Abstract
Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) is an inducer of cancer cell death that holds promise in cancer therapy. Cancer cells are more susceptible than normal cells to the cell-death-inducing effects of TRAIL. However, a variety of cancer cells are resistant to TRAIL through complex mechanisms. Here, we investigate the effects of inhibition of eukaryotic initiation factor 2 subunit α (eIF2α) dephosphorylation on TRAIL-induced apoptosis in hepatoma cells. Treatment of hepatoma cells with salubrinal, an inhibitor of eIF2α dephosphorylation, enhances TRAIL-induced eIF2α phosphorylation, CCAAT/enhancer-binding protein homologous protein (CHOP) expression and caspase activation. Salubrinal enhances TRAIL-induced apoptosis, which could be abrogated by caspase inhibitor. Overexpression of phosphomimetic eIF2α (S51D) enhances TRAIL-induced CHOP expression, caspase 7 and PARP cleavage and apoptosis. By contrast, overexpression of phosphodeficient eIF2α (S51A) abrogates the stimulation of TRAIL-induced apoptosis by salubrinal. Moreover, knockdown of growth arrest and DNA damage-inducible protein 34 (GADD34), which recruits protein phosphatase 1 to dephosphorylate eIF2α, enhances TRAIL-induced eIF2α phosphorylation, CHOP expression, caspase activation and apoptosis. Furthermore, the sensitization of hepatoma cells to TRAIL by salubrinal is dependent on CHOP. Knockdown of CHOP abrogates the stimulation of TRAIL-induced caspase activation and apoptosis by salubrinal. Combination of salubrinal and TRAIL leads to increased expression of Bim, a CHOP-regulated proapoptotic protein. Bim knockdown blunts the stimulatory effect of salubrinal on TRAIL-induced apoptosis. Collectively, these findings suggest that inhibition of eIF2α dephosphorylation may lead to synthetic lethality in TRAIL-treated hepatoma cells.
Similar content being viewed by others
Main
TNF-related apoptosis-inducing ligand (TRAIL) is a member of TNF superfamily that has diverse roles in physiological or pathological conditions.1 A major function of TRAIL is induction of apoptosis. The effects of TRAIL are mediated by cell membrane receptor, including death receptors TRAILR1 (also known as DR4) and TRAILR2 (also known as DR5). Binding of TRAIL to TRAILR1 or TRAILR2 leads to receptor oligomerization and recruitment of the FAS-associated protein with death domain (FADD) to death domain motifs in the carboxyl terminus of the receptor thereby recruiting caspase-8 or caspase-10 to form a multiprotein death-inducing signaling complex.2 By contrast, the decoy receptors TRAILR3 and TRAILR4 may inhibit TRAIL-induced apoptosis.3 TRAIL is a promising anticancer agent, because it can induce the apoptosis of tumor cells.4, 5, 6 Preclinical studies in animal models demonstrate that recombinant TRAIL suppressed some types of human tumor xenografts without apparent systemic toxicity.4, 5
Although TRAIL is an attractive agent that kills cancer cells other than normal cells,7 the sensitivity of cancer cells to TRAIL-induced apoptosis may vary among different types of cancer, subpopulations of a given tumor or individual cancer patients. The mechanisms underlying TRAIL resistance are complex. Resistance to TRAIL can occur as a result of disturbance in the signaling pathways triggered by TRAIL, such as dysfunction of the death receptor DR4 and DR5,8 defects in the FADD and caspase-8 and activation of PI3K/Akt pathway.9, 10, 11 Akt may inhibit Bid expression thereby leading to resistance to TRAIL-induced apoptosis in ovarian cancer cells.12 In addition, cell-surface galectin-3 confers resistance to TRAIL by impeding trafficking of death receptor in metastatic colon adenocarcinoma cells.13 Galectin-3 also sensitizes Bt-549 human breast cancer cells to TRAIL by upregulating PTEN and inactivating Akt.14 On the other hand, overexpression of galectin-3 in human bladder carcinoma cells activates Akt and confers resistance to TRAIL.15
Eukaryotic initiation factor 2 subunit α (eIF2α) has a key role in the process of mRNA translation.16 Phosphorylation of eIF2α reduces its activity, thereby impairing general protein synthesis, whereas increasing the synthesis of certain transcription factors and their targets. Phosphorylation of eIF2α at serine 51 is an important adaptive response to diverse stresses and stimuli such as endoplasmic reticulum (ER) stress, ultraviolet radiation, viral infection, TNFα or TRAIL.17, 18 Previous studies demonstrate that TNF or TRAIL-induced eIF2α phosphorylation is dependent on the double-stranded RNA-regulated protein kinase (PKR).18 As an important part of integrative stress response, phosphorylation of eIF2α may act as a double-edge sword in cell fate decisions. Upon ER stress, phosphorylation and inactivation of eIF2α is a transient process. Initially, phosphorylation of eIF2α may be cytoprotective as a result of reduced burden for the ER or other cellular machinery. Phosphorylation of eIF2α leads to increased synthesis of activating transcription factor 4 (ATF4) thereby increasing the expression of growth arrest and DNA damage-inducible protein 34 (GADD34), which recruits protein phosphatase 1 to eIF2α and dephosphorylates eIF2α.19 This in turn results in the recovery of protein synthesis and restoration of homeostasis. Prolonged or unresolved ER stress, however, can induce cell death, which may involve eIF2α phosphorylation and increased ATF4 and CCAAT/enhancer-binding protein homologous protein (CHOP) expression. Thus, sustained phosphorylation of eIF2α may induce cell death, depending on the cell types or context. Salubrinal, a selective inhibitor of eIF2α dephosphorylation, reportedly inhibits ER stress-induced apoptosis in neural cells.20 However, phosphorylation of eIF2α or treatment with salubrinal enhances proteasome inhibitior-induced apoptosis in leukemia cells and multiple myeloma cells.21, 22, 23 In addition, selective inhibition of eIF2α dephosphorylation causes pancreatic beta-cell dysfunction and apoptosis.24
Hepatoma cells are quite resistant to TRAIL, and TRAIL alone poorly induce apoptotic cell death.25 Given that TRAIL induces eIF2α phosphorylation, we wish to determine the effects of selective inhibition of eIF2α dephosphorylation on TRAIL-induced apoptosis. Here we report that salubrinal or GADD34 knockdown enhances TRAIL-induced hepatoma cell apoptosis in caspase-dependent manner. Treatment with salubrinal leads to an increase in TRAIL-induced eIF2α phosphorylation, CHOP and Bim expression. CHOP or Bim knockdown blunts the stimulation of TRAIL-induced apoptosis by salubrinal.
Results
Salubrinal enhance TRAIL-induced eIF2α phosphorylation and CHOP expression
Previous study indicated that salubrinal induced eIF2α phosphorylation and its downstream CHOP expression without affecting the transcription-dependent branch of the UPR.20 CHOP is one of the components of the ER stress-mediated apoptosis pathway.26 To determine the effects of salubrinal on eIF2α phosphorylation and other UPR elements in hepatoma cells, HepG2 cells were treated with salubrinal ranging from 10 to 100 μ M. The results of western blotting analysis revealed that salubrinal induced the phosphorylation of eIF2α and upregulation of CHOP in a dose-dependent manner, while the expression of ER-resident chaperone GRP78 was not affected (Figure 1a). We also investigated whether TRAIL would induce ER stress or eIF2α phosphorylation in hepatoma cells. HepG2 cells and BEL-7402 cells were treated with different doses of TRAIL for 24 h. The results showed that eIF2α phosphorylation was induced by TRAIL in a dose-dependent manner, while GRP78 expression was unaffected by TRAIL (Figures 1b and c). Compared with HepG2 cells, BEL-7402 cells were more susceptible to TRAIL-induced eIF2α phosphorylation.
Salubirnal enhances TRAIL-induced eIF2α phosphorylation and CHOP expression. (a) HepG2 cells were treated with salubrinal at the indicated dosages for 24 h. Total proteins were harvested and subjected to western blotting analysis of GRP78, P-eIF2α, CHOP and β-actin. (b) HepG2 cells were treated with TRAIL at the indicated dosages for 24 h. Total proteins were harvested and subjected to western blotting analysis of GRP78, P-eIF2α and β-actin. (c) BEL-7402 cells were treated with TRAIL at the indicated dosages for 24 h. Total proteins were harvested and subjected to western blotting analysis of GRP78, P-eIF2α and β-actin. (d) HepG2 cells were treated with 25 μ M salubrinal, 100 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of P-eIF2α, CHOP and β-actin. (e) BEL-7402 cells were treated with 25 μ M salubrinal, 25 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of P-eIF2α, CHOP and β-actin. (f) HepG2 cells were treated with 25 μ M salubrinal, 100 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of DR5 and β-actin
To detect the effect of combination of salubrinal and TRAIL on eIF2α phosphorylation and CHOP expression, HepG2 and BEL-7402 cells were treated with 25 μ M salubrinal, TRAIL or both for 24 h. Combination of salubrinal and TRAIL resulted in significant increase in eIF2α phosphorylation and CHOP expression compared with that in cells treated with salubrinal or TRAIL alone (Figures 1d and e). These results demonstrated that salubrinal enhanced TRAIL-induced eIF2α phosphorylation and CHOP expression.
CHOP reportedly upregulates DR5 expression.27 To detect whether salubrinal affected DR5 expression, HepG2 cells were treated with salubrinal, TRAIL or both for 24 h. Western blotting analysis revealed that the combined treatment with salubrinal and TRAIL did not induce any significant changes in the protein levels of DR5 (Figure 1f). These data indicated that salubrinal did not enhance TRAIL-induced apoptosis through upregulating DR5.
Salubrinal potentiates the inhibition of hepatoma cells survival by TRAIL
Previous studies demonstrated that salubrinal inhibited ER stress-induced apoptosis in neural cells but enhanced proteasome inhibitior-induced apoptosis in leukemia cells and multiple myeloma cells.20, 21, 22, 23 To determine whether combination of salubrinal and TRAIL could synergistically inhibit hepatoma cells survival, HepG2 and BEL-7402 cells were treated with or without salubrinal, TRAIL or both, followed by observing colonies formation 2 weeks after the treatment. Clonogenic assays on HepG2 and BEL-7402 cells demonstrated that salubrinal significantly enhanced the inhibitory effect of TRAIL on cell survival (Figure 2). Whereas treatment with salubrinal alone did not affect colony formation, there were much less colonies formed in cells treated with both salubrinal and TRAIL than that in cells treated with TRAIL alone. Meanwhile, the survived colonies in cells treated with both salubrinal and TRAIL tended to be much smaller than that in cells treated with TRAIL alone (Supplementary Tables S1 and S2).
Salubrinal potentiates the inhibition of hepatoma cell survival by TRAIL. (a) HepG2 cells were seeded in a six-well plate at 2000 cells per well. The next day, the cells were treated with salubrinal (25 μ M), TRAIL (50 ng/ml to 100 ng/ml) or both for 48 h and allowed to grow in the abscense of drugs for 2 weeks. The survived colonies are shown. The relative colony formation is plotted. Cell clonogenicity in the vehicle-treated group was set as 100%. (b) BEL-7402 cells were seeded in a six-well plate at 2000 cells per well. The next day, the cells were treated with salubrinal (25 μ M), TRAIL (50 ng/ml to 100 ng/ml) or both for 48 h and allowed to grow in the abscense of drugs for 2 weeks. The survived colonies are shown. The relative colony formation is plotted. Cell clonogenicity in the vehicle-treated group was set as 100%. Columns, mean of three individual experiments; bars: S.D. A representative of three experiments is shown
Salubrinal potentiates TRAIL-induced caspase activation and apoptosis
To detect whether salubrinal potentiated TRAIL-induced apoptosis, HepG2 and BEL-7402 cells were treated with salubrinal, increasing doses of TRAIL or both for 24 h, followed by Hoechst 33342 staining. Treatment of HepG2 and BEL-7402 cells with 25 μ M salubrinal had little effects on cell apoptosis. Combination of salubrinal and 50 or 100 ng/ml of TRAIL resulted in significantly higher apoptotic rate than TRAIL alone (Figure 3). The accuracy of Hoechst 33324 staining in detecting apoptosis was validated by another assay, terminal nucleotidyl transferase-mediated nick end labeling (TUNEL; Supplementary Figures S1 and S2).
Salubrinal potentiates TRAIL-induced apoptosis. (a) HepG2 cells were treated with salubrinal (25 μ M), TRAIL (50–100 ng/ml) or both for 24 h, and apoptosis was assessed by Hoechst 33342 staining. The apoptotic cells with strong fluorescence, fragmented or condensed nuclei were observed under fluorescent microscopy. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. (b) BEL-7402 cells were treated with salubrinal (25 μ M), TRAIL (12.5–25 ng/ml) or both for 24 h, and apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. A representative of three experiments is shown
Caspases are major effectors of TRAIL-induced apoptosis. To investigate whether salubrinal can enhance TRAIL-induced caspase activation, HepG2 cells were treated with salubrinal, TRAIL or both for 24 h, followed by western blotting analysis of the cleavage of an effector caspase, caspase-7, and its substrate PARP. Treatment with salubrinal or TRAIL alone weakly induced caspase-7 and PARP cleavage. However, combination of salubrinal and TRAIL resulted in a dramatic increase in caspase-7 and PARP cleavage (Figure 4a). Similar effects were detected in BEL-7402 cells (Figure 4b).
Salubrinal stimulates TRAIL-induced poly ADP-ribose polymerase (PARP) cleavage through eIF2α phosphorylation. (a) HepG2 cells were treated with 25 μ M salubrinal, 100 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of cleaved caspase-7, PARP and β-actin. (b) BEL-7402 cells were treated with 25 μ M salubrinal, 25 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of cleaved caspase-7, PARP and β-actin. (c) HepG2 cells were transfected with expression plasmid pEGFP-N1-eIF2α (WT) or eIF2α mutant pEGFP-N1-eIF2α (S51D). Twenty-four hours later, cells were treated with or without 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, eIF2α, cleaved caspase 7 and PARP. In parallel, the apoptosis rate was detected by Hoechst 33342 staining. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. (d) HepG2 cells were transfected with expression plasmid pEGFP-N1- eIF2α (WT) or eIF2α mutant (pEGFP-N1-eIF2α (S51A). Twenty-four hours later, cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, eIF2α, cleaved caspase 7 and PARP. In parallel, the apoptosis rate was detected by Hoechst 33342 staining. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D.
To determine whether eIF2α mutant (S51D), a mimic of phosphorylated eIF2α (S51), has a similar effect to salubrinal, HepG2 cells were transfected with wild-type eIF2α or eIF2α mutant (S51D) plasmid, followed by treatment with or without TRAIL. Overexpression of eIF2α mutant (S51D) enhanced TRAIL-induced CHOP expression, caspase 7 and PARP cleavage and apoptosis (Figure 4c and Supplementary Figure S3). These data suggest that eIF2α mutant (S51D) can mimic the effect of salubrinal on TRAIL-induced apoptosis.
To further determine the role of eIF2α phosphorylation in salubrinal’s sensitization of hepatoma cells to TRAIL-induced apoptosis, we transfected wild-type eIF2α or eIF2α S51A allele, a phosphorylation-deficient mutant of eIF2α, into HepG2 cells, followed by treatment with or without salubrinal and TRAIL. Overexpression of phosphorylation-deficient eIF2α inhibited TRAIL-induced CHOP expression, caspase 7 and PARP cleavage and abrogated the stimulatory effect of salubrinal on TRAIL-induced CHOP expression, caspase 7 and PARP cleavage and apoptosis (Figure 4d and supplementary Figure S4). These data suggest that the stimulation of TRAIL-induced apoptosis by salubrinal is due, at least in part, to inhibition of eIF2α dephosphorylation.
Additionally, treatment with caspase inhibitor abrogated the effects of TRAIL and salubrinal on apoptosis in both HepG2 and BEL-7402 cells (Figure 5). These data indicated that salubrinal potentiated TRAIL-induced apoptosis through caspase activation.
Salubrinal stimulates TRAIL-induced apoptosis in a caspase-dependent manner. (a) HepG2 cells were treated with or without 25 μ M salubrinal, 100 ng/ml TRAIL and 20 μ M z-VAD-fmk for 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. (b) BEL-7402 cells were treated with or without 25 μ M salubrinal, 25 ng/ml TRAIL and 20 μ M z-VAD-fmk for 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. A representative of three experiments is shown
GADD34 knockdown sensitizes hepatoma cells to TRAIL
Earlier reports indicated that GADD34 played an important role in the UPR.19, 28 GADD34 interacted with protein phosphatase 1 to dephosphorylate eIF2α, leading to a negative feedback loop and translational recovery, which mitigated cellular stress.19, 28, 29 Salubrinal selectively inhibited the GADD34/PP1-mediated dephosphorylation of eIF2α.20 To detect whether GADD34 knockdown potentiated TRAIL-induced eIF2α phosphorylation, CHOP expression, caspase 7 and PARP cleavage, HepG2 and BEL-7402 cells were transfected with the control or GADD34 siRNA, followed by treatment with or without TRAIL for 24 h. GADD34 knockdown did not induce eIF2α phosphorylation and CHOP expression but promoted TRAIL-induced eIF2α phosphorylation, CHOP expression,caspase-7 and PARP cleavage (Figures 6a and b). These data indicated that GADD34 knockdown could potentiate the induction of eIF2α phosphorylation, CHOP expression and caspase-7 activation by TRAIL.
GADD34 knockdown sensitizes hepatoma cells to TRAIL. (a) HepG2 cells were transfected with negative control siRNA (siCtrl) or siRNA to GADD34 (siGADD34). Forty-eight hours later, the cells were treated with 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of P-eIF2α, CHOP, cleaved caspase-7, PARP, GADD34 and β-actin. (b) BEL-7402 cells were transfected with negative control siRNA (siCtrl) or siRNA to GADD34 (siGADD34). Forty-eight hours later, the cells were treated with 25 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of P-eIF2α, CHOP, cleaved caspasse-7, PARP, GADD34 and β-actin. (c) HepG2 cells were transfected with negative control siRNA (siCtrl) or siRNA to GADD34 (siGADD34). Forty-eight hours after transfection, the cells were treated with or without 100 ng/ml TRAIL and 20 μ M z-VAD-fmk for 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars: S.D. (d) BEL-7402 cells were transfected with negative control siRNA (siCtrl) or siRNA to GADD34 (siGADD34). Forty-eight hours after transfection, the cells were treated with or without 25 ng/ml TRAIL and 20 μ M z-VAD-fmk for 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D.
Next, we investigated the effects of GADD34 knockdown on TRAIL-induced apoptosis. Knockdown of GADD34 significantly enhanced TRAIL-induced apoptosis in both HepG2 and BEL-7402 cells. Treatment with caspase inhibitor abrogated the effects of TRAIL and GADD34 knockdown on apoptosis (Figures 6c and d). These data suggest that GADD34 knockdown could sensitize hepatoma cells to TRAIL by promoting caspase activation.
Upregulation of CHOP contributes to the enhancement of TRAIL-induced apoptosis by salubrinal
To determine the functional significance of salubrinal-induced CHOP upregulation, we investigated whether downregulation of CHOP by small interfering RNA (siRNA) could inhibit the synergistic effects of salubrinal and TRAIL on hepatoma cells. HepG2 cells were transfected with the control or CHOP siRNA, followed by treatment with salubrinal, TRAIL or both for 24 h. CHOP knockdown led to significant decrease in apoptosis induced by combination of salubrinal and TRAIL but had little effects on cells treated with TRAIL alone (Figures 7a and b). Knockdown of CHOP in HepG2 cells significantly inhibited caspase 7 and PARP cleavage induced by the combined treatment with salubrinal and TRAIL (Figure 7c). CHOP knockdown in BEL-7402 cells also led to significant decrease in caspase 7 and PARP cleavage and apoptosis induced by combination of salubrinal and TRAIL (Figure 8). Taken together, these results suggested that salubrinal-induced CHOP expression was indispensable for enhancing TRAIL sensitivity in hepatoma cells.
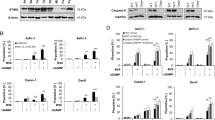
Upregulation of CHOP by salubrinal contributes to the enhancement of TRAIL-induced apoptosis in HepG2 cells. (a) HepG2 cells were transfected with negative control siRNA (siCtrl) or siRNA to CHOP (siCHOP). Forty-eight hours later, HepG2 cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Apoptosis was assessed by Hoechst 33342 staining. (b) The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars: S.D. (c) HepG2 cells were transfected with negative control siRNA (siCtrl) or siRNA to CHOP (siCHOP). Forty-eight hours later, HepG2 cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, caspase-7, PARP and β-actin
Upregulation of CHOP by salubrinal contributes to the enhancement of TRAIL-induced apoptosis in BEL-7402 cells. (a) BEL-7402 cells were transfected with negative control siRNA (siCtrl) or siRNA to CHOP (siCHOP). Forty-eight hours later, BEL-7402 cells were treated with or without 25 μ M salubrinal and 25 ng/ml TRAIL for further 24 h. Apoptosis was assessed by Hoechst 33342 staining. (b) The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars, S.D. (c) BEL-7402 cells were transfected with negative control siRNA (siCtrl) or siRNA to CHOP (siCHOP). Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 25 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, caspase-7, PARP and β-actin
Bim is a CHOP-regulated proapoptotic protein that mediates the stimulatory effect of salubrinal on TRAIL-induced apoptosis
Previous study suggests that Bim is a CHOP-regulated proapoptotic protein that mediates ER stress-induced apoptosis.30 To determine the effect of salubrinal and TRAIL on Bim expression, HepG2 and BEL-7402 cells were treated with or without salubrinal and TRAIL, followed by western blotting analysis of Bim expression. Combination of salubrinal and TRAIL significantly upregulated Bim expression in both cell lines (Figures 9a and b). In addition, CHOP knockdown abrogated the upregulation of Bim by salubrinal and TRAIL (Figures 9c and d), indicating that salubrinal and TRAIL induced Bim through CHOP.
Combination of salubrinal and TRAIL leads to CHOP-dependent upregulation of Bim. (a) HepG2 cells were treated with 25 μ M salubrinal, 100 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of Bim and β-actin. (b) BEL-7402 cells were treated with 25 μ M salubrinal, 25 ng/ml TRAIL or both for 24 h. Total proteins were harvested and subjected to western blotting analysis of Bim and β-actin. (c) HepG2 cells were transfected with siCtrl or siRNA to CHOP (siCHOP). Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, Bim and β-actin. (d) BEL-7402 cells were transfected with siCtrl or siRNA to CHOP (siCHOP). Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 25 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of CHOP, Bim and β-actin
Finally, we investigated whether Bim mediated the stimulatory effect of salubrinal on TRAIL-induced apoptosis. Western blotting analysis demonstrated that Bim knockdown blunted the stimulatory effect of salubrinal on TRAIL-induced PARP cleavage in both HepG2 and BEL-7402 cells (Figures 10a and b). Moreover, Bim knockdown abrogated the enhancement of TRAIL-induced apoptosis by salubrinal in HepG2 and BEL-7402 cells (Figures 10c and d). These data demonstrate that Bim mediates the stimulatory effect of salubrinal on TRAIL-induced apoptosis.
Bim knockdown blunts the stimulatory effect of salubrinal on TRAIL-induced apoptosis. (a) HepG2 cells were transfected with siCtrl or siRNA to Bim (siBim). Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of Bim expression and PARP cleavage. (b) BEL-7402 cells were transfected with siCtrl or siRNA to Bim (siBim). Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 25 ng/ml TRAIL for further 24 h. Total proteins were harvested and subjected to western blotting analysis of Bim expression and PARP cleavage. (c) HepG2 cells were transfected with negative control siCtrl or siBim. Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 100 ng/ml TRAIL for further 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars: S.D. (d) BEL-7402 cells were transfected with negative control siCtrl or siBim. Forty-eight hours later, the cells were treated with or without 25 μ M salubrinal and 25 ng/ml TRAIL for further 24 h. Apoptosis was assessed by Hoechst 33342 staining. The apoptosis rate is plotted. Columns, mean percentage of strongly Hoechst-positive cells; bars: S.D.
Discussion
TRAIL is one of the apoptosis-inducing proteins that have been tested in treating tumors.31 As an extracellular signal, TRAIL can elicit oxidative stress that mediates TRAIL-induced apoptosis.32 RNAi-based screening has revealed that some genes can either inhibit or enhance TRAIL-induced apoptosis.33 Previous study demonstrates that TRAIL induces eIF2α phosphorylation, a modifier of global protein synthesis. However, it is unclear how eIF2α phosphorylation would affect TRAIL-induced apoptosis. In this study, we show that inhibition of eIF2α dephosphorylation enhances TRAIL-induced apoptosis. Salubrinal, an inhibitor of GADD34-PP1c complex that is responsible for eIF2α dephosphorylation, enhances TRAIL-induced eif2α phosphorylation and apoptosis. Similar effects are observed in GADD34-knockdown cells.
It remains unclear how TRAIL induces eIF2α phosphorylation. Although TRAIL may induce reactive oxygen species (ROS) accumulation and ROS is one of the inducers of eIF2α phosphorylation,34 treatment with antioxidant does not prevent TRAIL-induced eIF2α phosphorylation (data not shown). eIF2α phosphorylation is also a component of integrated response to ER stress that usually induces GRP78, an ER-resident chaperone that has critical roles in physiological and pathological stress.35 However, TRAIL induces eIF2α phosphorylation without upregulating GRP78 expression. Thus, it is less likely that TRAIL induces eIF2α phosphorylation as a result of ER stress. Phosphorylation of eIF2α represents a kind of cellular stress response leading to reduced protein synthesis. Reduced translation would create a time window to repair cell damage or recover from stress. From this point of view, phosphorylation of eIF2α is cytoprotective. However, prolonged shutdown of protein synthesis would also be deleterious, because it may break cellular homeostasis. Therefore, eIF2α phosphorylation usually couples with induction of GADD34 expression, which in turn recruits PP1c to dephosphorylate eIF2α and restores protein synthesis. In this study, TRAIL upregulated GADD34 expression in BEL-7402 cells but not in HepG2 cells, possibly owing to the relatively high basic levels of GADD34 and to the relatively low induction of eIF2α phosphorylation by TRAIL in HepG2 cells.
Although salubrinal protects cells from ER stress-induced apoptosis in neural cells, it sensitizes leukemia cells to ER stressor and proteasome inhibition.22 Also, the pro-apoptotic function of eIF2α phosphorylation in response to TRAIL and proteasome inhibition contrasts with the anti-apoptotic activities of eIF2α kinases described for other stress conditions. For example, treatment of PEK−/− MEF cells with the ER stress inducer thapsigargin led to increased apoptosis compared with its wild-type counterpart.36 Loss of GCN2 leads to elevated apoptosis in MEF cells following exposure to UV irradiation.37 GCN2 phosphorylation of eIF2α in response to UV irradiation facilitates activation of NF-κB but does not lead to detectable accumulation of the pro-apoptotic factor CHOP. These studies suggest that the effects of salubrinal on stress-induced apoptosis may be cell type-specific or context-dependent. The primary eIF2α kinase activated by proteasome inhibition is GCN2, while the primary eIF2α kinase activated by TRAIL is PKR.21, 38 Loss of eIF2α phosphorylation contributes to reduced levels of caspase activation and diminished apoptosis in response to MG132 treatment, while inhibition of eIF2α dephosphorylation by salubrinal leads to increased caspase activation and apoptosis in response to proteasome inhibition.22 Similar to proteasome inhibitors, inhibition of eIF2α dephosphorylation leads to increased apoptosis in response to TRAIL via caspase-dependent mechanism. Another study by Moon et al.39 also demonstrated that salubrinal could potentiate apoptosis induced by a combination of guggulsterone and TRAIL via DR5 upregulation, while the effect of salubrinal on apoptosis induced by TRAIL alone was not reported.
Induction of CHOP function is one important reason for the pro-apoptotic function of eIF2α kinases or salubrinal during proteasome inhibition. Inhibition of eIF2α dephosphorylation also enhances TRAIL-induced CHOP expression. Although knockdown of CHOP does not prevent TRAIL-induced apoptosis, it abrogates the potentiation of TRAIL-induced apoptosis by salubrinal. These data suggest that CHOP is responsible for the stimulation of TRAIL-induced apoptosis by salubrinal. The mechanism underlying CHOP-induced apoptosis is complex. CHOP may induce cell death by promoting protein synthesis and oxidation in the stressed ER.40 In addition, studies examining the mechanism of CHOP-induced apoptosis identified numerous target genes, including Bcl2, GADD34, ER oxidoreductin 1 (ERO1α), tribbles-related protein 3 (TRB3) and Bim.30, 41 Although CHOP mainly upregulates gene expression, Bcl2 is otherwise downregulated by CHOP.42 Also, CHOP reportedly enhances DR5 expression.27 However, neither salubrinal nor combination of salubrinal and TRAIL affects DR5 expression. Thus, salubrinal does not potentiate TRAIL-induced apoptosis by upregulating DR5 expression.
This study demonstrates that maintaining eIF2α phosphorylation through drugs like salubrinal can potentiate killing of hepatoma cells by TRAIL. We therefore propose that salubrinal or similar compounds affecting phosphorylation of eIF2α may serve as a potential strategy to maximize TRAIL efficiency and decrease tumor cell resistance to this agent. Except for bortezomib and TRAIL, it would be interesting to test whether salubrinal can enhance the therapeutic effects of other anti-cancer agents.
Materials and Methods
Reagents and antibodies
Salubrinal was purchased from Enzo Life Sciences Inc. (San Diego, CA, USA) and dissolved in DMSO to 10 μ M and stored at −20 °C. Recombinant Human TRAIL/Apo2L was purchased from PeproTech Inc. (Rocky Hill, NJ, USA) and prepared by reconstituting in deionized water and by diluting in Dulbecco’s Minimal Essential Medium (DMEM) to 50 μg/ml and stored at −20 °C. The caspase inhibitor z-VAD-fmk was purchased from Beyotime Institute of Biotechnology (Nantong, Jiangsu, China). Anti-GRP78, anti-DR5, anti-phosphorylated eIF2α, anti-CHOP, anti-caspase-7 and anti-PARP antibodies were purchased from Cell Signaling Technology (Beverly, MA, USA). Anti-GADD34 antibody was purchased from GeneTex Inc. (Irvine, CA, USA). Anti-Bim antibody was purchased from Epitomics Inc. (Burlingame, CA,USA). Anti-β-actin antibody was purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA).
Plasmid and cloning
The EGFP-tagged expression plasmids for eIF2α and its mutants (pEGFP-N1-eIF2α; pEGFP-N1-eIF2α-S51D; pEGFP-N1-eIF2α-S51A) were generated by PCR subcloning of the corresponding eIF2α cDNA in expression plasmids kindly provided by Dr. Roland M. Schmid (Department of Medicine 2, Technical University of Munich, Munich, Germany) into pEGFP-N1. Integrity of subcloned sequences was verified by automated DNA sequencing (GENEWIZ, Suzhou, China).
Cell culture
Hepatoma cell lines HepG2 and BEL-7402 were obtained from Cell Lines Bank, Chinese Academy of Science (Shanghai, China). The cells were maintained in DMEM containing 10% fetal bovine serum, 100 U/ml of penicillin and 100 μg/ml streptomycin sulfate and incubated at 37 °C in a humidified atmosphere of 5% CO2.
Transient transfection
For transfection, 40 000 cells were plated into six-well plates and incubated for 1 day. LipofecTAMINE 2000 reagent (Invitrogen, Carlsbad, CA, USA) was diluted in 250 μl of Opti-MEM I reduced serum medium and incubated at room temperature for 5 min. In addition, pEGFP-N1-eIF2α (S51D) was diluted in 250 μl of Opti-MEM I reduced serum medium and mixed with the pre-diluted LipofecTAMINE 2000. The mixture was incubated at room temperature for 20 min. 4 μg Plasmid was added into each well and incubated at 37° C. Cells were transfected at a density of 70–80% and used for experiments 24 h after transfection.
RNA interference
The target sequences used for knockdown of CHOP, GADD34 and Bim were 5'-GCCUGGUAUGAGGACCUGC-3′, 5′-GGACACUGCAAGGUUCUGA-3′, 5′-CAACCACUAUCUCAGUGCA-3′, respectively. The negative control siRNA was purchased from Ribobio Co., Ltd. (Guangzhou, China). The double-stranded siRNA duplex was dissolved in DEPC-treated water. For transfection, 40000 cells were plated into 6-well plates and incubated for one day. LipofecTAMINE 2000 reagent (Invitrogen) was diluted in 250 μl of Opti-MEM I reduced serum medium and incubated at room temperature for 5 min. In addition, siRNA duplex was diluted in 250 μl of Opti-MEM I reduced serum medium and mixed with the pre-diluted LipofecTAMINE 2000. The mixture was incubated at room temperature for 20 min. Inall, 50 nM of siRNA was added into each well and incubated at 37 °C.
Western blotting
Cells were washed twice with phosphate-buffered saline and harvested with cold RIPA lysis buffer containing protease inhibitors. Cell lysates were collected from culture plates using a rubber policeman, and protein was collected by centrifugation. Protein concentrations were determined by bicinchoninic acid protein assay (Pierce Biotechnology, Rockford, IL, USA). Thirty micrograms of proteins were boiled in 4 × loading buffer for 10 min, then loaded into 10% Tris-HCl-Polyacrylamide gels and transferred to PVDF membrane (Millipore Corporation, Billerica, MA, USA). Membranes were incubated with primary antibodies and appropriate HRP-secondary antibodies. Detection was performed with chemiluminescent agents.
Clonogenic assay
To determine the effect of salubrinal and TRAIL on the clonogenicity, hepatoma cells were plated in six-well plates at 2000 cells per well. After 24 h, the cells were treated with or without salubrinal and TRAIL for 48 h. After being rinsed with fresh medium, cells were allowed to grow in the absence of drugs for 14 days to form colonies. The cells were fixed and stained with Giemsa dye liquor to visualize cell colonies.
Cell apoptosis assay
Cell apoptosis was assessed by Hoechst 33342 or TUNEL assay (Beyotime Institute of Biotechnology). For Hoechst 33342 staining, replicate cultures of hepatoma cells were plated in tissue culture plates. The cells were treated with or without salubrinal and TRAIL for 24 h and then incubated with Hoechst 33342 solution at 4 °C for 10 min, followed by examining under a fluorescence microscope (Leica Microsystems Gmbh, Wetzlar, Germany). High blue fluorescence and condensed or fragmented nuclei can be observed in apoptotic cells, while weak fluorescence was observed in live cells. Quantification of apoptotic cells was performed by taking the images in random fields and counting at least 200 cells in four random fields in each well.
For TUNEL assay, replicate cultures of hepatoma cells were plated in tissue culture plates. The next day, the cells were treated with or without salubrinal and TRAIL for 24 h. Then the cells were fixed in 4% paraformaldehyde for 60 min. Next, the cells were permeabilized using 0.25% Triton X-100 in PBS for 2 min, followed by incubating with TUNEL solution and DAPI solution at 37 °C for 60 min without light. The apoptotic cells were examined under a fluorescence microscope. Green fluorescence can be observed in the nuclei of apoptotic cells. All cells show blue fluorescence with DAPI. Quantification of all the cells and apoptotic cells in the same fields was performed by taking the images in random fields and counting at least 200 cells in four random fields in each well.
Statistical analysis
One-way ANOVA with least significant difference post hoc test was used to test for the differences in cell viability and apoptosis rate. All statistical tests were two-tailed, and difference was considered statistically significant when P<0.05.
Abbreviations
- ATF4:
-
activating transcription factor 4
- CHOP:
-
CCAAT/enhancer-binding protein homologous protein
- eIF2α:
-
eukaryotic initiation factor 2 subunit α
- ER:
-
endoplasmic reticulum
- GADD34:
-
growth arrest and DNA damage-inducible protein 34
- TRAIL:
-
tumor necrosis factor-related apoptosis-inducing ligand
References
Johnstone RW, Frew AJ, Smyth MJ . The TRAIL apoptotic pathway in cancer onset, progression and therapy. Nat Rev Cancer 2008; 8: 782–798.
Falschlehner C, Emmerich CH, Gerlach B, Walczak H . TRAIL signalling: decisions between life and death. Int J Biochem Cell Biol 2007; 39: 1462–1475.
Ashkenazi A, Dixit VM . Apoptosis control by death and decoy receptors. Curr Opin Cell Biol 1999; 11: 255–260.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor–related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Wang S . The promise of cancer therapeutics targeting the TNF-related apoptosis-inducing ligand and TRAIL receptor pathway. Oncogene 2008; 27: 6207–6215.
Bastiaens P . Systems biology: when it is time to die. Nature 2009; 459: 334–335.
Thomas LR, Johnson RL, Reed JC, Thorburn A . The C-terminal tails of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) and Fas receptors have opposing functions in Fas-associated death domain (FADD) recruitment and can regulate agonist-specific mechanisms of receptor activation. J Biol Chem 2004; 279: 52479–52486.
Kuang AA, Diehl GE, Zhang J, Winoto A . FADD is required for DR4-and DR5-mediated apoptosis lACK of TRAIL-induced apoptosis in FADD-deficient mouse embryonic fibroblasts. J Biol Chem 2000; 275: 25065–25068.
Rehemtulla A, Ross BD . A review of the past, present, and future directions of neoplasia. Neoplasia 2005; 7: 1039–1046.
Larribere L, Khaled M, Tartare-Deckert S . PI3K mediates protection against TRAIL-induced apoptosis in primary human melanocytes. Cell Death Differ 2004; 11: 1084–1091.
Goncharenko-Khaider N, Lane D, Matte I, Rancourt C, Piché A . The inhibition of Bid expression by Akt leads to resistance to TRAIL-induced apoptosis in ovarian cancer cells. Oncogene 2010; 29: 5523–5536.
Mazurek N, Byrd JC, Sun Y, Hafley M, Ramirez K, Burks J et al. Cell-surface galectin-3 confers resistance to TRAIL by impeding trafficking of death receptors in metastatic colon adenocarcinoma cells. Cell Death Differ 2012; 19: 523–533.
Mazurek N, Sun YJ, Liu KF, Gilcrease MZ, Schober W, Nangia-Makker P et al. Phosphorylated galectin-3 mediates tumor necrosis factor-related apoptosis-inducing ligand signaling by regulating phosphatase and tensin homologue deleted on chromosome 10 in human breast carcinoma cells. J Biol Chem 2007; 282: 21337–21348.
Oka N, Nakahara S, Takenaka Y, Fukumori T, Hogan V, Kanayama H et al. Galectin-3 inhibits tumor necrosis factor–related apoptosis-inducing ligand–induced apoptosis by activating Akt in human bladder carcinoma cells. Cancer Res 2005; 65: 7546–7553.
Sánchez I, Dynlacht BD . New insights into cyclins, CDKs, and cell cycle control. Semin Cell Dev Biol 2005; 16: 311–321.
Deng J, Harding HP, Raught B, Gingras AC, Berlanga JJ, Scheuner D et al. Activation of GCN2 in UV-irradiated cells inhibits translation. Curr Biol 2002; 12: 1279–1286.
Jeffrey IW, Bushell M, Tilleray VJ, Morley S, Clemens MJ . Inhibition of protein synthesis in apoptosis differential requirements by the tumor necrosis factor α family and a DNA-damaging agent for caspases and the double-stranded RNA-dependent protein kinase. Cancer Res 2002; 62: 2272–2280.
Novoa I, Zeng H, Harding HP, Ron D . Feedback inhibition of the unfolded protein response by GADD34-mediated dephosphorylation of eIF2α. J Cell Biol 2001; 153: 1011–1022.
Boyce M, Bryant KF, Jousse C, Long K, Harding HP, Scheuner D et al. A selective inhibitor of eIF2α dephosphorylation protects cells from ER stress. Science 2005; 307: 935–939.
Jiang HY, Wek RC . Phosphorylation of the α-subunit of the eukaryotic initiation factor-2 (eIF2α) reduces protein synthesis and enhances apoptosis in response to proteasome inhibition. J Biol Chem 2005; 280: 14189–14202.
Drexler HC . Synergistic apoptosis induction in leukemic cells by the phosphatase inhibitor salubrinal and proteasome inhibitors. PLoS One 2009; 4: e4161.
Schewe DM, Aguirre-Ghiso JA . Inhibition of eIF2α dephosphorylation maximizes bortezomib efficiency and eliminates quiescent multiple myeloma cells surviving proteasome inhibitor therapy. Cancer Res 2009; 69: 1545–1552.
Cnop M, Ladriere L, Hekerman P, Ortis F, Cardozo AK, Dogusan Z et al. Selective inhibition of eukaryotic translation initiation factor 2α dephosphorylation potentiates fatty acid-induced endoplasmic reticulum stress and causes pancreatic β-cell dysfunction and apoptosis. J Biol Chem 2007; 282: 3989–3997.
Chen KF, Yeh PY, Hsu C, Hsu CH, Lu YS, Hsieh HP et al. Bortezomib overcomes tumor necrosis factor-related apoptosis-inducing ligand resistance in hepatocellular carcinoma cells in part through the inhibition of the phosphatidylinositol 3-kinase/Akt pathway. J Biol Chem 2009; 284: 11121–11133.
Oyadomari S, Mori M . Roles of CHOP/GADD153 in endoplasmic reticulum stress. Cell Death Differ 2003; 11: 381–389.
Yamaguchi H, Wang HG . CHOP is involved in endoplasmic reticulum stress-induced apoptosis by enhancing DR5 expression in human carcinoma cells. J Biol Chem 2004; 279: 45495–45502.
Novoa I, Zhang Y, Zeng H, Jungreis R, Harding HP, Ron D . Stress-induced gene expression requires programmed recovery from translational repression. EMBO J 2003; 22: 1180–1187.
Brush MH, Weiser DC, Shenolikar S . Growth arrest and DNA damage-inducible protein GADD34 targets protein phosphatase 1α to the endoplasmic reticulum and promotes dephosphorylation of the α subunit of eukaryotic translation initiation factor 2. Mol Cell Biol 2003; 23: 1292–1303.
Puthalakath H, O'Reilly LA, Gunn P, Lee L, Kelly PN, Huntington ND et al. ER stress triggers apoptosis by activating BH3-only protein Bim. Cell 2007; 129: 1337–1349.
Wang S . TRAIL: a sword for killing tumors. Curr Med Chem 2010; 17: 3309–3317.
Lee MW, Park SC, Kim JH, Kim IK, Han KS, Kim KY et al. The involvement of oxidative stress in tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL)-induced apoptosis in HeLa cells. Cancer Lett 2002; 182: 75–82.
Aza-Blanc P, Cooper CL, Wagner K, Batalov S, Deveraux QL, Cooke MP . Identification of modulators of TRAIL-induced apoptosis via RNAi-based phenotypic screening. Mol Cell 2003; 12: 627–637.
Loghlen AO, Perez-Morgado MI, Salinas M, Martın ME . Reversible inhibition of the protein phosphatase 1 by hydrogen peroxide. Potential regulation of eIF2α phosphorylation in differentiated PC12 cells. Arch Biochem Biophys 2003; 417: 194–202.
Pfaffenbach KT, Lee AS . The critical role of GRP78 in physiologic and pathologic stress. Curr Opin Cell Biol 2011; 23: 150–156.
Harding HP, Zhang Y, Bertolotti A, Zeng H, Ron D . Perk is essential for translational regulation and cell survival during the unfolded protein response. Mol cell 2000; 5: 897–904.
Jiang HY, Wek RC . GCN2 phosphorylation of eIF2α activates NF-κB in response to UV irradiation. Biochem J 2005; 385: 371–380.
Clemens MJ . Targets and mechanisms for the regulation of translation in malignant transformation. Oncogene 2004; 23: 3180–3188.
Moon DO, Park SY, Choi YH, Ahn JS, Kim GY . Guggulsterone sensitizes hepatoma cells to TRAIL-induced apoptosis through the induction of CHOP-dependent DR5: involvement of ROS-dependent ER-stress. Biochem Pharmacol 2011; 82: 1641–1650.
Marciniak SJ, Yun CY, Oyadomari S, Novoa I, Zhang Y, Jungreis R et al. CHOP induces death by promoting protein synthesis and oxidation in the stressed endoplasmic reticulum. Genes Dev 2004; 18: 3066–3077.
Szegezdi E, Logue SE, Gorman AM, Samali A . Mediators of endoplasmic reticulum stress-induced apoptosis. EMBO Rep 2006; 7: 880–885.
McCullough KD, Martindale JL, Klotz LO, Aw TY, Holbrook NJ . Gadd153 sensitizes cells to endoplasmic reticulum stress by down-regulating Bcl2 and perturbing the cellular redox state. Mol Cell Biol 2001; 21: 1249–1259.
Acknowledgements
We thank Dr. Roland M Schmid for the eIF2α mutant (S51D and S51A) plasmids. This work was supported in part by the National Natural Science Foundation of China (No. 81272215; 81001587; 30900554).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by A Oberst
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Teng, Y., Gao, M., Wang, J. et al. Inhibition of eIF2α dephosphorylation enhances TRAIL-induced apoptosis in hepatoma cells. Cell Death Dis 5, e1060 (2014). https://doi.org/10.1038/cddis.2014.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.24
Keywords
This article is cited by
-
Inhibition of PPP1R15A alleviates osteoporosis via suppressing RANKL-induced osteoclastogenesis
Acta Pharmacologica Sinica (2024)
-
Heme-regulated inhibitor: an overlooked eIF2α kinase in cancer investigations
Medical Oncology (2022)
-
Sephin1 Reduces Prion Infection in Prion-Infected Cells and Animal Model
Molecular Neurobiology (2020)
-
Targeting KPNB1 overcomes TRAIL resistance by regulating DR5, Mcl-1 and FLIP in glioblastoma cells
Cell Death & Disease (2019)
-
Inhibition of eIF2α dephosphorylation accelerates pterostilbene-induced cell death in human hepatocellular carcinoma cells in an ER stress and autophagy-dependent manner
Cell Death & Disease (2019)