Abstract
We describe the detailed clinical and molecular characterization of three patients (aged 7, 84/12 and 31 years) with overlapping microdeletions in 19p13.12, extending to 19p13.13 in two cases. The patients share the following clinical features with a recently reported 10-year-old girl with a 19p13.12 microdeletion: mental retardation (MR), psychomotor and language delay, hearing impairment, brachycephaly, anteverted nares and ear malformations. All patients share a 359-kb deleted region in 19p13.12 harboring six genes (LPHN1, DDX39, CD97, PKN1, PTGER1 and GIPC1), several of which may be MR candidates because of their function and expression pattern. LPHN1 and PKN1 are the most appealing; LPHN1 for its interaction with Shank family proteins, and PKN1 because it is involved in a variety of functions in neurons, including cytoskeletal organization. Haploinsufficiency of GIPC1 may contribute to hearing impairment for its interaction with myosin VI. A behavioral phenotype was observed in all three patients; it was characterized by overactive disorder associated with MR and stereotyped movements (ICD10) in one patient and hyperactivity in the other two. As Ptger1-null mice show behavioral inhibition and impulsive aggression with defective social interaction, PTGER1 haploinsufficiency may be responsible for the behavioral traits observed in these patients.
Similar content being viewed by others
Introduction
De novo microdeletions detected by genome-wide array analysis at 19p13 have been reported in few patients,1, 2, 3, 4, 5 all of whom had malformations and mental retardation (MR). We present the detailed clinical and molecular description of two new cases (cases 1 and 3) with partially overlapping 19p13 microdeletions identified by investigating patients with syndromic MR using array comparative genomic hybridization (aCGH). We also re-analyzed at higher resolution and obtained the clinical follow-up of one published case1 (case 2) with a cryptic 19p13.12 deletion totally overlapping that of case 1. The shortest region of overlap (SRO) between the three cases is 359 kb in size and contains candidate genes potentially responsible for their common phenotypic features (psychomotor, mental, language delay and sensorineural hearing loss). Genes outside the SRO are also discussed.
Materials and methods
Patients
All patients are described in detail in the section below. The clinical features of our subjects and of patients with overlapping 19p13.12 deletions reported by Jensen et al,2 Lysy et al,4 and Auvin et al5 are summarized in Table 1.
Case 1
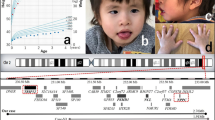
The proband is a male, the second child of healthy non-consanguineous parents aged 31 (mother) and 36 years (father) at his birth (Figure 1). Delivery was at 36 weeks of gestation by cesarean section. Birth weight was 2400 g (3–10th centile), and OFC was 32 cm (3–10th centile). Length and APGAR scores were not available. At birth, some dysmorphic features were noted: malformed ears, submucous cleft palate, bilateral cutaneous syndactyly of fingers II and III and toes and hypospadias. A standard karyotype was performed with normal results.
Photographs of case 1 at the age of 6 months (a), 8 years (b) and 17 years (c), and frontal (d) and lateral (e) view at the age of 31 years. Note the brachycephaly (a, d, e), tall forehead (a, c, d), ptosis (a–c), anteverted nares (a–d), long philtrum (a–d), thin upper lip (a–e), synophrys (a–d), short neck (c), anteverted ears (a, c, d) with thin helices (e) as well as symmetrically arranged, vaguely outlined nodules and partly confluent plaques accompanied by a fuzzy erythema on the forehead at age 31 years (d, e).
Early motor milestones were severely delayed: he acquired head control at 2 years and was able to walk without support at 3 years. He showed moderate MR and language delay. Hyperactivity was also observed.
At the age of 11 years, a diagnosis of praecox puberty was made. Cardiac ultrasound showed mild aortic and mitral valve regurgitation. Brain CT scans were normal. During adolescence, repetitive and stereotyped behavior worsened, with iterative questions and very limited interests. He developed catastrophic behavior as a response to minimal environmental changes, with auto-aggressive and sometimes hetero-aggressive acting out. A diagnosis of overactive disorder associated with MR and stereotyped movements (ICD 10) was made and specific drug therapy (clomipramine, chlorpromazine and fluvoxamine) was started with progressive improvement.
At 24 years, the patient showed several generalized clonic seizures with falls after an increase in the fluvoxamine dosage. Despite its potential pro-convulsive effects, antipsychotic therapy could not be tapered and antiepileptic chronic treatment was started with sodium valproate. Complete seizure control was achieved with a follow-up period of 8 years. EEG recordings showed a global slowing of background activity without epileptiform abnormalities.
At the age of 27 years, Auditory Brain Middle Latency Responses (ABR and MLR) and Behavioral Audiometry (VRA) exams revealed bilateral sensorineural hearing loss. The measurements obtained by pure tone audiometry (500–1000 and 2000 Hz) were 65 dBHL (right ear) and 80 dBHL (left ear). Tympanometry results were normal in the right ear, whereas the left ear was classified as type B, according to the classification of Jerger.6 Bilateral over-the-ear hearing aids were positioned. Over follow-up period of 4 years, the situation has remained stable. Scoliosis was also present at this age.
At the patient's last clinical evaluation (31 years), the following features were observed: short stature (height 150 cm; −3 SD), overweight (83 kg: BMI of 36.4 kg/m2; +3 SD), and a relative small head circumference (OFC 53.5 cm; −1.3 SD); generalized hypertrichosis; nystagmus; brachycephaly, synophrys, symmetrically arranged, vaguely outlined nodules and partly confluent plaques accompanied by a fuzzy erythema on the forehead, arched eyebrows, mild blepharophimosis, small/thin nasal root, anteverted nares, anteverted ears with thin helices, short neck (Figure 1d and e), brachydactyly (Supplementary Figure 1); bilateral cutaneous syndactyly of fingers II–IV; and bilateral cutaneous syndactyly of toes II and III. Neurological examination revealed no focal signs. He walked with small steps. Subclinical hypothyroidism and hyperlipidemia were also present. By abdominal ultrasound, mild hepatic steatosis was diagnosed; echocardiography confirmed mild aortic and mitral valve regurgitation.
Case 2
This patient was previously included in a MR cohort study (patient 11/3);1 in this study we provide a more detailed clinical and molecular characterization.
This female patient is the only child of unrelated parents, both 26 years old when she was born. Her father was known to suffer from epilepsy since childhood.
Pregnancy was initially uneventful. After week 21, IUGR was noted on repeated ultrasound examinations. Delivery took place at 32+1 weeks by cesarean section. Birth weight was in the lower normal range (between 3rd and 10th centile), length was at the third centile and microcephaly was noted (head circumference below the third centile; −2.4 SD). Shortly after birth, a mandibular neonatal tooth and a skin tag located on the tragus of the right ear were noticed.
Within the first weeks of life, weight and length dropped below the third centile. At the age of 1 month, a persisting ductus Botalli was surgically treated. Motor development was moderately delayed (unsupported sitting at 15 months and first attempts to crawl at 16 months) and there was no speech.
Abnormal results of repeated hearing tests raised the suspicion of bilateral sensorineural hearing loss. She developed muscular hypotonia and gastroesophageal reflux, and a persistent foramen ovale was noted. Cranial MRI scan, EEG and eye examination at the age of 11 months revealed no abnormalities. No evidence for metabolic disorders was found.
At the age of 16 months, the patient appeared slightly hypotonic and brachycephaly was observed. Facial dysmorphisms (synophrys, almond-shaped eyes with a bilateral epicanthic fold, long lashes, mild ptosis, flattened nasal bridge, anteverted nostrils, long philtrum and small mouth with thin upper lip) were noticed. Ears were small and posteriorly rotated and the neck was short. An atypical Simian crease was noted on her right hand.
At the age of 84/12 years, the patient had developed hyperopia and moderate MR with marked hyperactivity. Microcephaly (−3.2 SD) and brachycephaly were observed, and height and weight were within the normal range. In addition to the facial dysmorphisms described earlier, her face now appeared round and slightly flattened, and a small nasal root and thin lower lip were noted. Philtrum and neck now had a normal length. The two lateral incisors were missing in her upper jaw.
Case 3
The proband is a male, the second child of healthy, non-consanguineous parents. There is no family history of malformations or MR.
He was born by spontaneous delivery at 41 weeks of gestational age after an uncomplicated pregnancy. APGAR scores were 9/10/10. His birth weight was 2740 g (<3rd centile), length 47 cm (<3rd centile) and head circumference 34 cm (10–25th centile).
At 3 days, the patient presented remarkable hypotonia, and recurrent apneas accompanied by bradycardia without desaturation were observed. He was therefore admitted to a neonatal intensive care unit, in which a cerebral ultrasound showed a right choroid plexus cyst and subependymal hemorrhage. ECG, echocardiography and fundoscopy were also performed with normal results.
At 2 and 5 years, brain MRI scans were carried out revealing no morphological abnormalities. EMG showed normal conductance in the muscles tested and in peripheral nerves.
At the age of 5 years and 1 month, he showed a focal epileptic syndrome. He sporadically presented with focal left-sided seizures. Because of the extremely low frequency of the seizures, chronic anti-epileptic treatment was never started. EEG recordings showed epileptiform abnormalities over the right central areas.
The following distinctive features were observed: brachycephaly, small mouth with thin upper lip, long philtrum, anteverted ears, irregular teeth, bilateral epicanthic folds, brachydactyly, pectus excavatum, scoliosis, muscle hypotonia and cryptorchidism. Severe speech and neuromotor delay were also noted. His behavioral phenotype was characterized by hyperactivity.
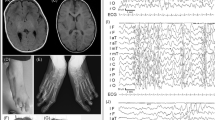
At last evaluation (7 years old; Figure 2), the patient showed moderate–severe speech and neuromotor retardation. His measurements were all within normal range. Bilateral conductive hearing loss was present.
Molecular studies
Routine karyotyping by G-banding analysis at 400–550 bands was performed using standard procedures. Array-CGH was performed with the Agilent oligo-platform at a resolution of ∼100 Kb (Agilent kit 44B, Agilent, Santa Clara, CA, USA) in all three cases. The procedures for DNA digestion, labeling and hybridization were executed according to the manufacturer's instructions. Array slides were analyzed by the Agilent scanner (G2505C) with Feature Extraction software (v10.1.1.1). Graphical overviews were obtained using the Agilent DNA analytics software (v4.0.81).
Genotyping of polymorphic loci, quantitative PCR (qPCR), long-range PCR (LR-PCR) and DNA sequencing were performed as described.7 The map used for determining the genomic positions of probes was Genome Browser Assembly hg18 (http://genome.ucsc.edu). On the basis of the aCGH characterization of the deletions, we identified the regions between last non-deleted and first deleted probes on each side of the breakpoint. For case 1, we selected four approximately evenly spaced primers on non-repeated portions of the distal breakpoint region and three primers on the proximal region, and performed LR-PCRs with all possible distal/proximal primer combinations. One combination generated a 4.5-kb fragment that was entirely sequenced. For case 2, we selected three qPCR primer pairs in non-repeated portions of the distal breakpoint region, two pairs in the proximal region, and narrowed both regions down to 3 to 4 kb. We then performed LR-PCR and generated a 5-kb fragment that was entirely sequenced. For case 3, we selected four evenly spaced qPCR primer pairs in the distal breakpoint region and eight pairs in the proximal region. By qPCR, we narrowed both regions down to 6–8 kb, and then selected three LR-PCR primers on each side and tested all combinations. The shortest resulting fragment (2.5 Kb) was again entirely sequenced.
Results
Molecular analysis
In the absence of an etiological diagnosis, aCGH was performed to detect submicroscopic imbalances in cases 1 and 3 and to redefine the chromosomal breakpoints in the case reported by Engels et al1 (case 2) with the same array platform (Agilent kit 44B; Figure 3b).
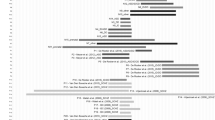
Molecular characterization of the deletions. (a) Map of the 19p13.12–p13.13 region; the deletions are represented by black bars, and the region of overlap between the deletions is shaded in gray. Only the genes discussed in the text are shown. According to the Database of Genomic Variants (projects.tcag.ca/variation), several copy-number variations (CNVs), mostly deletions, are found in the 19p13.12–p13.13 regions. Although some of them (no. 50073, 50074 and 10543) may be extremely rare CNVs, others, including no. 5089 and 5090 in the SRO region, have been detected at very high frequency exclusively in one report using BAC-array analysis,30 and may therefore be somewhat unreliable. (b) Aligned aCGH profile details of the deletions; the deleted regions are shaded in gray. (c) Sequence alignment of the breakpoint junctions; the 3-bp microhomology in case 1 and informative mismatching bases in cases 2 and 3 are shown in bold.
The results were:
Case 1: deletion of 1.9 Mb in chromosomal band 19p13.12 with breakpoints at 14.119–14.135 Mb and 16.053–16.071 Mb.
Case 2 (Engels et al1: case 11/03): deletion of 2.1 Mb in 19p13.12 with breakpoints at 13.965–13.933 Mb and 16.053–16.118 Mb.
Case 3: deletion of 1.5 Mb in 19p13.12–p13.13 with breakpoints at 12.870–12.878 Mb and 14.154–14.166 Mb.
Typing of informative polymorphic chromosome 19 markers D19S432 and D19S558 in the deleted interval in cases 1, 3 and their parents showed that in both patients the deletion was of maternal origin. For case 2, parental samples were not available to assess the origin of the deletion.
Breakpoint cloning and sequencing
The breakpoints characterized by aCGH were further defined by qPCR, amplified by LR-PCR and sequenced in all three cases. The positions and sequences of all junctions are shown in Figure 3c.
In case 1, a 3-bp ATC microduplication, evidence of microhomology-related non-homologous end joining (NHEJ), is present at the junction. The distal breakpoint falls in the 3′-end of the LPHN1 gene, whereas the proximal breakpoint is in intron 2 of TPM4.
In cases 2 and 3, DNA sequence analysis revealed that all breakpoints were located in Alu repeat elements and that both deletions originated by Alu-mediated homologous recombination. In case 3, the distal breakpoint is in intron 3 of the SYCE2 gene, whereas the proximal breakpoint is located between GIPC1 and DNAJB1. In case 2, the distal breakpoint is in intron 2 of the RFX1 gene, and the proximal breakpoint in intron 7 of the TPM4 gene. The proximal breakpoints of cases 1 and 2 are only 15 kb apart. In cases 1 and 2, the rearrangement does not generate any functional fusion gene because the genes involved are in opposite orientation. None of the breakpoints is located within or near segmental duplications.
Discussion
In this study, we report three patients with partially overlapping microdeletions in 19p13.12. Case 1 was analyzed using aCGH because of his abnormal behavior with catastrophic conduct characterized by auto- and hetero-aggressivity.
Case 2 was briefly reported by Engels et al1 and belonged to a cohort of patients selected for aCGH because of unexplained MR. Case 3 was studied for MR and difficult-to-manage hyperactivity.
A comparison of the clinical findings in the three cases is hampered by the fact that the ages of the patients at the times of last evaluation were very different (ie, 7 years, 8412 years and 31 years) and that the oldest patient's history is incomplete for the first months and years of his life. Changes of phenotype with age are well documented in patients with chromosomal imbalances.8 To increase the numbers of patients and to bridge the age gap, we compared the phenotypes of our patients with the recently reported case2 of a 10-year-old girl carrying a 2.52-Mb deletion in 19p13.12–p13.13 overlapping the deleted region in our patients (Figure 3a).
The clinical features common to our patients and to the case reported by Jensen et al2 are moderate-to-severe psychomotor delay, language delay, hearing loss and facial dysmorphisms including brachycephaly, anteverted nares, as well as ear malformations. Mild congenital cardiac anomalies and/or rhythm disturbance were observed at birth in all cases, and resolved spontaneously in the first years of life, with the exception of mild aortic and mitral valve incompetence that was still present at the age of 31 years in case 1 (Table 1).
The four cases share a 358 802-bp SRO in which six annotated genes lie: LPHN1/CIRL1, CD97, DDX39, PKN1, PTGER1 and GIPC1 (Figure 3a).
Two additional recently published cases carry 664 Kb5 and 3 Mb4 deletions of 19p13.13–p13.2 partially overlapping the distal portion (including the SYCE2, NFIX and CACNA1A genes) of the deletion detected in our case 3 (Figure 3a). There is no overlap between these deletions and those in our cases 1 and 2.
LPHN1/CIRL1 encodes the latrophilin-1 precursor/calcium-independent receptor for α-latrotoxin, a member of the latrophilin subfamily of G-protein coupled receptors (GPCR) and may function in both cell adhesion and signal transduction. Interestingly, LPHN1 interacts with the PSD-95/discs large/ZO-1 (PDZ) domain of the multidomain protein family ProSAP/SSTRIP/Shank.9 Shank1 knockout mice show smaller dendritic spines in CA1 pyramidal neurons and weaker synaptic transmission, along with altered learning and memory.10 Haploinsufficiency or point mutations of SHANK3, another member of the SHANK family, results in language delay and/or social communication disorders.11, 12 Deletion of LPHN1 may be potentially responsible for the cognitive and language delay and might contribute to the behavioral phenotype observed in our cases.
The GIPC1 gene, also named synectin (MIM 605072), encodes a scaffolding protein that regulates cell surface receptor expression and trafficking13 and may be involved in G protein-linked signaling. The GIPC1 gene is annotated in the G2C cognition database (human gene ID: G00001513; mouse gene ID: G00000264; http://www.genes2cognition.org). Haploinsufficiency of GIPC1 may also contribute to the mental and language delay in our patients. In addition, the GIPC1 gene may be a good candidate for the hearing impairment observed in all four patients, as its product binds to myosin VI,14 encoded by the MYO6 gene. The C442Y missence mutation in this gene may compromise myosin VI function in a dominant-negative manner and cause autosomal dominant nonsyndromic hearing loss.15 However, the patient described by Lysy et al4 had bilateral hearing loss, despite the fact that his deletion did not extend to the GIPC1 gene. Furthermore, no hearing loss was described in the Gipc1 knockout mouse.16, 17 Thus, the role of GIPC1 with respect to hearing impairment still remains to be clarified.
PKN1 (MIM 601032) codes for a protein belonging to the protein kinase C superfamily and is implicated in a variety of functions in neurons, including cytoskeleton organization.18 The recent demonstration that deregulation of PKN1 activity disrupts neurofilament organization and axonal transport19 suggests that its haploinsufficiency may have an effect on the mental and language delay in our patients.
The PTGER1 gene (MIM 176802) codes for a protein belonging to the G protein-coupled receptor family. Ptger1-null mice show behavioral inhibition, manifested as impulsive aggression with defective social interaction, impaired cliff avoidance and an exaggerated acoustic startle response when under social or environmental stress.20 Haploinsufficiency of this gene might be responsible for the severe behavioral traits in case 1 and the hyperactivity in cases 2 and 3.
On the basis of the current knowledge, deletion of CD97 (MIM 601211) and DDX39 should not have any effect on patients’ phenotype.
Several recent papers have shown that deletion or duplication of the region flanking a gene might also be disease causing.21, 22, 23 Thus, we also considered genes flanking the SRO as potentially responsible for the abnormal phenotypes.
On the proximal side (Figure 3a), SLC1A6 (MIM 600637) codes for a protein that transports L-glutamate, L- and D-aspartate, and seems to act as a symport by cotransporting sodium. It is expressed in the brain and in cell bodies of Purkinje cells. Association studies of polymorphisms in glutamate transporter genes showed that SLC1A6 is a susceptibility locus for schizophrenia.24 SLC1A6 is deleted in cases 1 and 2, and in the case of Jensen et al.2 Its haploinsufficiency could be responsible for cognitive and language delay in all subjects and may contribute to the severe behavioral phenotype in case 1.
On the distal side (Figure 3a), PRKACA (MIM 601639) codes for a catalytic subunit of the cAMP-dependent protein kinase that is required for long-term potentiation (LTP) in neonatal tissue.25 LTP is thought to be critically involved not only in learning and memory, but also in neural circuit formation and refinement. We suggest that haploinsufficiency of PRKACA may be responsible for the cognitive and language delay in cases 2 and 3, and in the subject reported by Jensen et al,2 and possibly also in case 1 if a functionally null allele for PRKACA was created by a position effect or by disruption of regulatory elements.
Mutations in the CACNA1A gene (MIM 601011) have been reported in association with epilepsy and status epilepticus26, 27, 28 and its haploinsufficiency has been suggested to be responsible for epilepsy and infantile spasms in the case of Auvin et al,5 as well as for the abnormal electroencephalography in the case of Lysy et al.4 CACNA1A deletion is most likely responsible for the spontaneous focal seizures in case 3, whereas a positional effect concomitant to antipsychotic treatment might have evoked the epileptic seizure observed in case 1.
NFIX (MIM 164005) has an essential role in the development of the brain and skeleton.29 Nfix-deficient mice show enlargement of the lateral and third brain ventricles and partial agenesis of the corpus callosum; in addition, they show kyphotic deformation of the spine and impaired endochondral ossification in the vertebrae and femur.29 The case of Lysy et al4 showed kyphosis, artogryphosis of lower limb craniosynostosys and moderate ventriculomegaly;4 Auvin et al5 reported advanced bone age. As our case 3 does not show any of these malformations, the phenotype associated with NFIX haploinsufficiency may occur with variable penetrance.
In conclusion, we described two new cases of 19p13.12–p13.13 deletions, refined the breakpoints of another1 and compared the data with three published patients.2, 4, 5 We found that patients with a 358 802-bp common deleted portion at 19p13.12 are characterized by psychomotor and language delay, MR, bilateral sensorineural and/or conductive hearing loss, brachycephaly, anteverted nares and ear malformations, as well as mild congenital cardiac anomalies at birth. Patients with deletion of CACNA1A show epilepsy or epileptiform abnormalities on EEG recording. Several genes within the deletion region may be associated with MR because of their function and expression pattern. The most appealing among them are LPHN1, for its interaction with the SHANK proteins, and PKN1. The GPC1 gene may be a candidate for the hearing impairment observed in all four patients. Case 1 is the most peculiar subject because of his difficult-to-manage aggressive behavioral phenotype, which was most pronounced after adolescence. In cases 2 and 3 hyperactivity was observed. Haploinsufficiency of the PTGER1 gene may be responsible for these behavioral traits.
However, it is also possible that specific phenotypic traits are because of a multigene effect associated with several of the numerous genes included in each deletion.
Additional cases, with different ages and breakpoints, will help to define the phenotype and clarify the role of each deleted gene.
References
Engels H, Brockschmidt A, Hoischen A et al: DNA microarray analysis identifies candidate regions and genes in unexplained mental retardation. Neurology 2007; 68: 743–750.
Jensen DR, Martin DM, Gebarski S et al: A novel chromosome 19p13.12 deletion in a child with multiple congenital anomalies. Am J Med Genet A 2009; 149A: 396–402.
Aten E, den Hollander N, Ruivenkamp C et al: Split hand-foot malformation, tetralogy of Fallot, mental retardation and a 1 Mb 19p deletion-evidence for further heterogeneity? Am J Med Genet A 2009; 149A: 975–981.
Lysy PA, Ravoet M, Wustefeld S et al: A new case of syndromic craniosynostosis with cryptic 19p13.2-p13.13 deletion. Am J Med Genet A 2009; 149A: 2564–2568.
Auvin S, Holder-Espinasse M, Lamblin MD, Andrieux J : Array-CGH detection of a de novo 0.7-Mb deletion in 19p13.13 including CACNA1A associated with mental retardation and epilepsy with infantile spasms. Epilepsia 2009; 50: 2501–2503.
Jerger JF : Clinical experience with impedence audiometry. Arch Otolaryngol 1970; 92: 311–324.
Bonaglia MC, Giorda R, Massagli A, Galluzzi R, Ciccone R, Zuffardi O : A familial inverted duplication/deletion of 2p25.1-25.3 provides new clues on the genesis of inverted duplications. Eur J Hum Genet 2009; 17: 179–186.
Van Buggenhout GJ, Pijkels E, Holvoet M, Schaap C, Hamel BC, Fryns JP : Cri du chat syndrome: changing phenotype in older patients. Am J Med Genet 2000; 90: 203–215.
Kreienkamp HJ, Zitzer H, Gundelfinger ED, Richter D, Bockers TM : The calcium-independent receptor for alpha-latrotoxin from human and rodent brains interacts with members of the ProSAP/SSTRIP/Shank family of multidomain proteins. J Biol Chem 2000; 275: 32387–32390.
Hung AY, Futai K, Sala C et al: Smaller dendritic spines, weaker synaptic transmission, but enhanced spatial learning in mice lacking Shank1. J Neurosci 2008; 28: 1697–1708.
Bonaglia MC, Giorda R, Mani E et al: Identification of a recurrent breakpoint within the SHANK3 gene in the 22q13.3 deletion syndrome. J Med Genet 2006; 43: 822–828.
Durand CM, Betancur C, Boeckers TM et al: Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 2007; 39: 25–27.
Lee NY, Ray B, How T, Blobe GC : Endoglin promotes transforming growth factor beta-mediated Smad 1/5/8 signaling and inhibits endothelial cell migration through its association with GIPC. J Biol Chem 2008; 283: 32527–32533.
Aschenbrenner L, Lee T, Hasson T : Myo6 facilitates the translocation of endocytic vesicles from cell peripheries. Mol Biol Cell 2003; 14: 2728–2743.
Melchionda S, Ahituv N, Bisceglia L et al: MYO6, the human homologue of the gene responsible for deafness in Snell's waltzer mice, is mutated in autosomal dominant nonsyndromic hearing loss. Am J Hum Genet 2001; 69: 635–640.
Chittenden TW, Claes F, Lanahan AA et al: Selective regulation of arterial branching morphogenesis by synectin. Dev Cell 2006; 10: 783–795.
Naccache SN, Hasson T, Horowitz A : Binding of internalized receptors to the PDZ domain of GIPC/synectin recruits myosin VI to endocytic vesicles. Proc Natl Acad Sci USA 2006; 103: 12735–12740.
Mukai H : The structure and function of PKN, a protein kinase having a catalytic domain homologous to that of PKC. J Biochem 2003; 133: 17–27.
Manser C, Stevenson A, Banner S et al: Deregulation of PKN1 activity disrupts neurofilament organisation and axonal transport. FEBS Lett 2008; 582: 2303–2308.
Matsuoka Y, Furuyashiki T, Yamada K et al: Prostaglandin E receptor EP1 controls impulsive behavior under stress. Proc Natl Acad Sci USA 2005; 102: 16066–16071.
Benko S, Fantes JA, Amiel J et al: Highly conserved non-coding elements on either side of SOX9 associated with Pierre Robin sequence. Nat Genet 2009; 41: 359–364.
Dathe K, Kjaer KW, Brehm A et al: Duplications involving a conserved regulatory element downstream of BMP2 are associated with brachydactyly type A2. Am J Hum Genet 2009; 84: 483–492.
Kurth I, Klopocki E, Stricker S et al: Duplications of noncoding elements 5′ of SOX9 are associated with brachydactyly-anonychia. Nat Genet 2009; 41: 862–863.
Deng X, Shibata H, Takeuchi N et al: Association study of polymorphisms in the glutamate transporter genes SLC1A1, SLC1A3, and SLC1A6 with schizophrenia. Am J Med Genet B Neuropsychiatr Genet 2007; 144B: 271–278.
Yasuda H, Barth AL, Stellwagen D, Malenka RC : A developmental switch in the signaling cascades for LTP induction. Nat Neurosci 2003; 6: 15–16.
Jouvenceau A, Eunson LH, Spauschus A et al: Human epilepsy associated with dysfunction of the brain P/Q-type calcium channel. Lancet 2001; 358: 801–807.
Beauvais K, Cavé-Riant F, De Barace C et al: New CACNA1A gene mutation in a case of familial hemiplegic migraine with status epilepticus. Eur Neurol 2004; 52: 58–61.
Kors EE, Melberg A, Vanmolkot KR et al: Childhood epilepsy, familial hemiplegic migraine, cerebellar ataxia, and a new CACNA1A mutation. Neurology 2004; 63: 1136–1137.
Driller K, Pagenstecher A, Uhl M et al: Nuclear factor I X deficiency causes brain malformation and severe skeletal defects. Mol Cell Biol 2007; 27: 3855–3867.
Wong KK, deLeeuw RJ, Dosanjh NS et al: A comprehensive analysis of common copy-number variations in the human genome. Am J Hum Genet 2007; 80: 91–104.
Acknowledgements
We thank the families for their cooperation. This work was supported by Italian Ministry of Health grant RF-AOM-2007-636538 to OZ.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Bonaglia, M., Marelli, S., Novara, F. et al. Genotype–phenotype relationship in three cases with overlapping 19p13.12 microdeletions. Eur J Hum Genet 18, 1302–1309 (2010). https://doi.org/10.1038/ejhg.2010.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.115
Keywords
This article is cited by
-
Adhesion G protein-coupled receptor gluing action guides tissue development and disease
Journal of Molecular Medicine (2022)
-
Chromosomal microarray analysis in the genetic evaluation of 279 patients with syndromic obesity
Molecular Cytogenetics (2018)
-
Transcriptome profiling analysis reveals the role of latrophilin in controlling development, reproduction and insecticide susceptibility in Tribolium castaneum
Genetica (2018)
-
Malan syndrome: Sotos-like overgrowth with de novo NFIX sequence variants and deletions in six new patients and a review of the literature
European Journal of Human Genetics (2015)
-
A new case of de novo 19p13.2p13.12 deletion in a girl with overgrowth and severe developmental delay
Molecular Cytogenetics (2014)