Abstract
p53 is an important tumor suppressor and stress response mediator. Proper control of p53 level and activity is tightly associated with its function. Posttranslational modifications and the interactions with Mdm2 and Mdm4 are major mechanisms controlling p53 activity and stability. As p53 protein is short-lived and hardly detectable in unstressed situations, less is known on its basal level expression and the corresponding controlling mechanisms in vivo. In addition, it also remains obscure how p53 expression might contribute to its functional regulation. In this study, we established bacterial artificial chromosome transgenic E.coli β-galactosidase Z gene reporter mice to monitor p53 expression in mouse tissues and identify important regulatory elements critical for the expression in vivo. We revealed preferentially high level of p53 reporter expressions in the proliferating, but not the differentiated compartments of the majority of tissues during development and tissue homeostasis. In addition, tumors as well as regenerating tissues in the p53 reporter mice also expressed high level of β-gal. Furthermore, both the enhancer box sequence (CANNTG) in the p53 promoter and the 3′ terminal untranslated region element were critical in mediating the high-level expression of the reporter. We also provided evidence that cellular myelocytomatosis oncogene was a critical player regulating p53 mRNA expression in proliferating cells and tissues. Finally, we found robust p53 activation preferentially in the proliferating compartment of mouse tissues upon DNA damage and the proliferating cells exhibited an enhanced p53 response as compared with cells in a quiescent state. Together, these results suggested a highly regulated expression pattern of p53 in the proliferating compartment controlled by both transcriptional and posttranscriptional mechanisms, and such regulated p53 expression may impose functional significance upon stress by setting up a precautionary mode in defense of cellular transformation and tumorigenesis.
Similar content being viewed by others
Main
p53, encoded by p53 (also termed as TP53 gene or transformation related protein 53 gene (Trp53)), is one of the most important tumor suppressors. Constant degradation in normal embryos or tissues mediated by the E3-ligase activity of Mdm2 renders p53 protein difficult to be detected,1 whereas Mdm4 mainly inhibits the transcriptional activity of p53.2 Repression of these two inhibitors by stress stimuli is the major posttranslational regulatory mechanism controlling p53 activity and stability.3 Once activated, p53 is able to mediate a plethora of responses including inhibition of cellular proliferation important for tumor suppression.
A number of studies demonstrated the importance of p53 gene dosage and expression level in affecting its activity and the cellular responses. Apoptosis increased with p53 levels in cultured cells.4 In mice, additional 1–2 copies of wild type p53 did not impact mouse development, growth and senescence, but significantly enhanced the sensitivity to γ-irradiation and strengthened the resistance to tumorigenesis.5 On the other hand, p53 haploinsufficiency was observed in a variety of situations including tumorigenesis and stress responses. Both individuals harboring a p53 germline mutation in the Li–Fraumeni families and p53 heterozygous mice exhibited increased incidence of tumors, some of which apparently did not lose the wild-type allele. In some instances, p53 heterozygosity was able to rescue the lethal phenotype caused by deletion of Mdm26 or Mdm4,7 again demonstrating a dosage dependent phenotype. In relevance to human breast cancer, methylations in CG rich region of p53 promoter impaired its transcription and protein levels,8 and impaired expression of the transcriptional factor HOXA5 also decreased p53 mRNA level in the tumor tissues.9 All these implied the importance of p53 expression and its regulatory mechanisms for stress responses and tumor suppression.
Early studies indicated a ubiquitous tissue expression pattern of p53 mRNA,10 consistent with p53’s role as a tumor suppressor of tissues from different origins. However, the studies on distribution of p53 mRNA in sub-tissue compartments have been scarce and scattered. In situ hybridization found high-level p53 mRNA expression during embryonic development11 and a striking p53 mRNA expression pattern in postnatal rat brain, with intensive signals in subventricular zone, rostral migratory stream and external granular layer (EGL), where new neurons were produced.12 In NIH3T3 cells, p53 mRNA level fluctuated with cell cycle progression.13, 14 F9 cells, an embryonic carcinoma stem cell line, expressed high levels of p53 mRNA, which dropped to low levels after induction of differentiation.15, 16 Meanwhile, many efforts focused on identifying and analyzing the cis-elements and transacting factors important for p53 expression. Cellular myelocytomatosis oncogene (c-Myc),17 NF-kappaB,18, 19 E2F1,20 C/EBP beta,21 EGR-1,22 and Ets-1/223 were among the many transcription factors shown to regulate p53 promoter activity. Other studies also addressed the posttranscriptional regulatory mechanisms for p53 expression. Cis-elements including AU rich element, CPE (cytoplasmic polyadenylation element) and micro RNA-binding sites were found present in p53 3′ terminal untranslated region (3′UTR) and regulated reporter p53 mRNA stability and/or translational efficiency in either a positive or negative manner. A recent study also found that ectopic human p53 5′UTR and 3′UTR in H1299 cells could bind each other through paired bases and strengthened translational efficiency of the reporter transcript upon DNA damage.24 However, the in vivo relevance, contribution and concerted nature of these regulations on p53 expression remain incompletely understood.
Proliferation is the fundamental cellular process closely linked to development, homeostasis and cancer. There are evidence suggesting that fast proliferating cells are more sensitive to stresses and p53 activation. Conventional radiotherapy and chemotherapy often lead to severe side effects in cancer patients mainly affecting fast renewing tissues. Intriguingly, loss of Mdm2 in a p53 hypomorphic background resulted in p53 stabilization preferentially in the proliferating compartments of the postnatal mice, leaving important clues about p53 basal expression.6 More works are needed to decipher or distinguish the underlying mechanisms influencing the p53 regulatory disparities.
In this study, we established bacterial artificial chromosome (BAC) transgenic reporter mice to model basal p53 expression in intact tissues during development, homeostasis, regeneration and tumorigenesis. We also evaluated the role of p53 promoter and 3′UTR elements in governing p53 expression patterns. Our results clearly demonstrated a preferential high-level expression of p53 reporter in the proliferating compartments of multiple tissues as dictated by the enhancer box sequence (E-box) and 3′UTR regulatory elements. Importantly, we provided evidence that the regulation of basal level expression was closely correlated with the robustness of p53 response under stress conditions.
Results
Generation of BAC transgenic reporter mice in monitoring p53 expression
To probe p53 mRNA expression pattern in mouse tissues, we performed in situ hybridization (ISH) in skins of postnatal day 1 mice, cerebellum of P7 mice and small intestine of mice at 2 months of age. p53 mRNA was at high levels in the hair follicle and basal layer of the skin, EGL of the cerebellum and intestinal crypts (Figure 1a), which are actively proliferating; in contrast, p53 mRNAs were at much lower levels in supra basal layer of the skin, internal granular layer (IGL) of the cerebellum and villus of the small intestine (Figure 1a), which belong to the differentiated compartments.
Generation and validation of BAC transgenic reporter mice to monitor p53 expression. (a) In situ hybridization of p53 mRNA using sense and antisense probes on postnatal day 1 mouse skin, P7 cerebellum and 2-month-old mouse intestine. Boxed areas in left panel were shown in the middle panel. Crypts in intestine were also shown in the middle bottom panel. Scale bar=100 μm. (b) Schematic representations of the p53 gene locus in the BAC and its modifications with the insertions of lacZ reporter in the ATG (exon 2) of full length p53 and EGFP reporter in the ATG (exon 5) of an isoform (Δ157) together with alterations in the regulatory elements in three BAC transgenic lines. ChIP primers used for detecting wild-type E-box-binding in tissues and cells (F1, R1 or F2, and WTR2); ChIP primers for detecting mutant E-box-binding in tissues and cells (F2, MutantR2). (c) β-gal staining of E10.5 mouse embryos from PZU, PMZU and PZS transgenic reporter lines with 1–4 BAC copies. Embryos were whole-mount stained in 1 mg/ml X-gal staining solution with gentle shaking for 24 h at 4 °C. Scale bar=400 μm. (d) A comparison of p53 reporter expression in p7 skin, p7 cerebellum and 2-month-old mouse small intestine by β-gal staining with p53 mRNA in situ hybridization. Scale bar=50 μm
To study p53 gene expression pattern more systematically, and to determine the regulatory mechanisms critical for basal p53 expression in vivo, we established BAC transgenic mice in which a E.coli β-galactosidase Z gene (LacZ) reporter gene (with nuclear localization signal) followed by the p53 3′UTR and simian vacuolating virus 40 polyadenylation sequences (SV40pA) was inserted immediately after the start codon (ATG) in the exon 2 of the BAC p53 gene locus. In addition, we disrupted a reported p53Δ157 isoform with start codon in p53 exon 525, 26 with an enhanced green fluorescence protein (EGFP)-human growth hormone polyadenylation sequences cassette in the same transgenic construct. No obvious EGFP expression was detected, and thus was not further investigated in this study. This transgenic model was designated as p53PZU (or PZU) to monitor basal p53 expression by β-galactosidase (β-gal) activity in vivo (Figure 1b). To study the transcriptional control of p53 expression, we focused on a previously identified conserved E-box-binding motif close to the transcriptional start site in p53 promoter.17 Chromatin immunoprecipitation (ChIP) analysis demonstrated a direct binding of c-Myc on this E-box sequence in proliferating cells but not in quiescent cells (Supplementary Figure 1). To address the possible regulatory role of the E-box in the p53 promoter, p53PMZU (or PMZU) transgenic mice were similarly established as p53PZU except for a mutated E-box in the BAC p53 promoter from CACGTG to GGTGTG (Figure 1b). To assess the importance of p53 3′UTR in the regulation of basal p53 expression, we also established p53PZS(or PZS) BAC transgenic mice in which the inserted LacZ reporter gene was followed only by the SV40pA to monitor the expression of p53 without its own 3′UTR sequence (Figure 1b). With X-gal incubation, all five lines of PZU embryos exhibited clearly visible β-gal staining, the level of which correlated with their corresponding transgenic BAC copy numbers (Figure 1c). Notably, all the embryos from transgenic lines of PMZU and PZS, including the high copy number lines, exhibited severely diminished β-gal staining (Figure 1c). Comparisons of LacZ expressions in the skin, cerebellum and small intestine of the PZU mice with the p53 mRNA ISH revealed highly consistent patterns (Figure 1d), validating our transgenic approach in monitoring endogenous p53 mRNA expression.
High level of p53-LacZ reporter expression in the proliferating compartments of mouse tissues
To reveal p53 expression pattern and levels in mouse tissues, we performed X-gal staining in tissues of PZU mice at postnatal day 7–8, 2–3 months and 13–15 months of age, respectively. Intense β-gal staining was observed in multiple tissue compartments including germinal center of spleen, spermatogenous cells of seminiferous tubule in testis, dentate gyrus (DG) of hippocampus, islet of postnatal and adult pancreas, ependymocytes of choroid plexus, EGL of the postnatal cerebellum, renal tubule and glomerulus, and pulmonary alveoli (Figure 2a). In contrast, β-gal was expressed at low or undetectable levels in the differentiated cells toward the center of the seminiferous tubule, mature neurons outside the DG region in adult hippocampus, acinous cells in adult pancreas and the IGL of cerebellum (Figure 2a), which were all non-proliferating in nature. Meanwhile, direct comparisons in the same tissues revealed that β-gal-staining intensities decreased when mice aged, especially in spleen, choroid plexus, kidney and lung (Figure 2a). Further, the whole mount X-gal staining on de-skinned mice also demonstrated attenuated β-gal expression in mouse muscle and skeleton with age (Supplementary Figure 2a). As few exceptions, the non-proliferating hepatocytes in the adult liver and purkinje cells of the cerebellum also demonstrated detectable levels of β-gal staining in the PZU mice (Figures 2a and d).
Preferential p53 reporter expression in the proliferating compartments during mouse postnatal development, tissue homeostasis, regeneration and tumorigenesis. (a) β-gal staining of tissues including spleen, testis, dentate gyrus, pancreas, choroid plexus, cerebellum, kidney, lung and liver at P7, 2 months and 15 months of age respectively in PZU mice. Frozen slices of tissues were stained in 1 mg/ml X-gal staining solution at 37 °C for 10 h, except for 3 h in testis. Scale bar=200 μm. (b) β-gal staining of skins at the anagen (p20) or telogen (p24) phase of the hair cycle in PZU mice. Frozen slices of skins were stained in 1 mg/ml X-gal staining solution at 37 °C for 10 h. Boxed areas were shown in the right panel. Scale bar=200 μm. (c) Double immunofluorescence of β-gal and Ki67 in p8 cerebellum and adult small intestine of PZU mice. Scale bar=200 μm. (d) β-gal staining of liver before and 48 h after hepatotectomy. Frozen slices of livers were stained in 1 mg/ml X-gal staining solution at 37 °C for 7 h in PZU mice. Boxed areas were shown in the right panel. Scale bar=200μm. (e) β-gal staining of small intestine in PZU mice and polyps in PZU; Apcmin mice
The preferential expressions of p53-LacZ reporter in proliferating compartment of tissues were further evaluated in the hair follicles of mouse skin which undergo step-wise growing and quiescent cycles. There were abundant strong β-gal-staining positive cells at the anagen stage of the hair cycle at P20, but only few at the quiescent telogen stage at P24 (Figure 2b). To study the link of proliferating or quiescent state with p53-LacZ expression, we performed X-gal staining in PZU MEFs either at proliferating or quiescent state. Upon withdraw of serum, proliferating MEFs gradually moved to a quiescent state. In parallel, PZU MEFs with high β-gal-staining intensity dropped from 46.2 to 2.6%, whereas cells with low-level β-gal increased from 18.7 to 68.3% (Supplementary Figure 2b). Next, we performed double immunofluorescence of ki-67 and β-gal in cerebellum and small intestine of the p7 PZU mice. β-gal positive cells were largely overlapped with those of Ki67 positive proliferating cells, establishing a direct link between high p53-LacZ expression and cellular proliferating state in multiple tissue compartments (Figure 2c).
Similar to the self-renewal of homeostatic tissues, tissue repair and regeneration also features cellular proliferation. β-gal expression was examined in the liver of PZU mice under hepatotectomy. In spite of the detectable basal level of β-gal staining, an enhanced β-gal expression was observed on the proliferation peak at 48 h (Figure 2d) as demonstrated by Ki67 staining (Supplementary Figure 2c), paralleling the level of endogenous p53 mRNA expression (Supplementary Figure 2d).
Tumorigenesis is marked by uncontrolled proliferation. To study p53 reporter expression during tumorigenesis, we crossed the PZU mice with Apcmin mice with a mutation in the Apc gene and studied β-gal expression in the adenomas developed in the small intestine of the PZU; Apcmin/+ mice. Most cells in these tumors expressed high levels of β-gal (Figure 2e), which paralleled with Ki67 immunostaining (Supplementary Figure 2e), whereas high levels of β-gal was only confined to the crypts in normal intestine (Figure 2e).
Collectively, high level of β-gal expression was specifically observed in the proliferating cell compartments in a variety of homeostatic or pathological contexts in the PZU mice, suggesting p53 expression is highly regulated in the proliferating compartments.
p53 reporter expression is controlled by both the p53 promoter E-box element and 3′UTR
With the E-box-mutated PMZU mice and 3′UTR-deleted PZS mice, we addressed the respective roles of the conserved E-box and the p53 3′UTR sequence on the enhanced reporter expression in the proliferating cells during development and homeostasis. Significantly diminished β-gal staining was observed in the PMZU embryos as compared with the PZU embryos with the same BAC copy number (Figure 1c). At both P7-8 and 2–3 months of age, β-gal staining was all reduced in the PMZU mice as compared with the PZU mice in a panel of tissues examined, including spleen, testis, hippocampus, pancreas, choroid plexus, postnatal cerebellum, skin, lung and liver (Figures 3a and b). Immunohistochemistry (IHC) with β-gal antibody also revealed decreased β-gal expressions in the small intestine and bone marrow of the PMZU mice, whereas IHC of Ki67 remained the same between the PMZU and PZU mice (Figures 3c and d). Consistently, in the PMZU mice, the LacZ mRNA level normalized to that of endogenous p53 also dropped by 50–73% in embryos, postnatal and adult tissues as compared with those of the PZU mice (Figures 4a and b). These results indicated an important role of the E-box on p53-LacZ reporter expression in vivo.
The E-box element in p53 promoter was critical in driving high level of p53 reporter expression in proliferating cell compartments. (a and b) β-gal staining of tissues at P7 (a) and 2 months (b) of age in PZU and PMZU mice. Scale bar=200 μm. (c and d) IHC of Ki67 and β-gal on intestine and bone marrow from P7 (c) and 2-month-old (d) PZU and PMZU mice. Scale bar=200 μm
Disrupting c-Myc binding on the conserved E-box in p53 promoter downregulated p53 expression. (a) RT-PCR results of LacZ mRNA expression as normalized to p53 mRNA in embryos at 10.5 day from PZU and PMZU mice. (b) RT-PCR results of LacZ mRNA expression as normalized to p53 mRNA in small intestine, skin and kidney from PZU and PMZU mice. (c) RT-PCR results of lacZ mRNA expression normalized to 18 s RNA in a time course upon serum stimulation in MEFs from PZU and PMZU mice with the same BAC copy number. (d) ChIP analysis on c-Myc binding on the endogenous p53 promoter (wild-type E-box) or p53 promoter in transgenic BAC (mutant E-box) in the E11.5 embryo of the PMZU mice. (e) ChIP analysis on c-Myc and USF binding on the endogenous p53 promoter (wild type E-box) or p53 promoter in transgenic BAC (mutant E-box) in small intestine and liver of 2-month-old PMZU mice. (f) ChIP analysis on c-Myc binding on the endogenous p53 promoter (wild-type E-box) or p53 promoter in transgenic BAC (mutant E-box) in the MEFs from the PMZU mice under the treatment of Myc inhibitor 10058-F4 for 6 h (upper panel) and ChIP analysis on c-Myc binding on the p53 promoter (wild-type E-box) in the MEFs from the PZU mice under the treatment of Myc inhibitor 10058-F4 for 6 h (lower panel). (g)RT-PCR results of LacZ and p53 mRNA expressions normalized to β-actin in MEFs from PZU and PMZU mice under the treatment of 10058-F4 for 8 h. (h) β-gal staining of MEFs from the PZU and PMZU mice under the treatment of 10058-F4 for 10 h. MEFs were stained in 1 mg/ml X-gal staining solution at 37 °C for 10 h. Scale bar=200 μm. Values are means±S.E.M.s. *P<0.05; **P<0.01; ***P<0.001 (t-test)
Cell cycle exit, reentry and progression can be manipulated in cell culture systems. p53 mRNA was elevated 8 h after serum stimulation and reached its peak at 16 h, paralleling cell cycle progression13, 14 (Supplementary Figure 3). LacZ mRNA from PZU MEFs also exhibited a similar elevation upon serum stimulation, whereas LacZ mRNA level from the PMZU mice remained unchanged (Figure 4c), again suggesting a role of E-box in coordinating cell cycle progression and p53 expression. c-Myc is a major Leucine Zipper-bHLH transcription factor important in promoting proliferation and cell cycle progression in many types of cells. To study whether the reduction of β-gal expression in proliferating tissues of PMZU mice is due to the loss of binding of c-Myc or other transcriptional factors to the mutant E-box, ChIP assay was performed in developing embryos and adult intestine, as well as the non-proliferating adult liver in the PMZU mice containing both the wild-type E-box from the endogenous p53 locus and the mutant E-box from the transgenic reporter. As expected, c-Myc could bind wild-type E-box in the endogenous p53 promoter, but not the mutant E-box in transgenic p53 promoter in the proliferating tissues of the PMZU mice (Figures 4d and e). However, little c-Myc binding was observed on endogenous p53 promoter in liver (Figure 4e). In contrast, upstream stimulatory factor-1 (USF-1), another bHLH transcription factor, could bind the E-box in endogenous p53 promoter but not the mutant E-box in liver of the PMZU mice (Figure 4e). Notably, no USF-1 binding to endogenous p53 promoter in small intestine was observed (Figure 4e). These results suggested a differential binding of transcription factors to the p53 E-box element in tissues. To reveal the functional significance of c-Myc binding in regulating p53 expression, specific Myc–Max dimerization inhibitor 10058-F4 was used to treat proliferating MEFs. 10058-F4 significantly impaired c-Myc binding on the wild-type E-box in both PMZU and PZU MEFs (Figure 4f) and decreased both LacZ mRNA expression and β-gal staining in PZU MEFs (Figures 4g and h). Furthermore, although 10058-F4 treatment also resulted in decreased endogenous p53 mRNA levels in both PZU and PMZU MEFs, it did not further influence the levels of LacZ mRNA and β-gal staining in PMZU MEFs (Figures 4g and h). Together, these results indicated a significant role of c-Myc in upregulating p53 reporter expression through the E-box element, suggesting that c-Myc may be one of the major transcriptional factors driving high level of p53 mRNA expression in proliferating cells.
Compared with PZU mice, PZS mice lacking p53 3′UTR exhibited no or very low-level β-gal expression visible only in limb bud of the E10.5 embryos with 1–4 BAC copies (Figure 1c). In addition, no or very weak β-gal staining was detected in multiple tissue compartments of the PZS mice with 2 or 4 BAC copies at both P7 (Figure 5a) and 2 months of age (Figure 5b). Only the PZS mice with 4 BAC copies exhibited weak β-gal staining in the spermatogenous cells of somniferous tubules in testis and dentate gyrus of hippocampus (Figures 5a and b). These results indicated an essential role of p53 3′UTR in supporting high level of β-gal expressions in the proliferating tissue compartments. Interestingly, in both embryos and a variety of adult tissues examined, LacZ mRNA levels of the PZS mice were not significantly different from those of the PZU mice (Supplementary Figure 4), indicating that p53 3′UTR does not significantly influence LacZ mRNA levels.
p53 3′UTR was essential in supporting high level of p53 reporter expression. (a) β-gal staining of multiple tissues including spleen, testis, dentate gyrus, pancreas, choroid plexus, cerebellum, skin, kidney, lung and liver in P7 PZU and PZS mice. Slices of frozen tissues were stained in 1 mg/ml X-gal staining solution at 37 °C for 10 h, except for 3 h for testis. Scale bar=200 μm. (b) β-gal staining of multiple tissues including spleen, testis, dentate gyrus, pancreas, choroid plexus, cerebellum, skin, kidney, lung and liver in 2-month-old PZU and PZS mice. Slices of frozen tissues were stained in 1 mg/ml X-gal staining solution at 37 °C for 10 h, except for 3 h for testis. Scale bar=200 μm. (c)Western blot analysis of β-gal expression in MEFs from PZU, PMZU and PZS mice with the two copies of transgenic BAC
Finally, we directly compared the β-gal protein level in MEFs from the PZU, PMZU and PZS mice with two BAC copies. Western blot analysis showed that β-gal level dropped by >50% in PMZU MEF, and was almost undetectable in PZS MEFs (Figure 5c), consistent with the β-gal staining results in multiple tissues. These results further supported that p53 E-box and 3′UTR were critical in sustaining high-level expression of the reporter gene.
A functional link between cellular proliferating state and p53 response
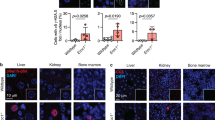
Our findings that p53 reporter expression was preferentially upregulated in proliferating cells suggested the possibility of a more robust p53 response upon stresses in proliferating cells compared with quiescent and differentiated cells. To directly explore the possible regulatory difference of p53 in vivo, B6 mice was either untreated or treated with 6 Gray of X-irradiation and immunostained for p53 protein in multiple tissue compartments. Consistent with previous results, p53 protein was undetectable in normal tissues from untreated mice (Figure 6a). In contrast, upon irradiation, p53 positive cells were readily detected in EGL of cerebellum in the postnatal mice, and in the basal cells of tongue and intestine crypt of both postnatal and adult mice (Figure 6a). In contrast, there was weak or no p53 staining in the more differentiated cell compartments of these tissues, such as IGL of the adult cerebellum, villi of small intestines and differentiated cells in the tongue epithelium (Figure 6a). Double staining with p53 and 5-Bromo-2-Deoxy Uridine (Brdu) antibodies further revealed that p53-positive-staining largely overlapped with that of Brdu in proliferating compartment of fast turn-over tissues including small intestine and testis (Figure 6b). Therefore, p53 protein was preferentially expressed in the proliferating tissue compartment upon induction, essentially recapitulating its gene expression pattern.
A direct link between cellular proliferation and p53-mediated stress response. (a) IHC of p53 in small intestine, cerebellum and tongue of p7 and 2-month-old C57BL/6J control mice or mice treated with X-irradiation for 4 h. Arrows depicted positive stained cells. Scale bar=200 μm. (b) Double immunostaining of p53 and BrdU in intestine and testis of 2-month-old C57BL/6J mice treated with X-irradiation for 4 h. (c) Western blot analysis of p53 and p21 expression on quiescent and proliferating 3T3 cells treated with Doxorubicin. (d) Western blot analysis of p53 and p21 expression on quiescent and proliferating MEFs treated with Cisplatin. (e) Immunofluorescence of p53 or p21 in quiescent and proliferating MEFs treated with Cisplatin or Doxorubicin. Scale bar=200 μm
To directly analyze p53 functional differences in cells at different proliferating state, quiescent and proliferating cells were treated with either Doxorubicin or Cisplatin. p53 protein was undetectable regardless of the cell cycle status in untreated controls (Figures 6c and d). However, upon treatment, p53 as well as its transcriptional target gene p21 was much more greatly elevated in the cycling cells (Figures 6c and d). Immunofluorescence on MEFs treated with Cisplatin also demonstrated much more elevated level of p53 activation and response in the cycling cells (Figure 6e). Thus, the concordant elevation in p53 expression may provide one possible functional basis for robust p53 responses selectively in the proliferating compartments upon stress.
Discussion
The regulatory and functional mechanisms of p53 have been under intensive investigations at molecular, cellular and organismal levels to better understand its roles in tumor suppression and stress responses. Aiming to study p53 expression pattern and regulatory mechanisms both under physiological contexts and in a comprehensive manner, we established an in vivo reporter system in BAC transgenic mice that were able to recapitulate endogenous p53 mRNA expression and identified a highly selective expression pattern for the reporter across diverse tissue compartments in both physiological and pathological conditions. We provided further evidence suggesting that the preferential expression of the reporter in proliferating compartments is critically dependent on both p53 promoter and 3′UTR elements through transcriptional or posttranscriptional mechanisms. In addition, this study also revealed a distinctive functional difference of p53 in proliferating compartments versus their differentiated or quiescent counterparts upon stress.
Previous studies on p53 mRNA expression and promoter analyses provided a number of relatively separated clues on p53 expression regulation in general and a few of them were implicated in proliferating cells. However, to our knowledge, there is no commonly recognized theme for p53 expression pattern in vivo and the importance and relevance remained to be further established for many of the cis-elements and transacting factors identified in regulating p53 expression. Here we took a transgenic approach in integrating different lines of studies and shed new insights on p53 expression and its regulatory mechanisms in a diverse set of conditions in vivo. Our results corroborated the existing results and argued strongly for a unified and distinctive expression pattern for p53 in vivo.
It becomes increasingly apparent that the studies on the regulatory mechanisms of gene expression can be better oriented and more meaningful with a clear understanding of the expression pattern of the gene. As an immediate early gene, c-Myc is induced earlier than p53 mRNA accumulation17, 27 and its mRNA level parallels with p53 mRNA level in cell lines.14, 28 Ectopic c-Myc or N-Myc was able to trans-activate reporters driven by an E-box from the p53 promoter in 3T3 and neuroblastoma cells.17, 29 In an effort to decipher the requirements of p53 expression in vivo, we found the E-box element in the p53 promoter and its binding by c-Myc contributed significantly to high-level expression of p53 reporters in the proliferating compartments. Notably, as regulators of cell cycle or proliferation, E2F1,20 C/EBP beta,21 EGR-122 and Ets-1/223 were also implicated in regulating p53 expression. In PMZU mice with high BAC copy numbers, weak β-gal staining was still detected in the proliferating compartments of tissues. In addition, the variations in the degree of reduction of lacZ expression in the PMZU mice of different ages also suggested a differential dependence on the E-box regulation. Thus, the strength of Myc activity, together with the contribution of other factors may help to coordinate p53 expression levels with the cellular proliferation state in multiple tissue compartments.
In spite of the dominant p53 reporter expression pattern identified in the vast majority of tissues examined, there were a few exceptions including the purkinje cells and hepatocytes with detectable levels of β-gal staining while being largely quiescent. Interestingly, the E-box mutation also abrogated the reporter expression in liver. Our ChIP analysis suggested that the moderate reporter expression in liver may be regulated by USF-1, which is known to be expressed in liver.30, 31 When the hepatocytes reentered the cell cycle, a strong increase in β-gal expression suggested that these cells were still subjected to the regulatory mechanisms distinguishing proliferating and quiescent states.
Replacement of p53 3′UTR with a commonly used SV40pA greatly reduced p53 reporter expression in the proliferating compartments, suggesting a general role of the 3′UTR as a positive regulatory elements in supporting p53 expression. RNA-binding factors such as HuR,32 Wig1,33 CPEB,34 and certain miRNAs were reported to regulate mRNA stability and/or translational efficiency by directly binding on cis-elements in p53 3′UTR.25, 26, 35 Although HuR and Wig1 could stabilize reporter mRNA and/or strengthen its translational efficiency upon DNA damage stress, CPEB did not significantly influence p53 mRNA level, but instead regulated the p53 mRNA polyA length and protein level through translation.34 In our study, reporter mRNA levels were not significantly altered in the PZS mice, thus pointing to the possible translational control in mediating the high-level expression mediated by the 3′UTR sequences.
Our results suggested concerted effects of both transcriptional and posttranscriptional regulations in supporting the selective p53 expression pattern in proliferating compartments. Known to be anti-proliferative, p53 protein is kept to undetectable levels in normal cells. However, this could also obscure many possible links between p53 expression and its function. For example, proliferative tissues are often more radiosensitive,36, 37 and while complex mechanisms may underlie such phenomenon, heightened p53 basal expression may have a role. Through genetic deletion of Mdm2 in a p53 hypomorphic background, we previously discovered that p53 protein stabilization and accumulation only appeared in the proliferating compartments of mice,6 a pattern fully recapitulated in the p53 reporter mice in this study. Here we demonstrated that DNA damage elicited a greater p53 response in the proliferating cells. Therefore, the posttranslational controls seem to act possibly in sequence with the transcriptional and posttranscriptional mechanisms in fine-tuning p53 activity in a variety of cell types depending on their proliferative and differentiation status.
In summary, this study took a genetic approach to address the long existing question of p53 expression and revealed novel insights suggesting a general intrinsic mechanism for upregulating p53 in proliferating cells and tissues. Linking to p53 function, the potential advantage and significance for such regulation at the expression level can be several folds: first, higher basal level of p53 would allow a fast and robust protein stabilization/activation and stress response upon stress; on the other hand, a retarded or weak p53 response may protect the cells from apoptosis in the terminally differentiated cells and reduce tissue damage; finally, the switches of cell proliferative state or fate constantly occur for stem cells and progenitors during homeostasis. The coordinated control of p53 expression may allow the uninterrupted monitoring of proliferation to fulfill its tumor suppressor functions.
As one of the most fundamental cellular processes, cellular proliferation and its regulation are an integral part of development, tissue homeostasis, tissue regeneration and tumorigenesis. Close monitoring of this process by a tumor suppressor controlled at multiple levels may set up a precautionary mode in the cells without causing much disturbance.
Materials and Methods
Mice
Mice were bred and maintained under specific pathogen-free conditions and experiments were conducted in accordance with the Institutional Animal Care and Use Committee at the animal facility of Model Animal Research Center of Nanjing University, China. p53PZU (PZU), p53PMZU(PMZU) and p53PZS(PZS) mice were established on C57BL/6J and CBA mixed background and were backcrossed to C57BL/6J background for three generations. PZU mice were crossed to C57BL/6J-ApcMin/+ mice obtained from Jackson Laboratory (Bar Harbor, ME, USA) to establish PZU; ApcMin/+ mice.
In situ hybridization
Tissues were quickly immersed into cold 4% PFA solution for 1–2 h, dehydrated with 30% sucrose in PBS (pH 7.4) at 4 °C overnight, embedded in OCT and cut into 12 μm slices, then sobbed onto slides covered by APES (amino-propyl-tri-ethoxy-silane) and dehydrated at 50 °C for 2 min and at RT in air for 2 h, and then preserved in −80 °C or treated immediately with a 479 bp biotin-tagged p53 mRNA sense or antisense probe. Slices were hybridized overnight at 65 °C (70 °C for small intestine) with the p53 RNA probe; after washed with gradient SSC solution, slices were incubated at 4 °C overnight with rabbit polyclonal antibody tagged with alkaline phosphatase (Roche, Tucson, AZ, USA; 1 : 500) against biotin; signals were developed with NBT/BCIP solution(Roche), and imaged by a bio-microscope (OLYMPUS, Tokyo, Japan). Probes were generated using an in vitro transcription kit (Roche). Primers to generate template for probes: 5′-AGTTCATTGGGACCATCCTGG-3′(F), 5′-CGTGCACATAACAGACTTGGC-3′(R).
Generation of BAC transgenic mice
p53 BAC (No.bMQ-441J16, Research Genetics, USA) was modified by homologous recombination to insert a LacZ reporter gene (from placZattB38) together with various alterations of p53 cis-regulatory elements to establish p53PZU, p53PMZU and p53PZS reporter constructs. The loxp and loxp511 sequences in the BACe3.6 backbone were then replaced with Amp and Kan prokaryotic expression cassette. The final BAC constructs were purified by Nucleo BondR max100 BAC extraction kit (MACHERERY-NAGEL, Düren, Germany), and were verified via sequencing and restriction enzyme mapping. BAC DNA was diluted to 1.0 ng/μl in microinjection buffer, and 1 pl DNA was injected into pronucleus of the fertilized mouse egg from CBA/ C57BL/6J crosses, as previously described.39 BAC transgenic copy number was estimated by comparing the photo density of bands representing p53 reporter transgene to those representing endogenous p53 allele as identified in Southern blot analysis.
X-gal staining
Embryos or tissues were fixed with cold 4% PFA for 1.5 h on ice. Embryos were stained in 1 mg/ml X-gal staining solution with gentle shaking for 24 h at 4 °C. Tissues were dehydrated, embedded in OCT and cut into 10μm slices, which were stained in 1 mg/ml X-gal staining solution for 3–10 h at either 25 or 37 °C (for comparing β-gal expression levels, staining conditions and time should be consistent between samples). X-gal staining for cells was similar as that for tissues slices except the fixation step: after washed with cold 1 × PBS, cells were fixed with 0.25% glutaraldehyde in 1 × PBS for 5 min as described.40
IHC and immunofluorecence
Tissues were dissected and fixed in cold 4% PFA over night at 4 °C, dehydrated by gradient alcohol (from 50 to 100%), rendered transparent in xylol and embedded in paraffin, then cut in a microtome to slices of 6 μm in thickness and affixed onto the APES coated slides. After deparaffinized and rehydrated, slices were performed with heat-induced epitope retrieval in sodium citrate (pH 6.0), blocked with normal goat serum block solution (Boster, Wuhan, China) for 1 h at room temperature. For IHC, slices were incubated with rabbit polyclonal antibody against β-gal (Invitrogen, Shanghai, China; A11131, 1 : 200), rat monoclonal antibody against Ki67 (Dako, Glostrup, Denmark; 1 : 100) or rabbit polyclonal antibody against p53 (Vector, Burlingame, CA, USA; CM5,1 : 500) for 18–22 h at 4 °C. β-gal or p53 was detected using Ultra Sensitive S-P (rabbit) kit (MaiXinBio, Fuzhou, China); Ki67 was incubated with the biotin-tagged goat anti-rat secondary antibody (Jackson, West Grove, PA, USA; 1 : 300) for 1 h at RT, then incubated with avidin-tagged horseradish peroxidase. All signals were developed with DAB agents (MaiXinBio) and then stained with hematoxylin. For immunofluorecence analysis, slices were incubated with chicken polyclonal antibody against β-gal (Abcam, Cambridge, MA, USA; ab9361, 1 : 200) and rat monoclonal antibody against Ki67 (Dako, 1 : 100) for 18–22 h at 4 °C, and then were incubated with the Cy3-tagged goat anti-chicken (Jackson, 1 : 500), FITC-tagged goat anti-rat antibodies (Jackson, 1 : 500) and the nucleus was stained with Hoechst (Sigma, St. Louis, MO, USA) for an hour at room temperature. Signals were imaged by either a bio- or confocal microscope (OLYMPUS).
ChIP analysis
For NIH3T3 cells: to make cells in a proliferating state, 2.2 × 106 NIH3T3 cells were cultured in 10-cm dish with 20% fetal bovine serum (FBS) in DMEM for 12 h. To make cells in a quiescent state, 2.2 × 106 NIH3T3 cells were cultured in 10-cm dish with 10% FBS in DMEM until cells reached 80–90% confluency; then cells were cultured with 0.05% FBS in DMEM for next 72 h. Cells were collected, sonicated and ChIP analysis was performed according to manufacturer’s instructions (Upstates) with Rabbit polyclonal antibody against c-Myc (Santa Cruz, Santa Cruz, CA, USA; sc-764 ×) or Rabbit IgG (Santa Cruz, Sc2763). Primer sequences used: 5′-CAGCTTTGTGCCAGGAGTCT-3′(F1), 5′-TAACTGTAGTCGCTACCTAC-3′(R1). For embryo and tissues: CHIP was performed as described41 with modifications on sonication: sonication for 1 s and pause for 2 s with the total time being 1 min for small intestine, 6 min for liver, and two 3.8 min for embryos. For MEFs: CHIP was performed similarly as 3T3 cells with a few differences: early passage MEFs from PZU or PMZU mice were plated on 10-cm dish at 1.5 × 106 density and cultured in 0.1% FBS in DMEM for 7–8 h, then cultured with 20% FBS in DMEM for 6 h. 10058-F4 (Sigma) was added to the medium at a final concentration of 180 μM, and MEFS were cultured for another 6 h and was sonicated for 4.8 min with sonication for 1 s, pause for 2 s.
Primers used for embryo, tissues and MEFs: wild type E-box, 5′- ACTTTTCACAAAGCGTTCCT-3′(F2), 5′-TTAGCCAGGGTGAGCACGTG-3′(WTR2); mutant E-box, 5′-ACTTTTCACAAAGCGTTCCT-3′(F2), 5′-TTAGCCAGGGTGAGCACACC-3′(MutantR2).
RNA and real time PCR
Total RNA samples were isolated using TRIzol (Invitrogen) and were reverse transcribed to cDNA using M-MLV Reverse Transcription kit (Invitrogen) following manufacturer’s instructions. Diluted cDNAs were used for Real Time PCR with SYBR Green reagents (Invitrogen) on an ABI Prism Step-One bio-analyzer (Foster City, CA, USA). Sequences of primers are available upon request. Expression data were normalized to β-actin mRNA, 18 s rRNA or endogeneous p53 mRNA expression. Expression changes were calculated using the ΔΔCt method and expressed as fold change over control. Experiments were repeated three times with similar results.
Western blotting
MEFs and 3T3 cells were lysed in RIPA buffer supplemented with complete Protease Inhibitor Cocktail Tablets (Roche). The membrane was incubated with anti- p53 (Vector, CM5, 1:200), anti-p21 (Santa Cruz, M19, 1:200), and appropriate secondary antibody (Pierce, Rockford, IL, USA; 1:2000) sequentially. Protein detection was performed using the ECL substrate (Thermo, Rockford, IL, USA) before exposure to film.
X-ray treatment
Mice were exposed to X-ray irradiation at 6 Gy dosage at a rate of 1.2 Gy per minute using Biological X-ray irradiator (RS2000, Rad Source, Brentwood, TN, USA). Tissues were collected 4 h after X-ray irradiation.
Statistical analysis
Data are expressed as mean±S.E.M. from triplicates. Statistical analyses were carried out using GraphPad Prism 5 software (GraphPad Software, La Jolla, CA, USA) and t-test analysis was performed between two groups. All P-values <0.05 were considered statistically significant.
Abbreviations
- Trp53 :
-
p53 gene or transformation related protein 53 gene
- BAC:
-
bacterial artificial chromosome
- c-MYC :
-
cellular myelocytomatosis oncogene
- USF-1:
-
upstream stimulatory factor-1
- E-box:
-
enhancer box sequence (CANNTG)
- ChIP:
-
chromatin immunoprecipitation
- 3′UTR:
-
3′ terminal untranslated region
- LacZ :
-
E.coli β-galactosidase Z gene
- EGFP:
-
enhanced green fluorescence protein
- SV40pA:
-
simian vacuolating virus 40 polyadenylation sequences
- FBS:
-
fetal bovine serum
- Apc min :
-
a mutation in the murine Apc gene
- Brdu:
-
5-Bromo-2-Deoxy Uridine
References
MacCallum DE, Hupp TR, Midgley CA, Stuart D, Campbell SJ, Harper A et al. The p53 response to ionising radiation in adult and developing murine tissues. Oncogene 1996; 13: 2575.
Francoz S, Froment P, Bogaerts S, De Clercq S, Maetens M, Doumont G et al. Mdm4 and Mdm2 cooperate to inhibit p53 activity in proliferating and quiescent cells in vivo. Proc Natl Acad Sci USA 2006; 103: 3232–3237.
Cheng Q, Chen J . Mechanism of p53 stabilization by ATM after DNA damage. Cell Cycle 2010; 9: 472.
Shaw P, Bovey R, Tardy S, Sahli R, Sordat B, Costa J . Induction of apoptosis by wild-type p53 in a human colon tumor-derived cell line. Proc Natl Acad Sci USA 1992; 89: 4495.
García-Cao I, García-Cao M, Martín-Caballero J, Criado LM, Klatt P, Flores JM et al. 'Super p53'mice exhibit enhanced DNA damage response, are tumor resistant and age normally. EMBO J 2002; 21: 6225–6235.
Liu G, Terzian T, Xiong S, Van Pelt CS, Audiffred A, Box NF et al. The p53–Mdm2 network in progenitor cell expansion during mouse postnatal development. J Pathol 2007; 213: 360–368.
Garcia D, Warr MR, Martins CP, Brown Swigart L, Passegué E, Evan GI . Validation of MdmX as a therapeutic target for reactivating p53 in tumors. Genes Dev 2011; 25: 1746.
Kang JH, Kim SJ, Noh DY, Park IA, Choe KJ, Yoo OJ et al. Methylation in the p53 promoter is a supplementary route to breast carcinogenesis: correlation between CpG methylation in the p53 promoter and the mutation of the p53 gene in the progression from ductal carcinoma in situ to invasive ductal carcinoma. Lab Invest 2001; 81: 573–579.
Raman V, Martensen SA, Reisman D, Evron E, Odenwald WF, Jaffee E et al. Compromised HOXA5 function can limit p53 expression in human breast tumours. Nature 2000; 405: 974–978.
Rogel A, Popliker M, Webb C, Oren M . p53 cellular tumor antigen: analysis of mRNA levels in normal adult tissues, embryos, and tumors. Mol Cell Biol 1985; 5: 2851–2855.
Schmid P, Lorenz A, Hameister H, Montenarh M . Expression of p53 during mouse embryogenesis. Development 1991; 113: 857–865.
Lookeren Campagne MV, Gill R . Tumor‐suppressor p53 is expressed in proliferating and newly formed neurons of the embryonic and postnatal rat brain: Comparison with expression of the cell cycle regulators p21Waf1/Cip1, p27Kip1, p57Kip2, p16Ink4a, cyclin G1, and the proto‐oncogene bax. J Comp Neurol 1998; 397: 181–198.
Reich NC, Levine AJ . Growth regulation of a cellular tumour antigen, p53, in nontransformed cells. Nature 1984; 308: 199–201.
Boggs K, Reisman D . Increased p53 transcription prior to DNA synthesis is regulated through a novel regulatory element within the p53 promoter. Oncogene 2006; 25: 555–565.
Dony C, Kessel M, Gruss P . Post-transcriptional control of myc and p53 expression during differentiation of the embryonal carcinoma cell line F9. Nature 1985; 317: 636–639.
Reich N, Oren M, Levine A . Two distinct mechanisms regulate the levels of a cellular tumor antigen, p53. Mol Cell Biol 1983; 3: 2143.
Reisman D, Elkind N, Roy B, Beamon J, Rotter V . c-Myc trans-activates the p53 promoter through a required downstream CACGTG motif. Cell Growth Differ 1993; 4: 57.
Qin ZH, Chen RW, Wang Y, Nakai M, Chuang DM, Chase TN et al. Nuclear factor κB nuclear translocation upregulates c-Myc and p53 expression during NMDA receptor-mediated apoptosis in rat striatum. J Neurosci 1999; 19: 4023–4033.
Hellin AC, Calmant P, Gielen J, Bours V, Merville MP . Nuclear factor-kappaB-dependent regulation of p53 gene expression induced by daunomycin genotoxic drug. Oncogene 1998; 16: 1187.
Choi M, Lee H, Rho HM . E2F1 activates the human p53 promoter and overcomes the repressive effect of hepatitis B viral X protein (Hbx) on the p53 promoter. IUBMB Life 2002; 53: 309–317.
Boggs K, Reisman D . C/EBPβ participates in regulating transcription of the p53 gene in response to mitogen stimulation. J Biol Chem 2007; 282: 7982.
Liu J, Grogan L, Nau MM, Allegra CJ, Chu E, Wright JJ et al. Physical interaction between p53 and primary response gene Egr-1. Int J Oncol 2001; 18: 863.
Venanzoni MC, Robinson LR, Hodge DR, Kola I, Seth A . ETS1 and ETS2 in p53 regulation: spatial separation of ETS binding sites (EBS) modulate protein: DNA interaction. Oncogene 1996; 12: 1199.
Chen J, Kastan MB . 5′–3′-UTR interactions regulate p53 mRNA translation and provide a target for modulating p53 induction after DNA damage. Genes Dev 2010; 24: 2146–2156.
Tian S, Huang S, Wu S, Guo W, Li J, He X . MicroRNA-1285 inhibits the expression of p53 by directly targeting its 3′ untranslated region. Biochem Biophys Res Commun 2010; 396: 435–439.
Swarbrick A, Woods SL, Shaw A, Balakrishnan A, Phua Y, Nguyen A et al. miR-380-5p represses p53 to control cellular survival and is associated with poor outcome in MYCN-amplified neuroblastoma. Nat Med 2010; 16: 1134–1140.
Thompson NL, Mead JE, Braun L, Goyette M, Shank PR, Fausto N . Sequential protooncogene expression during rat liver regeneration. Cancer Res 1986; 46: 3111.
Roy B, Beamon J, Balint E, Reisman D . Transactivation of the human p53 tumor suppressor gene by c-Myc/Max contributes to elevated mutant p53 expression in some tumors. Mol Cell Biol 1994; 14: 7805–7815.
Chen L, Iraci N, Gherardi S, Gamble LD, Wood KM, Perini G et al. p53 is a direct transcriptional target of MYCN in neuroblastoma. Cancer Res 2010; 70: 1377.
Casado M, Vallet VS, Kahn A, Vaulont S . Essential role in vivo of upstream stimulatory factors for a normal dietary response of the fatty acid synthase gene in the liver. J Biol Chem 1999; 274: 2009–2013.
Gregor PD, Sawadogo M, Roeder RG . The adenovirus major late transcription factor USF is a member of the helix-loop-helix group of regulatory proteins and binds to DNA as a dimer. Genes Dev 1990; 4: 1730–1740.
Mazan-Mamczarz K, Galbán S, López de Silanes I, Martindale JL, Atasoy U, Keene JD et al. RNA-binding protein HuR enhances p53 translation in response to ultraviolet light irradiation. Proc Natl Acad Sci USA 2003; 100: 8354.
Vilborg A, Glahder JA, Wilhelm MT, Bersani C, Corcoran M, Mahmoudi S et al. The p53 target Wig-1 regulates p53 mRNA stability through an AU-rich element. Proc Natl Acad Sci 2009; 106: 15756–15761.
Burns DM, Richter JD . CPEB regulation of human cellular senescence, energy metabolism, and p53 mRNA translation. Genes Dev 2008; 22: 3449.
Le MTN, Teh C, Shyh-Chang N, Xie H, Zhou B, Korzh V et al. MicroRNA-125b is a novel negative regulator of p53. Genes Dev 2009; 23: 862.
Hellström NAK, Björk‐Eriksson T, Blomgren K, Kuhn, H G . Differential recovery of neural stem cells in the subventricular zone and dentate gyrus after ionizing radiation. Stem Cells 2009; 27: 634–641.
Tada E, Parent J, Lowenstein D, Fike J . X-irradiation causes a prolonged reduction in cell proliferation in the dentate gyrus of adult rats. Neuroscience 2000; 99: 33–41.
Liberman LM, Stathopoulos A . Design flexibility in cis-regulatory control of gene expression: synthetic and comparative evidence. Dev Biol 2009; 327: 578–589.
Chandler KJ, Chandler RL, Broeckelmann EM, Hou Y, Southard-Smith EM, Mortlock DP . Relevance of BAC transgene copy number in mice: transgene copy number variation across multiple transgenic lines and correlations with transgene integrity and expression. Mamm Genome 2007; 18: 693–708.
Zuk PA, Zhu M, Mizuno H, Huang J, Futrell JW, Katz AJ et al. Multilineage cells from human adipose tissue: implications for cell-based therapies. Tissue Eng 2001; 7: 211–228.
Feng D, Liu T, Sun Z, Bugge A, Mullican SE, Alenghat T et al. A circadian rhythm orchestrated by histone deacetylase 3 controls hepatic lipid metabolism. Science 2011; 331: 1315–1319.
Acknowledgements
We thank NRCMM mouse facility for assistance in generating the BAC transgenic mice. This work was supported by grants from Natural Science Foundation of China (30871265, 31171305) to GL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Supplementary Information accompanies this paper on Cell Death and Disease website .
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chen, L., Zhang, G., Zhou, Y. et al. BAC transgenic mice provide evidence that p53 expression is highly regulated in vivo. Cell Death Dis 6, e1878 (2015). https://doi.org/10.1038/cddis.2015.224
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2015.224