHF et al (2012). Determinants of intrinsic aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 56: 5591–5602.
Langworthy NG, Renz A, Mackenstedt U, Henkle-Dührsen K, De C Bronsvoort MB, Tanya VN et al (2000). Macrofilaricidal activity of tetracycline against the filarial nematode Onchocerca ochengi: elimination of Wolbachia precedes worm death and suggests a dependent relationship. Proc R Soc B 267: 1063–1069.
Lee PA, Tullman-Ercek D, Georgiou G . (2006). The bacterial twin-arginine translocation pathway. Annu Rev Microbiol 60: 373–395.
Lessner FH, Venters BJ, Keiler KC . (2007). Proteolytic adaptor for transfer-messenger RNA-tagged proteins from α-proteobacteria. J Bacteriol 189: 272–275.
Lewis K . (2007). Persister cells, dormancy and infectious disease. Nat Rev Microbiol 5: 48–56.
Lewis K . (2012). Persister cells: molecular mechanisms related to antibiotic tolerance. Handb Exp Pharmacol 211: 121–133.
Li H, Durbin R . (2009). Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25: 1754–1760.
Li Z, Carlow CKS . (2012). Characterization of transcription factors that regulate the type IV secretion system and riboflavin biosynthesis in Wolbachia of Brugia malayi. PLoS One 7: e51597.
Lin XM, Yang JN, Peng XX, Li H . (2010). A novel negative regulation mechanism of bacterial outer membrane proteins in response to antibiotic resistance. J Proteome Res 9: 5952–5959.
Maisonneuve E, Shakespeare LJ, Jørgensen MG, Gerdes K . (2011). Bacterial persistence by RNA endonucleases. Proc Natl Acad Sci USA 108: 13206–13211.
Makepeace BL, Rodgers L, Trees AJ . (2006). Rate of elimination of Wolbachia pipientis by doxycycline in vitro increases following drug withdrawal. Antimicrob Agents Chemother 50: 922–927.
McClure JC, Crothers ML, Schaefer JJ, Stanley PD, Needham GR, Ewing SA et al (2010). Efficacy of a doxycycline treatment regimen initiated during three different phases of experimental ehrlichiosis. Antimicrob Agents Chemother 54: 5012–5020.
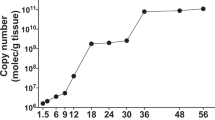
McGarry HF, Egerton GL, Taylor MJ . (2004). Population dynamics of Wolbachia bacterial endosymbionts in Brugia malayi. Mol Biochem Parasitol 135: 57–67.
McMeniman CJ, Lane AM, Fong AWC, Voronin DA, Iturbe-Ormaetxe I, Yamada R et al (2008). Host adaptation of a Wolbachia strain after long-term serial passage in mosquito cell lines. Appl Environ Microbiol 74: 6963–6969.
McMeniman CJ, Lane RV, Cass BN, Fong AWC, Sidhu M, Wang YF et al (2009). Stable introduction of a life-shortening Wolbachia infection into the mosquito Aedes aegypti. Science 323: 141–144.
McOrist S . (2000). Obligate intracellular bacteria and antibiotic resistance. Trends Microbiol 8: 483–486.
Michele TM, Ko C, Bishai WR . (1999). Exposure to antibiotics induces expression of the Mycobacterium tuberculosis sigF gene: implications for chemotherapy against mycobacterial persistors. Antimicrob Agents Chemother 43: 218–225.
Miller WJ, Ehrman L, Schneider D . (2010). Infectious speciation revisited: impact of symbiont-depletion on female fitness and mating behavior of Drosophila paulistorum. PLoS Pathog 6: e1001214.
Moreira LA, Iturbe-Ormaetxe I, Jeffery JA, Lu G, Pyke AT, Hedges LM et al (2009). A Wolbachia symbiont in Aedes aegypti limits infection with dengue, chikungunya, and Plasmodium. Cell 139: 1268–1278.
Ng WL, Kazmierczak KM, Robertson GT, Gilmour R, Winkler ME . (2003). Transcriptional regulation and signature patterns revealed by microarray analyses of Streptococcus pneumoniae R6 challenged with sublethal concentrations of translation inhibitors. J Bacteriol 185: 359–370.
Nguyen D, Joshi-Datar A, Lepine F, Bauerle E, Olakanmi O, Beer K et al (2011). Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 334: 982–986.
Nota B, Timmermans M, Franken O, Montagne-Wajer K, Marien J, De Boer ME et al (2008). Gene expression analysis of Collembola in cadmium containing soil. Environ Sci Technol 42: 8152–8157.
Nuñez PA, Soria M, Farber MD . (2012). The twin-arginine translocation pathway in α-proteobacteria is functionally preserved irrespective of genomic and regulatory divergence. PLoS One 7: e33605.
Rajapakse S, Rodrig C, Fernando SD . (2011). Drug treatment of scrub typhus. Trop Doct 41: 1–4.
Rasgon JL, Gamston CE, Ren X . (2006). Survival of Wolbachia pipientis in cell-free medium. Appl Environ Microbiol 72: 6934–6937.
Riegler M, Sidhu M, Miller WJ, O’Neill SL . (2005). Evidence for a global Wolbachia replacement in Drosophila melanogaster. Curr Biol 15: 1428–1433.
Robinson MD, McCarthy DJ, Smyth GK . (2010). EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26: 139–140.
Robinson MD, Oshlack A . (2010). A scaling normalisation method for differential expression analysis of RNA-seq data. Genome Biol 11: R25.
Roy CR, Mukherjee S . (2009). Bacterial FIC proteins AMP up infection. Sci Signal 2: pe14.
Schiefer A, Schmitz A, Schäberle TF, Specht S, Lämmer C, Johnston KL et al (2012). Corallopyronin a specifically targets and depletes essential obligate Wolbachia endobacteria from filarial nematodes in vivo. J Infect Dis 206: 249–257.
Schiefer A, Vollmer J, Lammer C, Specht S, Lentz C, Ruebsamen-Schaeff H et al (2013). The ClpP peptidase of Wolbachia endobacteria is a novel target for drug development against filarial infections. J Antimicrob Chemother 68: 1790–1800.
Sedgwick SG, Smerdon SJ . (1999). The ankyrin repeat: a diversity of interactions on a common structural framework. Trends Biochem Sci 24: 311–316.
Siozios S, Ioannidis P, Klasson L, Andersson SGE, Braig HR, Bourtzis K . (2013). The diversity and evolution of Wolbachia ankyrin repeat domain genes. PLoS One 8: e55390.
Specht S, Mand S, Marfo-Debrekyei Y, Debrah AY, Konadu P, Adjei O et al (2008). Efficacy of 2- and 4-week rifampicin treatment on the Wolbachia of Onchocerca volvulus. Parasitol Res 103: 1303–1309.
Stuen S, Bergstrom K . (2001). The effect of two different oxytetracycline treatments in experimental Ehrlichia phagocytophila infected lambs. Acta Vet Scand 42: 339–346.
Taylor MJ, Makunde WH, McGarry HF, Turner JD, Mand S, Hoerauf A . (2005). Macrofilaricidal activity after doxycycline treatment of Wuchereria bancrofti: a double-blind, randomised placebo-controlled trial. Lancet 365: 2116–2121.
Timmermans M, Ellers J . (2009). Wolbachia endosymbiont is essential for egg hatching in a parthenogenetic arthropod. Evol Ecol 23: 931–942.
Tobie JE, Beye HK . (1960). Fluorescence of tetracyclines in filarial worms. Proc Soc Exp Biol Med 104: 137–140.
VanBogelen RA, Neidhardt FC . (1990). Ribosomes as sensors of heat and cold shock in Escherichia coli. Proc Natl Acad Sci USA 87: 5589–5593.
Vizcaino JA, Cote RG, Csordas A, Dianes JA, Fabregat A, Foster JM et al (2013). The Proteomics identifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res 41: D1063–D1069.
Waldbauer JR, Rodrigue S, Coleman ML, Chisholm SW . (2012). Transcriptome and proteome dynamics of a light-dark synchronized bacterial cell cycle. PLoS One 7: e43432.
Watt G, Chouriyagune C, Ruangweerayud R, Watcharapichat P, Phulsuksombati D, Jongsakul K et al (1996). Scrub typhus infections poorly responsive to antibiotics in northern Thailand. Lancet 348: 86–89.
Werren JH, Baldo L, Clark ME . (2008). Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6: 741–751.
Wiśniewski JR, Zougman A, Mann M . (2009). Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J Proteome Res 8: 5674–5678.
Wu M, Sun LV, Vamathevan J, Riegler M, Deboy R, Brownlie JC et al (2004). Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: a streamlined genome overrun by mobile genetic elements. PLoS Biol 2: E69.
Yun SH, Choi CW, Park SH, Lee JC, Leem SH, Choi JS et al (2008). Proteomic analysis of outer membrane proteins from Acinetobacter baumannii DU202 in tetracycline stress condition. J Microbiol 46: 720–727.
Zug R, Hammerstein P . (2012). Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS One 7: e38544.
Acknowledgements
We gratefully acknowledge provision of the wMelPop-CLA-infected RML-12 cell line and pre-publication access to genomic data for wMelPop-CLA by Megan Woolfit, Iñaki Iturbe-Ormaetxe and Scott O’Neill (Monash University). We also thank Pia Koldkjær and Margaret Hughes (Centre for Genomic Research) for performing the RNA library preparations, and Sujai Kumar, Charlotte Repton and Mark Blaxter (University of Edinburgh) for permission to use the wLs and wDi genome assemblies in our orthologous cluster analysis. This study was supported by the 7th Framework programme of the European Commission (project identifier HEALTH-F3-2010-242131).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Darby, A., Christina Gill, A., Armstrong, S. et al. Integrated transcriptomic and proteomic analysis of the global response of Wolbachia to doxycycline-induced stress. ISME J 8, 925–937 (2014). https://doi.org/10.1038/ismej.2013.192
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.192
Keywords
This article is cited by
-
Omics analysis of Mycobacterium tuberculosis isolates uncovers Rv3094c, an ethionamide metabolism-associated gene
Communications Biology (2023)
-
Wolbachia: endosymbiont of onchocercid nematodes and their vectors
Parasites & Vectors (2021)
-
Best practices on the differential expression analysis of multi-species RNA-seq
Genome Biology (2021)
-
Association of Wolbachia with Gene Expression in Drosophila Testes
Microbial Ecology (2021)
-
Similarities and differences between 6S RNAs from Bradyrhizobium japonicum and Sinorhizobium meliloti
Journal of Microbiology (2020)