Abstract
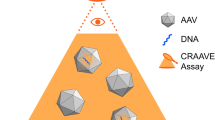
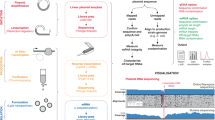
There is a recognised need for standardisation of protocols for vector genome analysis used in vector manufacturing, to establish dosage, in biodistribution studies and to detect gene doping in sport. Analysis of vector genomes and transgene expression is typically performed by qPCR using plasmid-based calibrants incorporating transgenic sequences. These often undergo limited characterisation and differ between manufacturers, potentially leading to inaccurate quantification, inconsistent inter-laboratory results and affecting clinical outcomes. Contamination of negative samples with such calibrants could cause false positive results. We developed a design strategy for synthetic reference materials (RMs) with modified transgenic sequences to prevent false positives due to cross-contamination. When such RM is amplified in transgene-specific assays, the amplicons are distinguishable from transgene’s amplicons based on size and sequence. Using human erythropoietin as a model, we produced certified RM according to this strategy and following ISO Guide 35. Using non-viral and viral vectors, we validated the effectiveness of this RM in vector genome analysis in blood in vitro. The developed design strategy could be applied to production of RMs for other transgenes, genes or transcripts. Together with validated PCR assays, such RMs form a measurement tool that facilitates standardised, accurate and reliable genetic analysis in various applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wilson JM . Bulls, bubbles, and biotech. Hum Gene Ther 2013; 24: 715–716.
Ayuso E, Blouin V, Lock M, McGorray S, Leon X, Alvira MR et al. Manufacturing and characterization of a recombinant adeno-associated virus type 8 reference standard material. Hum Gene Ther 2014; 25: 977–987.
Lock M, McGorray S, Auricchio A, Ayuso E, Beecham EJ, Blouin-Tavel V et al. Characterization of a recombinant adeno-associated virus type 2 Reference Standard Material. Hum Gene Ther 2010; 21: 1273–1285.
Baoutina A, Coldham T, Bains GS, Emslie KR . Gene doping detection: evaluation of approach for direct detection of gene transfer using erythropoietin as a model system. Gene Ther 2010; 17: 1022–1032.
Baoutina A, Coldham T, Fuller B, Emslie KR . Improved detection of transgene and nonviral vectors in blood. Hum Gene Ther Methods 2013; 24: 345–354.
Beiter T, Zimmermann M, Fragasso A, Hudemann J, Niess AM, Bitzer M et al. Direct and long-term detection of gene doping in conventional blood samples. Gene Ther 2011; 18: 225–231.
Ni W, Le Guiner C, Gernoux G, Penaud-Budloo M, Moullier P, Snyder RO . Longevity of rAAV vector and plasmid DNA in blood after intramuscular injection in nonhuman primates: implications for gene doping. Gene Ther 2011; 18: 709–718.
Lock M, Alvira MR, Chen SJ, Wilson JM . Absolute determination of single-stranded and self-complementary adeno-associated viral vector genome titers by droplet digital PCR. Hum Gene Ther Methods 2014; 25: 115–125.
Boehme P, Stellberger T, Solanki M, Zhang W, Schulz E, Bergmann T et al. Standard free droplet digital polymerase chain reaction as a new tool for the quality control of high-capacity adenoviral vectors in small-scale preparations. Hum Gene Ther Methods 2015; 26: 25–34.
Nixon G, Garson JA, Grant P, Nastouli E, Foy CA, Huggett JF . Comparative study of sensitivity, linearity, and resistance to inhibition of digital and nondigital polymerase chain reaction and loop mediated isothermal amplification assays for quantification of human cytomegalovirus. Anal Chem 2014; 86: 4387–4394.
Pavsic J, Zel J, Milavec M . Assessment of the real-time PCR and different digital PCR platforms for DNA quantification. Anal Bioanal Chem 2016; 408: 107–121.
Oakes B, Tai AK, Cingoz O, Henefield MH, Levine S, Coffin JM et al. Contamination of human DNA samples with mouse DNA can lead to false detection of XMRV-like sequences. Retrovirology 2010; 7: 109.
ISO. Guide 35, Reference Materials — General and Statistical Principles for Certification. 2006.
Praszeres DMF Product specifications and quality controlIn: Plasmid Biopharmaceuticals: Basics, Applications and Manufacturing. John Wiley & Sons: Hoboken, NJ, USA, 2011, pp 357–390.
Stenman J, Orpana A . Accuracy in amplification. Nat Biotechnol 2001; 19: 1011–1012.
Holden MJ, Madej RM, Minor P, Kalman LV . Molecular diagnostics: harmonization through reference materials, documentary standards and proficiency testing. Expert Rev Mol Diagn 2011; 11: 741–755.
Hou Y, Zhang H, Miranda L, Lin S . Serious overestimation in quantitative PCR by circular (supercoiled) plasmid standard: microalgal pcna as the model gene. PLoS One 2010; 5: e9545.
Haynes RJ, Kline MC, Toman B, Scott C, Wallace P, Butler JM et al. Standard reference material 2366 for measurement of human cytomegalovirus DNA. J Mol Diagn 2013; 15: 177–185.
Hayden RT, Gu Z, Sam SS, Sun Y, Tang L, Pounds S et al. Comparative evaluation of three commercial quantitative cytomegalovirus standards by use of digital and real-time PCR. J Clin Microbiol 2015; 53: 1500–1505.
White H, Deprez L, Corbisier P, Hall V, Lin F, Mazoua S et al. A certified plasmid reference material for the standardisation of BCR-ABL1 mRNA quantification by real-time quantitative PCR. Leukemia 2015; 29: 369–376.
Burke DG, Dong LH, Bhat S, Forbes-Smith M, Fu S, Pinheiro L et al. Digital polymerase chain reaction measured pUC19 marker as calibrant for HPLC measurement of DNA quantity. Anal Chem 2013; 85: 1657–1664.
Choi VW, Asokan A, Haberman RA, Samulski RJ . Production of recombinant adeno-associated viral vectorsIn: Current Protocols in Human Genetics. John Wiley & Sons, 2007, 53: pp 12.19.11–12.19.21.
Huang X, Miller W . A time-efficient, linear-space local similarity algorithm. Adv Appl Math 1991; 12: 337–357.
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H et al. Clustal W and Clustal X version 2.0. Bioinformatics (Oxford, England) 2007; 23: 2947–2948.
Zuker M . Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 2003; 31: 3406–3415.
Stothard P . The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques 2000; 28: 1104.
Owczarzy R, Tataurov AV, Wu Y, Manthey JA, McQuisten KA, Almabrazi HG et al. IDT SciTools: a suite for analysis and design of nucleic acid oligomers. Nucleic Acids Res 2008; 36: W163–W169.
Kibbe WA . OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res 2007; 35: W43–W46.
Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S et al. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem 2012; 84: 1003–1011.
Lizee G, Aerts JL, Gonzales MI, Chinnasamy N, Morgan RA, Topalian SL . Real-time quantitative reverse transcriptase-polymerase chain reaction as a method for determining lentiviral vector titers and measuring transgene expression. Hum Gene Ther 2003; 14: 497–507.
JCGM. JCGM 100 – Evaluation of Measurement data – Guide to the Expression of Uncertainty in Measurement (GUM 1995 with minor corrections); 2008.
Al-Soud WA, Radstrom P . Purification and characterization of PCR-inhibitory components in blood cells. J Clin Microbiol 2001; 39: 485–493.
Tsang JC, Lo YM . Circulating nucleic acids in plasma/serum. Pathology 2007; 39: 197–207.
Ziegler A, Zangemeister-Wittke U, Stahel RA . Circulating DNA: a new diagnostic gold mine? Cancer Treat Rev 2002; 28: 255–271.
Garcia ME, Blanco JL, Caballero J, Gargallo-Viola D . Anticoagulants interfere with PCR used to diagnose invasive aspergillosis. J Clin Microbiol 2002; 40: 1567–1568.
WADA. Guidelines — Blood Sample Collection 2014. Available from https://www.wada-ama.org/en/resources/world-anti-doping-program/guidelines-blood-sample-collection (last accessed 27 May 2016).
Baoutina A, Alexander IE, Rasko JE, Emslie KR . Developing strategies for detection of gene doping. J Gene Med 2008; 10: 3–20.
Ni W, Le Guiner C, Moullier P, Snyder RO . Development and utility of an internal threshold control (ITC) real-time PCR assay for exogenous DNA detection. PLoS One 2012; 7: e36461.
Fagone P, Wright JF, Nathwani AC, Nienhuis AW, Davidoff AM, Gray JT . Systemic errors in quantitative polymerase chain reaction titration of self-complementary adeno-associated viral vectors and improved alternative methods. Hum Gene Ther Methods 2012; 23: 1–7.
Acknowledgements
This work was supported by the World Anti-Doping Agency and the Australian Government through the Sport Anti-Doping Program of the Department of Health. We thank Michael Forbes-Smith for help with chromatography purification of RM3, Jacob McLaughlin for testing diluent evaporation from vials at 40 °C and Kate Griffiths, Vanessa Agon and Lindsey Mackay (all from National Measurement Institute) for critically reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Gene Therapy website
Supplementary information
Rights and permissions
About this article
Cite this article
Baoutina, A., Bhat, S., Zheng, M. et al. Synthetic certified DNA reference material for analysis of human erythropoietin transgene and transcript in gene doping and gene therapy. Gene Ther 23, 708–717 (2016). https://doi.org/10.1038/gt.2016.47
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gt.2016.47