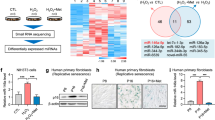
Abstract
Here we show that transgenic expression of miR-17 extends lifespan and inhibits cellular senescence. We propose that miR-17 acts as a critical regulator of cellular senescence and tumorigenesis. We demonstrate that miR-17 targets both ADCY5 and IRS1, upregulating the downstream signals MKP7, FoxO3, LC3B, and HIF1α, and downregulating mTOR, c-myc, cyclin D1, and JNK. Silencing either ADCY5 or IRS1 promoted autophagy and repressed cellular senescence and apoptosis. Repression of ADCY5 by miR-17 translocated membrane-bound RGS2 into the nucleus, promoting interactions of RGS2 with HIF1α and the MKP7 promoter, enhancing MKP7 transcription. ADCY5 repression by miR-17 also facilitated the translocation of EGFR and MKP7 from membrane into cytoplasmic and mitochondrial fractions. Importantly, we found that MKP7 inhibited senescence by dephosphorylating PRAS40 at Thr246 and mTOR at Ser2248, facilitating the interaction and loss of function of both molecules. Thus, the oncogenic miR-17 also acts pleiotropically to inhibit cellular senescence and extend longevity.
Similar content being viewed by others
Main
MicroRNAs (miRNAs) are short strands of RNAs of 18–24 nucleotides in length. Several miRNAs in 17∼92 cluster have been reported to play important roles in cancer development through the repression of tumor-associated genes.1, 2, 3, 4 Overexpression of the miR-17∼92 cluster has been shown to enhance tumor cell proliferation and reduce apoptosis by regulating cell-cycle progression.5, 6 Expression of miR-17, a miRNA that has been shown to downregulate the tumor suppressor PTEN,7, 8 is widely believed to be oncogenic. Interestingly, miR-17 has also been found to be downregulated in human aging cells.9 Lifespan is mainly affected by tissue aging and cellular senescence. Studies have revealed conservation of lifespan-regulating genes and their signaling networks. Decreased target of rapamycin (TOR) signaling and insulin/IGF-like signaling are two central events to extend lifespan.
TOR, or mTOR (mechanistic target of rapamycin) in mammals, is a cytoplasmic protein kinase that exists in all cells of all species.10, 11 The mTOR pathway is a sensor of nutrients, growth factors, hormones, mitogens, and cytokines.12, 13 In response to signals from these molecules, mTOR promotes cellular mass growth. Hyperstimulation and hyperactivation of mTOR lead to cellular senescence and tissue aging resulting in decreased lifespan. Importantly, inhibition of TOR or raptor, an essential component of TOR complex, extends lifespan dramatically.14, 15
The insulin/IGF-like signaling is widely accepted as a signaling pathway that plays a central role in the regulation of organism growth and lifespan.16 Lifespan is prolonged in yeast, flies, and rodents when the activity of the insulin/IGF-like signaling is reduced although these organisms are different in their shape and size.17 Worms with a mutation in daf-2, a Caenorhabditis elegans homolog of insulin/IGF receptor lead to active and healthy lives twice as long as those of wild-type worms.18, 19 Decreased function of insulin/IGF-like signaling by the Forkhead transcription factor can extend lifespan.20 Adenylate cyclase 5, a membrane-bound enzyme, can catalyze production of cyclic adenosine monophosphate (cAMP) and modulate glucose-stimulated insulin secretion and proinsulin-to-insulin conversion.21 Knocking out the ADCY5 gene (adenylate cyclase type 5) in mice may protect cardiac muscle from apoptosis and extend lifespan.22
In this study, we found that transgenic expression of miR-17 increased mouse lifespan by repressing expression of insulin receptor substrate 1 (IRS1) and ADCY5. IRS1 has been shown to play a key role in transmitting signals from the insulin and insulin-like growth factor 1 (IGF-1) receptors to intracellular pathways.23, 24 We also found that decreased ADCY5 promoted expression of the dual specificity phosphatase 16 (DUSP16/MKP7), an enzyme that inactivates their target kinases by dephosphorylating both the phosphoserine/threonine and phosphotyrosine residues including JNK (c-Jun N-terminal kinases) and ERK (extracellular signal-regulated kinase).25 Increased MKP7 appeared to play a similar role as insulin-inhibitory signaling. In addition, increased MKP7 inhibited mTOR and JNK activation, but increased HIF1α (hypoxia-inducible factor 1-alpha) expression. Increased HIF1α in turn promoted MKP7 transcription, resulting in the inhibition of cellular senescence and tissue aging. Through a number of signaling pathways, we propose that the oncogenic miR-17 can also act to downregulate cellular senescence and extend lifespan. This is an important step in understanding the implications of therapeutically targeting oncogenic miRNAs.
Results
Expression of miR-17 decreases senescence
We have reported that transgenic mice expressing miR-17 grew slower in the early stages of development.26 As these mice aged, however, we found that they developed liver tumors, which were pathologically similar to human hepatocellular carcinoma.8 When examining the lifespans of the miR-17 transgenic mice, we unexpectedly found that the miR-17 transgenic mice had significantly longer lifespans as compared with wild-type mice (Figure 1a).
Expression of miR-17 repressed organismal and cellular senescence. (a) Lifespans (survival days) of miR-17 transgenic and wild-type mice, showing longer lifespan of the miR-17 transgenic mice. (b) The femur of transgenic and wild-type mice was embedded in the same paraffin wax block. One section of 5 μm was obtained from every 100 μm coronally or sagittally. The miR-17 transgenic sections showed increased bone mass. (c) Left, sections of intestine, lung, and heart of transgenic and wild-type mice were subject to β-gal staining. Transgenic mice displayed less β-gal staining for all organs. Right, sections of skin were subject to β-gal staining. Transgenic mice showed less β-gal staining. (d) Left, the number of β-gal stained cells were counted and quantified. miR-17 expression decreased β-gal staining compared to the control. Asterisks indicate significant differences. *P<0.05; **P<0.01. Error bars, S.D. (n=6). Right, fewer miR-17-transfected MEF cells showed β-gal staining than the control after cultured in serum-free medium. (e) After being treated with H2O2 for 2 h, and then cultured in basal medium for 48 h, the miR-17-transfected MEFs showed less β-gal staining than the control. *P<0.05; **P<0.01. Error bars, S.D. (n=6)
We examined the femur bone mass and found that the aged miR-17 transgenic mice displayed increased bone mass as compared with the wild-type mice (Figure 1b). A number of organs, including intestine, lung, heart, and skin, were then subject to β-galactosidase (β-gal) staining, a molecular marker of senescence. It was found that the miR-17 transgenic mice displayed lower intensities of β-gal staining than the wild-type mice (Figure 1c).
To corroborate the role of miR-17 in senescence, we developed a mouse embryonic fibroblast (MEF) cell line stably transfected with either an miR-17 expression construct or a control vector. Expression of two mature miRNAs, miR-17-5p and miR-17-3p, was confirmed (Supplementary Figure S1a). Analysis of senescence by β-gal staining revealed that ectopic expression of miR-17 decreased senescence when the cells were cultured in a serum-free medium (Figure 1d) or treated with H2O2 for 2 h, followed by culturing in basal medium for 48 h (Figure 1e). We also found that the MEF cells transfected with miR-17 had a lower rate of apoptosis (Supplementary Figure S1b) and the cells survived longer (Supplementary Figure S1c) when cultured in serum-free medium or treated with H2O2.
miR-17 represses ADCY5 and IRS1 but promotes MKP7
Protein expression was examined to uncover proteins that might be mediating miR-17 function in senescence. MiR-17- and control vector-transfected MEFs were cultured in normal medium, serum-free medium, or medium treated with H2O2. We found that ADCY5, IRS1, PTEN, pmTOR, c-myc, cyclin D1, and pJNK were downregulated, while MKP7, FoxO3a (Forkhead box O3), LC3B, pAkt, GSK-3β (S9P; glycogen synthase kinase-3β serine-9), and HIF1α were upregulated (Figure 2a). Among the downregulated proteins, ADCY5 was a potential target of miR-17-5p and IRS1 was a potential target of miR-17-3p (Supplementary Figure S1d). To confirm direct targeting of ADCY5 by miR-17-5p and IRS1 by miR-17-3p, we generated luciferase constructs harboring a fragment of ADCY5 or IRS1 (Supplementary Figure S1e). The potential target sites were mutated. Luciferase assay confirmed that miR-17-5p could target ADCY5, while miR-17-3p could target IRS1 (Figure 2b). The target repression appeared to occur post-transcriptionally, since the levels of ADCY5 and IRS1 mRNAs were not affected by miR-17 transfection (Supplementary Figure S1f).
Targeting ADCY5 and IRS1 by miR-17. (a) The vector- or miR-17-transfected MEFs were cultured in 10% FBS or serum-free medium for 48 h, or treated with 75 μM for 2 h, and then cultured in 10% FBS/medium for 48 h, followed by western blot analysis, which showed downregulation of ADCY5, IRS1, PTEN, pmTOR, c-myc, cyclin D1, and pJNK, and upregulation of MKP7, FoxO3a, LC3B, pAkt, GSK-3β (S9P), and HIF1α. All membranes were reprobed with anti-β-actin antibody to confirm equal loading, which is shown beneath of each blot. (b) Left, decreased luciferase activities were observed in 293T cells co-transfected with miR-17-5p and luciferase reporter construct Luc-ADCY5 (luc-AC5), which was reversed when the miR-17-5p binding site was mutated (luc-AC5-mut). Right, decreased luciferase activities were observed in cells co-transfected with miR-17-3p and luc-IRS1, which was reversed when the miR-17-3p binding site was mutated (luc-IRS1-mut). **P<0.01. Error bars, S.D. (n=4). (c) Upper, decreased expression of ADCY5 and IRS1 was detected in MEFs transfected with miR-17-5p or/and -3p mimic (Mi) compared with control (N.C.). Lower, increased expression of ADCY5 and IRS1 was detected in both miR-17- and control vector-transfected MEF cells when the cells were transfected with miR-17-5p or/and -3p inhibitor (In) compared with control
To confirm miRNA targeting, we transfected MEFs with miR-17-5p and/or miR-17-3p mimics and found that ADCY5 was repressed in the presence of miR-17-5p, while IRS1 was repressed in the presence of miR-17-3p (Figure 2c, upper). When miR-17 and control vector-transfected MEF cells were transfected with miR-17-5p and/or miR-17-3p inhibitor compared with a control oligo, expression of ADCY5 and IRS1 was increased (Figure 2c, lower). Typical full-gel photos of ADCY5, IRS1, PTEN, pmTOR, c-myc, cyclin D1, pJNK, MKP7, FoxO3a, LC3B, pAKT, GSK-3β (S9P), HIF1α, and IR are provided in (Supplementary Figure S2).
It has been reported that ADCY5 knockout decreased cAMP levels.22 We analyzed cAMP levels in cells transfected with miR-17 or the control vector and found little difference between the cell lines (Supplementary Figure S3a). ADCY5 silencing was confirmed by western blotting (Supplementary Figure S3b). MEFs transfected with miR-17-5p mimic or siRNA (small interfering RNA) targeting ADCY5 produced lower levels of cAMP transiently, mainly in the fractions associated with the integral membrane protein, caveolin (Supplementary Figure S3c). MiR-17 expressing MEF cells produced ADCY5 at the highest levels among the nine isoforms analyzed, although both vector- and miR-17-transfected cells produced comparable levels of the adenylate cyclases (Supplementary Figure S3d).
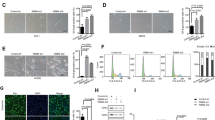
To test the roles of ADCY5 and IRS1 in mediating miR-17 function, we transfected MEFs with siRNAs targeting ADCY5, IRS1, and PTEN, as PTEN has been reported to be a target of miR-17-5p.7 It was confirmed that MEFs transfected with siRNAs targeting ADCY5, IRS1, and PTEN expressed lower levels of these proteins compared with control (Supplementary Figure S4a). Silencing ADCY5 or IRS1, but not PTEN, was found to repress senescence (decreased β-gal staining), when the cells were cultured in the serum-free medium or treated with H2O2 (Figure 3a; Supplementary Figure S4b). Western blot analysis revealed that when ADCY5 was silenced, levels of MKP7, FoxO3a, and LC3B, and HIF1α were upregulated, while c-myc, cyclin D1, and pmTOR were downregulated (Figure 3b). Silencing IRS1 increased FoxO3a and LC3B expression, while silencing PTEN upregulated pAKT and GSK3β (S9P) levels. Our results, combined with previously reported signaling pathways, allowed us to illustrate a signaling network, by which miR-17 functions (Figure 3c).
miR-17 repressed cellular senescence via promoting expression of MKP7. (a) The number of β-gal-stained MEFs decreased in cells transfected with siRNAs targeting ADCY5 and IRS1 compared with cells transfected with a control oligo or siRNA targeting PTEN, when the cells were cultured in serum-free medium for 4 days (left) or treated with 150 μM H2O2 (right). *P<0.05; **P<0.01. Error bars, S.D. (n=6). (b) Silencing ADCY5 upregulated levels of MKP7, FoxO3a, and LC3B, HIF1α, and pJNK, but downregulated c-myc, cyclin D1, and pmTOR. Silencing IRS1 promoted FoxO3a and LC3B expression. Silencing PTEN upregulated pAKT and GSK3β (S9P) levels. All membranes were reprobed with anti-β-actin antibody to confirm equal loading, which is shown beneath of each blot. (c) miR-17 repressed MEFs senescence by targeting ADCY5 and IRS, and enhancing expression of MKP7. (d) Silencing FoxO3a downregulated LC3B level. Silencing MKP7 increased levels of pJNK, pmTOR, c-myc, and cyclin D1, but downregulated HIF1α, FoxO3a, and LC3B. Silencing HIF1α had little effect on the expression of all proteins analyzed. (e) The number of β-gal-stained miR-17-transfected MEFs increased in cells treated with siRNAs targeting MKP7 and FoxO3a compared with cells treated the control oligo or siRNA targeting HIF1α, when the cells were treated with 150 μM H2O2 (n=6)
To confirm the downstream signaling of miR-17-5p, we silenced MKP7, FoxO3a, and HIF1α (Supplementary Figure S4c). Silencing MKP7 increased levels of pJNK, pmTOR, c-myc, and cyclin D1, but downregulated HIF1α, FoxO3a, and LC3B (Figure 3d). Silencing FoxO3a downregulated LC3B level, while silencing HIF1α had little effect on expression of all proteins analyzed. By functional analysis, we found that silencing MKP7 and FoxO3a, but not silencing HIF1α, increased cellular senescence, when the cells were treated with 150 μM H2O2 (Figure 3d, Supplementary Figure S4d) or cultured in the serum-free medium (Supplementary Figure S4e). In these conditions, silencing MKP7 and HIF1α, but not FoxO3a, decreased cell survival (Supplementary Figure S4f) but increased apoptosis (Supplementary Figure S4g), suggesting different effects of these molecules in senescence and cell death.
miR-17 regulates MKP7 subcellular translocation
To examine the distribution of MKP7, we isolated caveolin-associated membrane fractions from lysates of mock- and miR-17-transfected MEFs. While upregulation of MKP7 in the lysate of miR-17-transfected cells was confirmed, a significant translocation of MKP7 and epidermal growth factor receptor (EGFR) to non-caveolin fraction was observed (Figure 4a; Supplementary Figure S5a). Although the total levels of EGFR was not affected by miR-17 transfection (Figure 4a), we detected a clear translocation of EGFR to the mitochondria in the cells transfected with miR-17 with or without stress (Figure 4b). Expression of miR-17 also enhanced MKP7 expression in the mitochondrial fractions. In the nuclear fraction, EGFR localization was increased but MKP7 expression decreased when the cells were stressed in serum-free conditions or treated with H2O2, which was not found in mitochondrial fractions (Figure 4b). Such protein translocation by cell stress was not found for RGS2 (regulator of G-protein signaling 2). The increase in EGFR and decrease in MKP7 were validated by confocal microscopy (Figure 4c). Representative images are provided in (Supplementary Figure S5b; Supplementary Figure S6a).
Expression of miR-17 promoted MKP7 subcellular translocation. (a) Left, cell lysates from mock- and miR-17-transfected MEFs were probed with antibodies as indicated. Right, membranes isolated from the lysates were probed with antibodies as indicated. miR-17 transfection promoted translocation of MKP7 and EGFR to non-caveolin fraction. (b) Upper, mitochondria fraction was probed with antibodies against EGFR, MKP7, and VDAC1/Porin. Lower, nuclear extract of mock- and miR-17-transfected MEFs were probed with antibodies as indicated. Cells were cultured in serum-free medium for 48 h, or treated with 75 μM H2O2 for 2 h, then cultured for 48 h showed increased levels of EGFR but decreased levels of MKP7 in the nucleus compared with cells cultured in basal medium. (c) Upper, the MEFs were also stained with phalloidins (green) for cell structure, DAPI (blue) for nucleus, and red fluorescence showing expression of MKP7, RGS2, or EGFR. miR-17-transfected MEFs showed enhanced expression of MKP7, and distinguished translocation of RGS2 to the nucleus. However, both control and miR-17 MEFs showed depleted MKP7 and condensed EGFR in the nucleus, when cultured in serum-free medium or treated with H2O2. Lower, in addition, the cells were also stained with VADC1 (yellow) for mitochondria, and red staining showing expression of MKP7 or EGFR. miR-17 transfection enhanced expression of MKP7 and EGFR in the mitochondria
Confirmation of ADCY5 and IRS1 effects
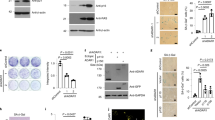
We then validated the effects of ADCY5 on mediating the function of miR-17 in senescence. We first confirmed that ectopic expression of ADCY5 decreased levels of MKP7, HIF1α, LC3B, and FoxO3a, but increased levels of pJNK, pmTOR, c-myc, and cyclin D1 (Figure 5a). ADCY5 expression facilitated translocation of EGFR, MKP7 and RGS2 to caveolin-associated fraction from the non-caveolin fraction (Figure 5a; Supplementary Figure S6b), but had little effect on IRS1 expression and translocation (Supplementary Figure S6c). Functionally, ectopic expression of ADCY5 increased β-gal-positive cells in the miR-17-transfected MEFs, when the cells were stressed (Figure 5b; Supplementary Figure S7a). Consistent with this was the decreased cell survival (Supplementary Figure S7b) and increased apoptosis (Supplementary Figure S7c) in the stressed cells.
Overexpression of ADCY5 or IRS1 abolished miR-17 repressed MEFs senescence and MKP7 subcellular translocation. (a) It was confirmed that ectopic expression of ADCY5 decreased levels of MKP7, HIF1α, LC3B, and FoxO3a, but increased levels of pJNK, pmTOR, c-myc, and cyclin D1 (left). In the membrane fraction, pEGFR disappeared in the non-caveolin preparation (middle). In the nucleus, EGFR levels were promoted, while MKP7 levels were suppressed in the stress conditions (right, cell treated with H2O2 or cultured in serum-free medium). (b) Ectopic expression of ADCY5 increased β-gal-positive cells in the miR-17-transfected MEFs, when the cells were cultured in serum-free medium for 4 days (left) or treated with 150 μM H2O2 (right). **P<0.01. Error bars, S.D. (n=6). (c) Ectopic expression of IRS1 decreased levels of LC3B and FoxO3a. (d) Ectopic expression of IRS1 increased β-gal-positive cells in the miR-17-transfected MEFs, when the cells were cultured in serum-free medium for 4 days (left) or treated with 200 μM H2O2 (right). **P<0.01. Error bars, S.D. (n=6). (e) Cell lysates (left) and membrane (right) from mock- and miR-17-transfected MEFs were probed with antibodies to pERK, ERK2, pMEK1, and MEK1. (f) Upper, total cell lysates and membrane fraction from si-ADCY5-transfected MEFs were subject to immunoblotting for expression of pERK, ERK2, pMEK1, MEK1, PTPN9, and PTPRF. Lower, miR-17-transfected cells were transiently transfected with IRS1 and a control vector. Expression of ERK and MEK was analyzed. (g) Diagram showing the relationship of miR-17, ADCY5, and IRS1
To validate the effects of ADCY5 on protein translocation, we knocked down ADCY5 as above and analyzed protein distribution in membrane, nuclei, and mitochondria. After confirming protein fractionation (Supplementary Figure S7d), we found that silencing ADCY5 promoted enrichment of MKP7, EGFR, pEGFR, and RGS2 in the non-caveolin fraction (Supplementary Figure S7e), an effect similar to miR-17 overexpression (Figure 4a). In addition, silencing ADCY5 also played a similar role as miR-17 overexpression for protein distribution in nuclei and mitochondria (Supplementary Figure S7f).
We then validated the effects of IRS1 using a similar approach. Cells transfected with IRS1 were subject to western blot analysis. The downstream signal molecules LC3B and FoxO3a were confirmed downregulated in the total cell lysates (Figure 5c). The lysates were then subject to fractionation (Supplementary Figure S8a). Unlike ADCY5, expression of IRS1 neither show any effect on translocation of MKP7, EGFR, and RGS2 to the non-caveolin fraction (Supplementary Figure S8b) nor played additional effect on cell stress (Supplementary Figure S8c). Nevertheless, expression of IRS1 increased cell senescence (Figure 5d), although it did not affect cell survival (Supplementary Figure S8d) and apoptosis (Supplementary Figure S8e).
It has been reported that ADCY5 knockdown increased lifespan in vivo and in vitro.22 The major mechanism is increased phosphorylation of MEK1 (MAPK/ERK kinase-1) and ERK. It also involved decreased accumulation of cAMP. Since ADCY5 was found to be repressed by miR-17 expression, we examined total cell lysate and membrane fraction of the miR-17- and vector-transfected cells for pERK and pMEK1, using PTPN9 (cytoplasmic protein) and PTPRF (membrane protein) as controls ensuring successful fractionation of proteins. We did not detect change of pERK and pMEK1 (Figure 5e). Nevertheless, when ADCY5 was suppressed by siRNA targeting ADCY5, significant upregulation of pERK and pMEK1 was detected, which was mainly due to the increase in non-caveolin fraction (Figure 5f, upper). We then overexpressed IRS1 in the miR-17-transfected cells and detected increased levels of pERK and pMEK1, again, mainly in the cytoplasm (Figure 5f, lower). These results suggested that while ADCY5 can repress phosphorylation of ERK and MEK1 as reported,22 IRS1 could promote this process (Figure 5g). As a consequence, expression of miR-17, which repressed both ADCY5 and IRS1, displayed no potent effect on ERK and MEK1 phosphorylation.
MKP7 represses mTOR function
It has been reported that mTOR plays important roles in cell differentiation, autophagy, and senescence27, 28 and phosphorylation at the Ser2248 is essential.29 However, dephosphorylated PRAS40 at Thr246 can bind to the dephosphorylated mTOR2248 and inhibit mTOR activity.30 We found that expression of miR-17 decreased levels of pmTOR and pPRAS40, while MKP7 level was promoted (Figure 6a). In co-immunoprecipitation assays, antibody against mTOR pulled down PRAS40 and MKP7, while antibody against PRAS40 pulled down MKP7, suggesting the interaction of mTOR and PRAS40 with MKP7.
Expression of MKP7 repressed mTORC1 function. (a) Lysates prepared from cells transfected with miR-17 or a control vector as indicated were subject to immunoprecipitation with anti-mTOR or anti-PRAS40 antibodies, followed by western blotting probed with antibodies as indicated. mTOR precipitation pulled down MKP7 and PRAS40. PRAS40 precipitation pulled down MKP7. (b) Lysates prepared from cells transfected with MKP7 or a vector as indicated were subject to immunoprecipitation with anti-mTOR or anti-PRAS40 antibodies, followed by western blot analysis. mTOR precipitation pulled down MKP7 and PRAS40. PRAS40 precipitation pulled down MKP7. (c) Lysates prepared from cells transfected with siRNA targeting MKP7 or a control oligo were subject to immunoprecipitation with anti-mTOR or anti-PRAS40 antibodies, followed by western blot analysis. mTOR precipitation pulled down trace amount of PRAS40 but not MKP7. PRAS40 precipitation pulled down decreased levels of MKP7 compared with the control. (d) Left, immunoprecipitated MKP7 and non-precipitated total lysate of vector- or MKP7-transfected MEFs were incubated with immunoprecipitated mTOR for 24 h. The reaction was terminated by adding × 2 Laemmli buffer, and the mix was subject to western blot analysis for pmTOR (Ser2248) expression. In the presence of MKP7, mTOR was dephosphorylated at Ser2248. Right, immunoprecipitated and non-precipitated MKP7 were incubated with immunoprecipitated PRAS40 for 24 h, followed by western blot analysis. MKP7 dephosphorylated pPRAS40 at Thr246. (e) Mechanism of MKP7 repressing mTOR function
MKP7 has been known to function in dephosphorylation of other proteins.31, 32 We overexpressed MKP7 in MEFs and found that the levels of pmTOR and pPRAS40 were greatly decreased (Figure 6b). In co-immunoprecipitation assays, anti-mTOR antibody pulled down PRAS40 and MKP7, while anti-PRAS40 antibody pulled down MKP7. To validate the dephosphorylating activity of MKP7, MEFs were transfected with siRNA targeting MKP7. This resulted in upregulation of pmTOR and pPRAS40 (Figure 6c).
We then tested whether MKP7 could directly dephosphorylate pmTOR by mixing the purified MKP7 from different preparation of cell lysate with the purified total mTOR. Incubation of both purified proteins resulted in dephosphorylation of pmTOR by MKP7, greater amount of MKP7 resulting in dephosphorylation of pmTOR (Figure 6d). Similarly, purified MKP7 resulted in decreased pPRAS40 level. These results suggest that MKP7 could directly dephosphorylate pmTOR and pPRAS40 and forming complexes with these two proteins (Figure 6e).
To confirm upregulation of MKP7 in the miR-17 transgenic mice, we analyzed expression of MKP7, ADCY5, and IRS1 in the organs of the miR-17 transgenic mice. In the heart, intestine, kidney, lung, skin, and spleen, we found that MKP7 was upregulated but ADCY5 and IRS1 were downregulated as compared with wild-type mouse organs (Figure 7a). To examine how MKP7 expression was upregulated, we isolated chromatins of mock- and miR-17-transfected MEFs and incubated digested chromatins with anti-RGS2, or anti-HIF1α antibodies. The precipitated DNA was subject to polymerase chain reaction (PCR) with specific primers flanking a piece of DNA sequence at the MKP7 promoter. It was found that miR-17 expression enhanced RGS2 and HIF1α binding to MKP7 promoter (Figure 7b). It appeared that expression of miR-17 repressed ADCY5, allowing RGS2 to translocate to nucleus, where it interacted with HIF1α, enhancing HIF1α binding to MKP7 promoter, and promoting MKP7 transcription. Increased levels of MKP7 in the cytoplasm and mitochondria dephosphorylated senescence-related proteins (e.g., pmTOR and pPRAS40), particularly during stress. Repressing ADCY5 also facilitated EGFR translocation into the cytoplasm and mitochondria. Increased mitochondrial EGFR and MKP7 in turn enhanced autophagy, but decreased senescence and apoptosis (Figure 7c). To validate the interaction of RGS2 and HIF1α, nuclear proteins were subject to co-immunoprecipitation. The assay showed that anti-RGS2 antibody could pull down HIF1α, anti-HIF1α antibody could pull down RGS2 (Figure 7d).
The role of MKP7 subcellular translocation in anti-senescence. (a) Sections of heart, intestine, kidney, lung, skin, and spleen of transgenic and wild-type mice were immunostained for expression of ADCY5, IRS1, and MKP7. Transgenic mice expressed high levels of MKP7 and low levels of ADCY5 and IRS1. (b) Chromatins of mock- and miR-17-transfected MEFs were isolated, digested and immunoprecipitated with rabbit IgG (negative control) and antibodies against histone H3 (positive control), RGS2, or HIF1α, followed by PCR or RT-PCR with specific primers flanking a piece of DNA sequence at the MKP7 promoter. miR-17 expression enhanced RGS2 and HIF1α binding to MKP7 promoter. (c) Expression of miR-17 represses ADCY5, which releases RGS2, and allowed it to translocate to nucleus and to bind to MKP7 promoter. Nuclear RGS2 can also bind to HIF1α and enhances HIF1α binding to MKP7 promoter, promoting MKP7 transcription. Increased levels of MKP7 in the cytoplasm and mitochondria dephosphorylate senescence-related proteins (e.g., pmTOR and pPRAS40), in particular during stress. Repressing ADCY5 also facilitates EGFR moving to the cytoplasm and mitochondria. Increased mitochondria levels of EGFR and MKP7 can enhance autophagy, but decrease senescence and apoptosis. (d) Nuclear preparation prepared from cells transfected with miR-17 or a control vector was subject to immunoprecipitation with anti-RGS2 or anti-HIF1α antibodies, followed by western blotting probed with antibodies as indicated. Anti-RGS2 antibody pulled down HIF1α, anti-HIF1α antibody pulled down RGS2
Discussion
In this study, we found that transgenic expression of miR-17 suppressed cellular and organismal senescence and prolonged lifespan in mice. In specific contexts, multiple miRNAs have been shown to both positively and negatively regulate senescence and lifespan.33, 34, 35, 36, 37 Previously, most studies focused on the effect of miRNAs on senescence at the cellular level, or investigated the longevity role of miRNAs in lower organisms.38, 39 Uniquely, our study systematically describes the functions of one specific miRNA in controlling senescence and aging at both the organismal and cellular levels. The precursor microRNA miR-17 can be processed to produce two mature miRNAs, miR-17-5p and miR-17-3p. In the transgenic mice and transfected cells, the levels of both mature miRNAs were comparable and were shown to target ADCY5 and IRS1, respectively.
ADCY5 has been shown to interact with the N-terminal domain of RGS2.40 We found that silencing ADCY5 released the membrane-bound RGS2, allowing it to be translocated to the nuclei where it interacted with HIF1α. The RGS2-HIF1α complex bound to the MKP7 promoter, increasing MKP7 transcription. It has been reported that ADCY5 knockout extended lifespan by upregulating MEK/ERK signaling pathway.22, 41 We found that enhanced expression of MKP7 is a major mechanism regulating lifespan and senescence. Our study demonstrates that enhancing MKP7 expression is an additional function of ADCY5, a functional target of miR-17-5p. MKP7, or Dusp16 is a member of the dual specificity protein phosphatase family. It can activate mitogen-activated protein kinase (MAPK) cascades and negatively regulates MAPK activity.42 We found that increased expression of MKP7 inhibited mTOR signaling by dephosphorylating mTOR at Ser2248 and PRAS40 at Thr246. The dephosphorylated mTOR and PRAS40 formed a complex resulting in quiescent mTOR activity. It has been shown previously that decreased mTOR activity inhibits cellular senescence.43, 44 Our results showed that increased MKP7 expression represents a different layer of function in the ability of the miR-17-ADCY5 axis in suppressing mTOR activity and inhibiting cellular senescence.
We also found that increased MKP7 expression played a similar role as repressing IRS1 expression, which enhanced expression of FoxO3a and LC3B. IRS1 signaling has been reported to be an evolutionarily conserved pathway regulating mammalian lifespan.45, 46 We found that enhanced expression of FoxO3 and LC3B could induce autophagy and inhibit senescence. Thus, there appeared to be a cross-talk between mTOR signaling and insulin signaling in reducing cellular senescence mediated by MKP7. Associated with this effect, MKP7 could also inhibit c-myc and cyclin D1 expression resulting in inhibited senescence.
In addition, we found that MKP7 could inhibit JNK activity and enhance HIF1α expression resulting in inhibiting cell apoptosis. This effect was similar to miR-17 repression of PTEN expression. PTEN has been shown to regulate variety of cell activities in the suppression of tumor growth.47 We have previously shown that miR-17 enhanced tumor growth by repressing PTEN expression.8 Here we confirmed that PTEN was targeted by miR-17 resulting in inhibitory apoptosis. Thus, MKP7 could function similarly to PTEN in modulating cellular apoptosis. Interestingly however, repression of PTEN by miR-17 was not found to be sufficient to modulate cellular senescence. We thus hypothesized that miR-17 was eliciting a tumorigenic and anti-senescence phenotype by pleiotropically targeting different signaling pathways.
Another molecule that mediated miR-17-ADCY5 signaling appeared to be EGFR. Silencing ADCY5 stimulated EGFR translocation to mitochondria, resulting in upregulation of ERK and MEK. This was shown to promote autophagy and inhibit senescence. Silencing ADCY5 also increased MKP7 levels in the mitochondria. Increased MKP7 would promote FoxO3a and LC3B activities, resulting in enhanced autophagy and decreased senescence. Under stressed conditions, EGFR showed significant nuclear translocation and MKP7 showed cytoplasm translocation, which might be related to EGFR-regulated stress transcription. This could act to enhance MKP7 function in dephosphorylating cytoplasmic proteins. The specific functions of mitochondrial, cytoplasmic and nuclear EGFR and MKP7 in autophagy, senescence, and apoptosis await further investigation.
ADCY5 knockout mice showed increased longevity as a result of activating Raf/MEK/ERK signaling and cAMP levels.22 In this study, we found that silencing ADCY5 only temporarily repressed cAMP levels in MEFs, and that stable knockouts of ADCY5 did not have any effects on cAMP levels. This may be due to the fact that there are nine isoforms of ADCY. Although we were able to detect a decrease in cAMP after transient ADCY5 transfection, stable transfection of ADCY5 did not detect clear reduction of cAMP. This might be the result of supplementary effect from other isoforms. We were also able to detect increased levels of pERK and pMEK in the cells transfected with siRNA targeting ADCY5. This is in agreement with previously reported results. However, in the cells transfected with miR-17, we did not detect change in pERK and pMEK levels. This appeared to be the opposite effect of silencing IRS1. Silencing IRS1 decreased pERK and pMEK levels, an opposite effect of silencing ADCY5. As a consequence, overexpression of miR-17, which silenced both IRS1 and ADCY5, produced little effects on pERK and pMEK levels. These results suggest that miR-17 inhibited senescence through mechanisms other than increasing pERK and pMEK levels.
In summary, we found that transgenic expression of miR-17 increased mouse lifespan by repressing expression of IRS1 and ADCY5. Decreased ADCY5 promoted MKP7 expression, resulting in inhibited mTOR and JNK activation, but increased HIF1α expression. Increased HIF1α in turn promoted MKP7 transcription that inhibited cellular senescence and tissue aging. Silencing ADCY5 produced a similar effect as miR-17 transfection, inducing nuclear localization of RGS2 that formed a complex with HIF1α and the MKP7 promoter, enhancing MKP7 transcription, which is a key mediator of inhibitory senescence.
Thus we identified miR-17 as a pleiotropic miRNA that regulates both tumorigenesis and senescence. We found that the processed mature miR-17-3p and miR-17-5p, targeted IRS1 and ADCY5, respectively. Downregulation of both these molecules acted to inhibit cellular senescence, through separate pathways. In particular, ADCY5 repression was found to translocate the membrane-bound RGS2 into the nucleus, which increased MKP7 transcription. This resulted in dephosphorylation of mTOR at Ser2248 and PRAS40 at Thr246, leading to quiescent mTOR activity and reduced cellular senescence.
In addition to describing the oncogenic role of miR-17 in two cellular senescence pathways, we identified MKP7 as a key mediator of mTOR signaling. Identifying and understanding the multiple pathways that miRNAs can target will be important in predicting the effects of therapeutically targeting or introducing single miRNAs.
Material and Methods
Materials
The monoclonal antibodies against ADCY5, pmTOR(S2448), mTOR, pPRAS40 (Thr246), PRAS40, FoxO3a, LC3B, GSK-3β (S9P), IR, PCDNA and pJNK, senescence β-gal staining kit, and SimpleChIP chromatin IP kit were obtained from Cell Signaling (Beverly, MA, USA). Polyclonal antibodies against MKP7, RGS2, pEGFR, EGFR, pERK, ERK2, pMEK1, MEK1, IRS1, Cav2, cyclin D1, and c-myc were obtained from Santa Cruz Biotechnology (Dallas, TX, USA). The monoclonal antibodies against HIF1α, pAKT, and β-actin were obtained from Abcam (San Francisco, CA, USA). Horseradish peroxidase-conjugated goat anti-mouse IgG and horseradish peroxidase-conjugated goat anti-rabbit IgG were obtained from Bio-Rad (Hercules, CA, USA). Immunoblotting was performed using the ECL western blot detection kit (enhanced chemiluminescence; Amersham Biosciences, Amersham, UK). miRNA RT and PCR kits were obtained from Qiagen (Valencia, CA, USA). cAMP assay kit was obtained from R&D (Minneapolis, MN, USA).
Constructs and primers
Plasmid miR-17 and control plasmid were generated previously. Both plasmids contain a pBluescript backbone, a CMV promoter driving green fluorescent protein (GFP) expression and a human H1 promoter driving pre-miR-17 or a non-related sequence serving as a control. The control plasmid was inserted a non-related sequence (5′-ATACAGTACTGTGATAACTGAAGTTTTTGGAAAAGCTTTAGTTATTAA-3′) instead of the pre-miR-17 sequence. To investigate the function of miR-17, the above two constructs were transfected into MEFs. After cell sorting through monitoring GFP signal by FACS, transfected stable cells were obtained.
The pMir-Report (Ambion, Austin, TX, USA), luciferase reporter vector was applied to generate the luciferase constructs. A fragment of the 3′-untranslated region (3′UTR) of mouse ADCY5 and IRS1 was cloned by RT-PCR. Two primers, musADCY5-SacI (5′-CCCGAGCTCTAGCAGATACCAGCCAGCGGTGCC-3′) and musADCY5-MluI (5′-CCACGCGTCAGAAGTTGCTTCTGAGTCAA-3′), were synthesized to clone the 3′UTR fragment of ADCY5, while the 3′UTR fragment of IRS1 was cloned with the primers musIRS1-SacI (5′-CCCGAGCTCACCTACCTTGTGTGTTCTGGGAAC-3′) and musIRS1-MluI (5′-CCACGCGTGTTGATCAGGGCAACTGGGCA-3′). The PCR products were digested with SacI and MluI and the fragments were inserted into the luciferase vector to obtain Luc-ADCY5 and Luc-IRS1. The mutant construct Luc-ADCY5-mut, mutation of the miR-17-5p binding site, was generated by PCR approach with primers musADCY5-SacI and musADCY5-MluI-mut (5′-CCACGCGTCAGAAGTTGCTTCTGAGTGTTAGTG-3′). The PCR products, using primers musIRS1-ScaI and musIRS1-MluI-mut (5′-CCACGCGTGTTGATCAGGGCAACTGGGCATGACCAGTG-3′), were applied to create the Luc-IRS1-mut construct. A non-related sequence, amplified from the coding sequence of the chicken versican G3 domain was used as a negative control as previously described.1
The plasmid vector pCMV6 containing mouse cDNA of Dusp16/MKP7 was obtained from Origene (Rockville, MD, USA), and pBABE containing mouse cDNA fragment of IRS1 was from Addgene (Cambridge, MA, USA). The plasmid vector PCDNA3.1(+)/myc-His B with mouse cDNA of ADCY5 was kindly provided by Dr. Yijuang Chern. Primer sequences used are shown in Supplementary Figure S8f.
Cell survival assay
Cells (2 × 105) were cultured in 10% FBS/DMEM (fetal bovine serum/Dulbecco’s modified Eagle’s medium) medium in culture dishes and maintained at 37°C for 12 h. After cell attachment, the medium was replaced with serum-free DMEM medium or 10% FBS/DMEM medium containing H2O2. Cells were harvested daily or at specific time points and cell number was counted by Coulter Counter (Beckman Coulter, Brea, CA, USA).
Hydrogen peroxide inducing senescence
Cells in basal medium were treated with indicated concentrations of hydrogen peroxide (H2O2) for 2 h. Washing with phosphate-buffered saline (PBS), the treated cultures were incubated in fresh growth medium without H2O2 for 48 h, and processed to β-gal staining.
Annexin V assay
An Annexin V-FITC apoptosis detection kit (Biovision, Milpitas, CA, USA) was used to detect cell apoptosis. Cells were collected and resuspended in binding buffer, and Annexin V-FITC and propidium iodide were added to each sample and incubated in the dark for 15 min. Annexin V-FITC binding was determined by flow cytometry using FITC signal detector (FL1) and propidium staining by the phycoerythrin emission signal detector (FL2).
Intracellular cAMP measurement
Cells were lysed and added to 96-well microtiter plates coated with antiserum of cAMP Enzyme Immunoassay kit (R&D). cAMP levels were measured according to the kit instructions. The optical densities in the plates were determined in a plate reader at 450 nm within 30 min.
Luciferase activity assay
A dual-luciferase reporter system developed by Promega (E1960, Madison, WI, USA) was used to perform luciferase activity assay. Briefly, U343 cells (ATCC, Manassas, VA, USA) were cultured on 12-well tissue culture plates at a density of 2 × 105 cells per well. Cells were co-transfected with the luciferase reporter constructs (Madison, WI, USA), corresponding miRNA mimics, and Renilla luciferase construct (Promega) for 5 h. After 24 h culture, the transfected cells were lysed by 150 μl of passive lysis buffer; 30 μl of lysates were mixed with 50 μl of LAR II, and then firefly luciferase activity was measured by a luminometer. For the internal control, 50 μl of Stop & Glo reagent (Madison, WI, USA) was added to the sample.
Immunofluorescence microscopy
Cells growing on BD culture slides (Franklin Lakes, NJ, USA) were fixed for 10 min in 3.7% formaldehyde, blocked with 10% goat serum, and then incubated with primary antibody in TBS containing 10% goat serum albumin overnight. The slides were washed and stained with goat anti-mouse Alexa 594 or goat anti-rabbit Alexa 647 (Life Technologies, Carlsbad, CA, USA) at room temperature for 1 h. Fluorescent phalloidins, Acti-stain 488 (Cytoskeleton, Denver, CO, USA) was used to stain F-actin to show cell structure. DNA staining was performed with DAPI. Confocal laser scanning microscopy was performed using an LSM 510 Meta microscope (Carl Zeiss, Oberkochen, Germany).
Western blot analysis
Cells were lysed, and the protein samples were subject to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) containing 5–12% acrylamide. Transblotting was processed onto a nitrocellulose membrane in × 1 Tris/glycine buffer containing 20% methanol at 66-V at 4°C for 2 h. The membrane was blocked in TBST (10 mM Tris-Cl, pH 8.0; 150 mM NaCl, 0.05% Tween-20, Sigma-Aldrich, St. Louis, MO, USA) containing 5% non-fat dry milk powder (TBSTM) for 0.5 h, and then incubated with primary antibodies at 4°C overnight. The membranes were washed with TBST (3 × 30 min) and then incubated with secondary antibodies for 1 h. After washing, the bound antibodies were visualized with an ECL detection kit (Amersham Biosciences).
Real-time PCR
For mature miRNA analysis, total RNAs were extracted from cells with mirVana miRNA Isolation Kit (Ambion, San Francisco, CA, USA) according to the manufacturer’s instructions. cDNA synthesis was performed with miScript Reverse Transcription Kit (Qiagen), using 1 μg RNA as template. PCRs were performed with miScriptSYBR GreenPCR Kit (Qiagen) using 1 μl cDNA as template. The primers used as real-time PCR controls were mouse-U6RNAf and mouse-U6RNAr.
Preparation of membrane fractions
Cells were lysed in 2 ml of 2-(N-morpholino) ethanesulfonic acid (MES)-buffered saline (MBS: 25 mM MES; 150 mM NaCl, pH 6.0) with protease inhibitors (PI, Roche Applied Science, Penzberg, Germany), and homogenized with a prechilled Dounce homogenizer (Thomas Scientific, Swedesboro, NJ, USA) with 20 strokes. Samples were centrifuged at 1000 × g for 5 min. The supernatant (2 ml) was adjusted to 45% sucrose by adding 2 ml of 90% sucrose in MBS. Following this, 4 ml 35% sucrose in MBS/Na2CO3 (250 mM) and 4 ml 5% sucrose MBS/Na2CO3 were in turn layered on top of the supernatant. After centrifuged at 39 000 r.p.m. for 16–10 h in a SW41 rotor (Beckman Instruments, Brea, CA, USA), a light-scattering band at the 5–35% sucrose interface was collected. Twelve (1 ml) fractions were collected, starting from the top of the gradient. For immunoblotting, an equal amount of total protein from each fraction (25 μg) was analyzed.
Nuclear extract preparation
Cells were lysed in 1 ml lysis buffer (20 mM Hepes, pH 7.2; 10 mM KCl; 2 mM MgCl2 and PI), and homogenized with a prechilled Dounce homogenizer with 20 strokes. Cell lysis was centrifuged at 4200 r.p.m. for 5 min. The pellet was washed with PBS three times, and resuspended in 100 μl lysis containing 0.5 M NaCl. After centrifugation at 13 200 r.p.m. for 10 min, supernatants containing nuclear extract were used for analysis.
Mitochondria purification preparation
Cells were lysed in 1 ml lysis buffer containing 1 mM DTT and 250 mM sucrose, and homogenized with a prechilled Dounce homogenizer with 20 strokes. Cell lysis was centrifuged at 1000 × g for 10 min. The supernatant was then centrifuged at 10 000 × g for 25 min. The pellet was resuspended in 10 mM Tris-HCl, pH 7.5, 250 mM sucrose and PI, and carefully layered onto a discontinuous sucrose gradient (6 ml 1.0 M sucrose over 6 ml 1.5 M sucrose, dissolved in 20 mM Tris-HCl, pH 7.5; 150 mM NaCl and 1 mM DTT). After ultracentrifugation at 60 000 × g for 30 min, the white band between the 1.0 and 1.5 M sucrose interface was collected and resuspended in a buffer containing 20 mM Tris-HCl, pH 7.5; 150 mM NaCl and PI.
Immunoprecipitation assays
Cells were washed in ice-cold PBS, and lysed in 1 ml lysis buffer. Equal amounts of protein were incubated with 5 μg primary antibody and 40 μl of 50% slurry of protein A-Sepharose (Sigma-Aldrich) at 4°C for 4 h. The pellet was washed three times with PBS and were resuspended in × 2 Laemmli buffer (Sigma-Aldrich, 0.125 M Tris-HCl, 4% SDS, 20% glycerol, 10% 2-mercaptoethanol, and 0.004% bromophenol blue, pH 6.8), followed by western blot analysis.
Chromotin immunoprecipitation assay
Chromotin immunoprecipitation (ChIP) was performed using SimpleChIP chromatin IP kit (Cell Signaling) according to the manufacturer’s instructions. Briefly, cells were treated with formaldehyde solution, and the chromatin was isolated, digested, and immunoprecipitated with antibody against RGS2 or HIF1α. The captured chromatin was eluted, uncrosslinked, and the DNA was recovered. ChIP DNA was subject to PCR using specific primers flanking a piece of DNA sequence of MKP7 promoter region, which has binding sites for RGS2 or HIF1α.
MKP7 dephosphorylation assay
Cells were lysed in 1 ml lysis buffer, and homogenized with a prechilled Dounce homogenizer with 20 strokes. Equal amounts of protein were incubated with 5 μg of anti-mTOR, PRAS40, or MKP7 antibody, and 40 μl of 50% slurry of protein A-Sepharose at 4°C for 4 h. The pellet was washed three times with PBS and were resuspended in 50 μl phosphatase buffer (50 mM Tris-HCl, pH 7.5; 1 mM MgCl2; 0.1 mM EDTA; and 100 μg/ml bovine serum albumin, 25 μg/ml leupeptin) in non-reducing conditions without β-mercaptoethanol. Approximately, 25 μl immunoprecipitated products of mTOR or PRAS40 were incubated with 50 μl immunoprecipitated products of MKP7 at 37°C for 24 h. The reaction was terminated by adding × 2 Laemmli buffer.
Senescence β-galactosidase staining
Staining with β-gal was performed using senescence-galactosidase staining kit (Cell Signaling) according to the manufacturer’s instructions. In brief, cells were fixed for 5 min in 3.7% formaldehyde, washed in PBS and stained in β-gal solution (1 mg/ml 5-bromo-4-chloro-3-indolyl-β-gal) at 37°C until staining become visible in either experiment or control plates. The number of positive cells was counted under a light microscope.
Tissues of wide-type and transgenic mice were freshly excised, rapidly frozen in liquid nitrogen, and mounted in OCT. Sections (5 μm) were placed on glass slides, fixed in 1% formalin in PBS at room temperature for 1 min, and immersed in β-gal staining solution at 37°C for 24 h. The samples were counterstained with eosin, and viewed under a bright field at 10–40 × magnification.
Tissue slide H&E staining, immunohistochemistry
Tissues were freshly excised and fixed in 10% formalin overnight, immersed in 70% ethanol, embedded in wax, and sectioned. The sections were de-paraffinized with xylene and ethanol and then boiled in a pressure cooker. After blocked with 10% goat serum for 1 h, all samples were incubated with primary antibody at 4°C overnight. The sections were incubated with biotinylated secondary antibody for 1 h, followed by avidin-conjugated horseradish peroxidase provided by the Vectastain ABC kit (Vector, PK-4000, Vector Laboratories, Burlingame, CA, USA). The slides were then stained with DAB followed by Mayer’s Hematoxylin (Sigma-Aldrich) counter staining and slide mounting.
Statistical analysis
All experiments were performed in triplicate and numerical data were subject to independent sample t-test. The levels of significance were set at *P<0.05 and **P<0.01.
Abbreviations
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- GFP:
-
green fluorescent protein
- siRNA:
-
small interfering RNA
- miRNA:
-
microRNA
- 3′UTR:
-
3′-untranslated region
- ADCY5:
-
adenylate cyclase type 5
- β-gal:
-
β-galactosidase
- cAMP:
-
cyclic adenosine monophosphate
- DUSP16/MKP7:
-
specificity protein phosphatase 16
- EGFR:
-
epidermal growth factor receptor
- ERK:
-
extracellular signal-regulated kinase
- ECL:
-
enhanced chemiluminescence
- FBS:
-
fetal bovine serum
- FoxO3:
-
Forkhead box O3
- GSK-3β:
-
glycogen synthase kinase-3 β
- GSK-3β (S9P):
-
glycogen synthase kinase-3β serine-9 phosphorylation
- H2O2:
-
hydrogen peroxide
- HIF1α:
-
hypoxia-inducible factor 1-alpha
- IGF-1:
-
insulin-like growth factor 1
- IRS1:
-
insulin receptor substrate 1
- JNK:
-
c-Jun N-terminal kinases
- MEK:
-
MAPK/ERK kinase
- mTOR:
-
mechanistic target of rapamycin
- MEFs:
-
Mouse embryonic fibroblasts
- PCR:
-
polymerase chain reaction
- SDS-PAGE:
-
sodium dodecyl sulfate-polyacrylamide gel electrophoresis
- RGS2:
-
regulator of G-protein signaling 2
References
Fang L, Du WW, Yang W, Rutnam ZJ, Peng C, Li H et al. MiR-93 enhances angiogenesis and metastasis by targeting LATS2. Cell Cycle 2012; 11: 4352–4365.
Mi S, Li Z, Chen P, He C, Cao D, Elkahloun A et al. Aberrant overexpression and function of the miR-17-92 cluster in MLL-rearranged acute leukemia. Proc Natl Acad Sci USA 2010; 107: 3710–3715.
Mu P, Han YC, Betel D, Yao E, Squatrito M, Ogrodowski P et al. Genetic dissection of the miR-17∼92 cluster of microRNAs in Myc-induced B-cell lymphomas. Genes Dev 2009; 23: 2806–2811.
Xiao C, Srinivasan L, Calado DP, Patterson HC, Zhang B, Wang J et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol 2008; 9: 405–414.
Ma Y, Zhang P, Wang F, Zhang H, Yang Y, Shi C et al. Elevated oncofoetal miR-17-5p expression regulates colorectal cancer progression by repressing its target gene P130. Nat Commun 2012; 3: 1291.
Mestdagh P, Bostrom AK, Impens F, Fredlund E, Van Peer G, De Antonellis P et al. The miR-17-92 microRNA cluster regulates multiple components of the TGF-beta pathway in neuroblastoma. Mol Cell 2010; 40: 762–773.
Li H, Yang BB . Stress response of glioblastoma cells mediated by miR-17-5p targeting PTEN and the passenger strand miR-17-3p targeting MDM2. Oncotarget 2012; 3: 1653–1668.
Shan SW, Fang L, Shatseva T, Rutnam ZJ, Yang X, Du W et al. Mature miR-17-5p and passenger miR-17-3p induce hepatocellular carcinoma by targeting PTEN, GalNT7 and vimentin in different signal pathways. J Cell Sci 2013; 126: 1517–1530.
Hackl M, Brunner S, Fortschegger K, Schreiner C, Micutkova L, Muck C et al. miR-17, miR-19b, miR-20a, and miR-106a are down-regulated in human aging. Aging Cell 2010; 9: 291–296.
Johnson SC, Rabinovitch PS, Kaeberlein M . mTOR is a key modulator of ageing and age-related disease. Nature 2013; 493: 338–345.
Laplante M, Sabatini DM . mTOR signaling in growth control and disease. Cell 2012; 149: 274–293.
Cota D, Proulx K, Smith KA, Kozma SC, Thomas G, Woods SC et al. Hypothalamic mTOR signaling regulates food intake. Science 2006; 312: 927–930.
Gleason CE, Lu D, Witters LA, Newgard CB, Birnbaum MJ . The role of AMPK and mTOR in nutrient sensing in pancreatic beta-cells. J Biol Chem 2007; 282: 10341–10351.
Jia K, Chen D, Riddle DL . The TOR pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span. Development 2004; 131: 3897–3906.
Vellai T, Takacs-Vellai K, Zhang Y, Kovacs AL, Orosz L, Muller F . Genetics: influence of TOR kinase on lifespan in C. elegans. Nature 2003; 426: 620.
Cohen E, Paulsson JF, Blinder P, Burstyn-Cohen T, Du D, Estepa G et al. Reduced IGF-1 signaling delays age-associated proteotoxicity in mice. Cell 2009; 139: 1157–1169.
Tatar M, Bartke A, Antebi A . The endocrine regulation of aging by insulin-like signals. Science 2003; 299: 1346–1351.
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R . A C. elegans mutant that lives twice as long as wild type. Nature 1993; 366: 461–464.
Kimura KD, Tissenbaum HA, Liu Y, Ruvkun G . daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science 1997; 277: 942–946.
Ogg S, Paradis S, Gottlieb S, Patterson GI, Lee L, Tissenbaum HA et al. The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 1997; 389: 994–999.
Prentki M, Matschinsky FM . Ca2+, cAMP, and phospholipid-derived messengers in coupling mechanisms of insulin secretion. Physiol Rev 1987; 67: 1185–1248.
Yan L, Vatner DE, O’Connor JP, Ivessa A, Ge H, Chen W et al. Type 5 adenylyl cyclase disruption increases longevity and protects against stress. Cell 2007; 130: 247–258.
Bohni R, Riesgo-Escovar J, Oldham S, Brogiolo W, Stocker H, Andruss BF et al. Autonomous control of cell and organ size by CHICO, a Drosophila homolog of vertebrate IRS1-4. Cell 1999; 97: 865–875.
Copps KD, Hancer NJ, Opare-Ado L, Qiu W, Walsh C, White MF . Irs1 serine 307 promotes insulin sensitivity in mice. Cell Metab 2010; 11: 84–92.
Musikacharoen T, Bandow K, Kakimoto K, Kusuyama J, Onishi T, Yoshikai Y et al. Functional involvement of dual specificity phosphatase 16 (DUSP16), a c-Jun N-terminal kinase-specific phosphatase, in the regulation of T helper cell differentiation. J Biol Chem 2011; 286: 24896–24905.
Shan SW, Lee DY, Deng Z, Shatseva T, Jeyapalan Z, Du WW et al. MicroRNA MiR-17 retards tissue growth and represses fibronectin expression. Nat Cell Biol 2009; 11: 1031–1038.
Robida-Stubbs S, Glover-Cutter K, Lamming DW, Mizunuma M, Narasimhan SD, Neumann-Haefelin E et al. TOR signaling and rapamycin influence longevity by regulating SKN-1/Nrf and DAF-16/FoxO. Cell Metab 2012; 15: 713–724.
Wullschleger S, Loewith R, Hall MN . TOR signaling in growth and metabolism. Cell 2006; 124: 471–484.
Rosner M, Siegel N, Valli A, Fuchs C, Hengstschlager M . mTOR phosphorylated at S2448 binds to raptor and rictor. Amino Acids 2010; 38: 223–228.
Vander Haar E, Lee SI, Bandhakavi S, Griffin TJ, Kim DH . Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 2007; 9: 316–323.
Masuda K, Katagiri C, Nomura M, Sato M, Kakumoto K, Akagi T et al. MKP-7, a JNK phosphatase, blocks ERK-dependent gene activation by anchoring phosphorylated ERK in the cytoplasm. Biochem Biophys Res Commun 2010; 393: 201–206.
Willoughby EA, Perkins GR, Collins MK, Whitmarsh AJ . The JNK-interacting protein-1 scaffold protein targets MAPK phosphatase-7 to dephosphorylate JNK. J Biol Chem 2003; 278: 10731–10736.
Boehm M, Slack F . A developmental timing microRNA and its target regulate life span in C. elegans. Science 2005; 310: 1954–1957.
Boehm M, Slack FJ . MicroRNA control of lifespan and metabolism. Cell Cycle 2006; 5: 837–840.
Ibanez-Ventoso C, Driscoll M . MicroRNAs in C. elegans aging: Molecular insurance for robustness? Curr Genomics 2009; 10: 144–153.
Liu N, Landreh M, Cao K, Abe M, Hendriks GJ, Kennerdell JR et al. The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature 2012; 482: 519–523.
Marino G, Ugalde AP, Fernandez AF, Osorio FG, Fueyo A, Freije JM et al. Insulin-like growth factor 1 treatment extends longevity in a mouse model of human premature aging by restoring somatotroph axis function. Proc Natl Acad Sci USA 2010; 107: 16268–16273.
Kato M, Chen X, Inukai S, Zhao H, Slack FJ . Age-associated changes in expression of small, noncoding RNAs, including microRNAs, in C. elegans. RNA 2011; 17: 1804–1820.
Smith-Vikos T, Slack FJ . MicroRNAs and their roles in aging. J Cell Sci 2012; 125: 7–17.
Salim S, Sinnarajah S, Kehrl JH, Dessauer CW . Identification of RGS2 and type V adenylyl cyclase interaction sites. J Biol Chem 2003; 278: 15842–15849.
De Lorenzo MS, Chen W, Baljinnyam E, Carlini MJ, La Perle K, Bishop SP et al. ‘Reduced malignancy as a mechanism for longevity in mice with adenylyl cyclase type 5 disruption’. Aging Cell 2014; 13: 102–110.
Willoughby EA, Collins MK . Dynamic interaction between the dual specificity phosphatase MKP7 and the JNK3 scaffold protein beta-arrestin 2. J Biol Chem 2005; 280: 25651–25658.
Iglesias-Bartolome R, Patel V, Cotrim A, Leelahavanichkul K, Molinolo AA, Mitchell JB et al. mTOR inhibition prevents epithelial stem cell senescence and protects from radiation-induced mucositis. Cell Stem Cell 2012; 11: 401–414.
Young AR, Narita M . Spatio-temporal association between mTOR and autophagy during cellular senescence. Autophagy 2011; 7: 1387–1388.
Selman C, Lingard S, Choudhury AI, Batterham RL, Claret M, Clements M et al. Evidence for lifespan extension and delayed age-related biomarkers in insulin receptor substrate 1 null mice. FASEB J 2008; 22: 807–818.
Selman C, Partridge L, Withers DJ . Replication of extended lifespan phenotype in mice with deletion of insulin receptor substrate 1. PLoS One 2011; 6: e16144.
Tibarewal P, Zilidis G, Spinelli L, Schurch N, Maccario H, Gray A et al. PTEN protein phosphatase activity correlates with control of gene expression and invasion, a tumor-suppressing phenotype, but not with AKT activity. Sci Signal 2012; 5: ra18.
Acknowledgements
This work was supported by a Discovery Grant from the Natural Sciences and Engineering Research Council of Canada (NSERC; 227937-2012) and grants from Canadian Institutes of Health Research (MOP-102635, MOP-111171) to BBY who is the recipient of a Career Investigator Award (CI 7418) from the Heart and Stroke Foundation of Ontario.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Calin
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Du, W., Yang, W., Fang, L. et al. miR-17 extends mouse lifespan by inhibiting senescence signaling mediated by MKP7. Cell Death Dis 5, e1355 (2014). https://doi.org/10.1038/cddis.2014.305
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.305
This article is cited by
-
Epigenetic changes during ageing and their underlying mechanisms
Biogerontology (2020)
-
MicroRNAs as modulators of longevity and the aging process
Human Genetics (2020)
-
miR-17-92 and miR-106b-25 clusters regulate beta cell mitotic checkpoint and insulin secretion in mice
Diabetologia (2019)
-
Functional similarities of microRNAs across different types of tissue stem cells in aging
Inflammation and Regeneration (2018)
-
Analysis of regulator of G-protein signalling 2 (RGS2) expression and function during prostate cancer progression
Scientific Reports (2018)