Abstract
Pro-survival signalling mediated by the androgen receptor (AR) is implicated as a key contributor to prostate carcinogenesis. As prostate tumours are characterized by nutrient-poor, hypoxic and acidified microenvironments, one mechanism whereby AR signalling may contribute to survival is by promoting adaptation to cellular stress. Here we have identified a novel role for AR in the inhibition of autophagy induced by serum withdrawal. This blockade is attributed to AR-mediated upregulation of the endoplasmic reticulum (ER) chaperone glucose-regulated protein 78/BiP (Grp78/BiP), and occurs independently of ER stress response pathway activation. Interestingly, AR activation did not affect serum starvation-induced mammalian target of rapamycin inhibition, illustrating that the adaptive role for androgens lies not in the ability to modulate nutrient sensing, but in the promotion of ER stability. Finally, we show that the adaptive advantage conferred by AR-mediated Grp78/BiP upregulation is temporary, as upon chronic serum starvation, AR activation delayed but did not suppress the onset of autophagy and cell death. This study reveals a novel mechanism whereby maintained AR signalling promotes temporary adaptation to cellular stress and in turn may contribute to the evasion of prostate tumour cell death.
Similar content being viewed by others
Main
Prostate cancer is a hormonally driven disease, dependent on endogenous androgens for its development. Androgen ablation is the most common form of therapeutic intervention for prostate cancer patients, resulting in widespread death of prostate epithelia.1 Many studies have attempted to delineate the crosstalk between androgen receptor (AR) and antiapoptotic signalling pathways;2, 3, 4, 5, 6 however, the contribution of AR towards macroautophagy (referred to hereafter as autophagy), a mechanism that can either promote cell survival or contribute to cell death, has only recently been appreciated.7, 8, 9, 10 Autophagy is a homeostatic process whereby a cell engulfs damaged and long-lived proteins, organelles and cytoplasmic components into autophagosomes. Autophagosomal fusion with lysosomes permits the proteolytic degradation of its contents to generate small reusable biomolecules.11, 12, 13 Autophagy is upregulated when cells are under stress, such as during nutrient starvation or hypoxia – conditions that are common in the poorly vascularized environment of a tumour – thus allowing for the replenishment of nutrients and energy or the clearance of cytotoxic compounds. In this context, autophagy can promote survival by allowing for temporary cellular adaptation to unfavourable conditions; however, upon chronic insult, this process can contribute towards cell death.14, 15, 16
One stimulus that can promote autophagy is endoplasmic reticulum (ER) stress. Although numerous signalling effectors have been implicated in linking these two processes, the precise molecular mechanisms remain elusive.17, 18, 19, 20 Inducers of ER stress such as nutrient starvation lead to the accumulation of unfolded proteins within the ER lumen.21, 22 Like autophagy, the ER stress response is initiated to minimize cellular insult, in this case by halting global translation, thereby reducing the protein overload within the ER.23
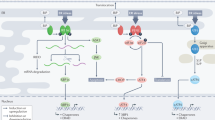
Three well-characterized pathways mediate ER stress signalling, collectively called the unfolded protein response (UPR).22, 23, 24 The master regulator of the UPR is the ER-resident chaperone glucose-regulated protein 78/BiP (Grp78/BiP). The accumulation of unfolded proteins titrates Grp78/BiP from binding to the luminal tails of the ER transmembrane proteins PRK-like ER kinase (PERK), inositol-requiring transmembrane kinase/endonuclease (IRE-1) and activating transcription factor 6 (ATF6). This allows for autophosphorylation of PERK, which in turn phosphorylates and inhibits eukaryotic initiation factor 2α (eIF2α). This inhibition, while preventing global mRNA translation, permits the translation of specific cap-independent ER stress response genes, such as ATF4.24 Secondly, Grp78/BiP dissociation results in the translocation of ATF6 to the Golgi, in which it is activated by proteolytic cleavage.25 Finally, IRE-1 autophosphorylation promotes the splicing of X-box-binding protein-1 (XBP-1) mRNA to its short form, XBP-1s.26 Taken together, the active transcription factors ATF6, XBP-1s and ATF4 regulate the expression of ER chaperones that enhance the folding capacity of the ER, including Grp78/BiP itself, as well as other stress genes such as CHOP and GADD34.22, 23 Although the UPR is an adaptive response, chronic ER stress will eventuate in cell death. Furthermore, ER stress-induced autophagy has been shown to either necessitate or impede apoptosis in a context-dependent manner, highlighting the sensitive interplay between these two processes in the determination of cell fate.18, 19, 27
Here, we have identified a novel role for androgens in the modulation of autophagy. Androgen treatment delayed the induction of serum starvation-induced autophagy by increasing the expression of Grp78/BiP, an effect that was negated by siRNA knockdown of this protein. Altered UPR signalling did not mediate AR inhibition of autophagy, suggesting that alternative signalling pathways downstream of Grp78/BiP may exist to modulate this key biological process. Critically, long-term androgen treatment significantly delayed both the onset of autophagy and cell death compared with cells serum-starved without androgens and this delay in cell death was blocked by Grp78/BiP knockdown. These results support a novel mechanism – modulation of autophagy by Grp78/BiP – by which AR signalling may promote adaptation to nutrient starvation and in turn evasion of cell death.
Results
Androgen treatment blocks serum starvation-induced autophagy
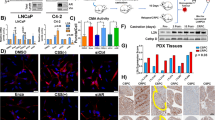
In order to model maintained AR activation in nutrient-poor conditions, we determined the effect of mibolerone, a non-metabolizable synthetic AR agonist, upon serum starvation-induced autophagy.28 Stimulation of AR by its agonists is known to induce AR expression, whereas androgen ablation reduces receptor levels.29, 30 In order to confirm activation of AR by mibolerone, LNCaP cells were cultured for 24 h in medium containing 10% fetal bovine serum (FBS), serum-free (SF) medium or serum-free medium supplemented with 10 ng/ml mibolerone, lysed and western blotted. Although serum starvation leads to the reduction of AR expression, mibolerone-treated cells exhibited protein levels of AR similar to that observed in cells maintained in FBS (Figure 1a). The expression of prostate-specific antigen (PSA), an androgen target gene, was also assessed as a marker of AR transcriptional activity.1 Similarly, PSA expression was specifically induced by mibolerone, confirming activation of AR in the presence of this agonist (Figure 1a).
Androgen treatment blocks serum starvation-induced autophagy. (a) Expression levels of androgen receptor (AR) and PSA in LNCaP cells maintained for 24 h in medium containing 10% fetal bovine serum (FBS), serum free (SF) medium or SF+10 ng/ml mibolerone (SF+mib). Cell lysates were western blotted as indicated, with actin as a loading control. (b) Representative confocal images of cells transiently expressing GFP-LC3 maintained as described in (a). (c) Quantification of GFP-LC3 puncta-positive cells depicted in (b). A positive cell contained five or more punctae. Fifty cells per condition per experiment were analysed. (d) Cells were treated for 0, 2 or 6 h with SF medium with or without bafilomycin and/or mibolerone. Lysates were western blotted with an anti-LC3 antibody, with actin as a loading control. The lipid-conjugated (LC3-II) and -unconjugated (LC3-I) forms are indicated by arrows. Graph depicts densitometric analysis of LC3-II band intensity, normalized to actin levels and expressed as fold change from untreated control. (e) Representative confocal images of cells stably expressing mRFP-GFP-LC3 treated as described in (a). Merged images depict overlay of GFP (green) and mRFP (red) images. (f) Quantification of mRFP+ and mRFP+/GFP+ punctae in cells shown in (e), 25 cells per condition per experiment. For (c), (d) and (f), n=3, error bars indicate mean S.E. *P<0.05 and **P<0.01, as indicated. For (b) and (e), scale bar=20 nm
To investigate the effect of mibolerone on serum starvation-induced autophagy, LNCaP cells were infected with an adenoviral plasmid encoding GFP-tagged LC3 and assessed by confocal microscopy. LC3 is a microtubule-associated protein that becomes lipidated and recruited to the autophagosomal membrane upon autophagy induction.11, 13 To confirm proper recruitment of GFP-LC3, cells were treated for 6 h with bafilomycin A, an inhibitor of the lysosomal V-ATPase proton pump. This perturbation of lysosomal function blocks autophagosomal trafficking, resulting in the accumulation of autophagosomes.31 Accordingly, bafilomycin-treated cells exhibit numerous GFP-LC3 punctae (Supplementary Figure S1).
In cells maintained in FBS, GFP-LC3 displayed a diffuse cytoplasmic localization, with few punctae, corresponding to the low basal levels of autophagy in these cells (Figure 1b). Upon serum starvation for 24 h, numerous GFP-LC3 punctae were detected; however, this was not observed in cells maintained in SF+mibolerone (Figure 1b). Quantification revealed that serum starvation significantly increased the number of punctae-positive cells, but the presence of mibolerone reduced this number to that comparable to basal levels (Figure 1c).
The absence of GFP-LC3 punctae in cells serum starved with mibolerone for 24 h can indicate an inhibition of autophagy induction.31 To explore this possibility, the lipidation of endogenous LC3 was assessed by western blot, as lipid conjugation of this protein results in an electrophoretic mobility shift.32 Cells were treated for 2 or 6 h in SF or SF+mibolerone in the presence or absence of bafilomycin A, to allow for the accumulation of lipidated LC3, and lysed. In cells not treated with bafilomycin A, only basal levels of the lipidated (LC3-II) form could be detected, presumably owing to the high turnover of LC3-II by lysosomal degradation or delipidation31 (Figure 1d). However, in the presence of bafilomycin A, an increase in accumulated LC3-II was detected in cell lysates maintained in SF compared with those in SF+mibolerone, shown to be significant by densitometric analysis (Figure 1d). This result suggests that mibolerone impacts upon the flux of autophagosomes at the induction stage.
In order to confirm that mibolerone blocks autophagy induction in serum-starved cells, we wished to visualize the trafficking of autophagosomes from induction to lysosomal fusion. To do so, LNCaP cells were stably infected with a plasmid encoding LC3 that is dually labelled with mRFP and GFP. As GFP fluorescence is pH-sensitive, this signal is lost upon fusion with the acidic lysosomal compartment. mRFP fluorescence, however, is unaffected, and thus the colocalization of GFP+ and mRFP+ punctae acts as a marker for a (pre-fusion) autophagosome, whereas punctae positive for only mRFP signifies a (post-fusion) autolysosome.33 To verify the expected localization of mRFP-GFP-LC3, cells were treated with bafilomycin A for 6 h, after which they showed almost complete colocalization of GFP and mRFP (Supplementary Figure S2).
In accordance with previous results (Figure 1b), cells expressing mRFP-GFP-LC3 displayed a mainly diffuse localization when maintained for 24 h in FBS (Figure 1e). In contrast, serum starvation resulted in the formation of both mRFP+/GFP+ and mRFP+ punctae, showing the maintained induction and trafficking of autophagosomes through to lysosomal fusion, and quantification of both puncta types revealed that this was a significant effect (Figure 1e and f). However in serum-starved cells supplemented with mibolerone, we observed numbers of both puncta types comparable to FBS (Figure 1e and f). These results, taken together with the analysis of LC3-II lipidation in Figure 1d, confirm that treatment with mibolerone blocks the induction of serum starvation-induced autophagy.
Androgen receptor activation blocks autophagy induction independently of mammalian target of rapamycin signalling
Mammalian target of rapamycin (mTOR) acts as a key sensor of growth factor activation, nutrient availability and energy status, and the inhibition of this protein induces autophagy.34 The activation status of the mTOR signalling pathway in serum-starved cells was determined by western blotting for phosphorylated mTOR and its downstream effectors S6 kinase, ribosomal protein S6 and the translation factor binding protein 4E-BP1. As predicted, the phosphorylation of mTOR and its effectors were reduced after serum starvation compared with cells maintained in FBS (Figure 2a). Densitometric analysis confirmed that this reduction was statistically significant (Figure 2b).
Androgen suppresses autophagy induction independently of mTOR signalling. (a) Cells that were maintained for 24 h in SF medium alone or with 10 ng/ml mibolerone (SF+mib), 1 μM rapamycin (SF+rapa) or in combination (SF+mib+rapa) were lysed and western blotted with the indicated antibodies, with actin as a loading control. All panels are representative of at least three independent experiments. (b) Graphs depict densitometric analysis of band intensities of p-mTOR, p-S6K, p-S6 and p-4E-BP1, normalized to total protein levels and expressed as fold change from untreated control. (c) Representative confocal images of cells transiently expressing GFP-LC3 maintained for 24 h in the indicated conditions. Scale bar=20 nm. (d) Quantification of GFP-LC3 punctae-positive cells described in (c). A positive cell contained 10 or more punctae. n=3, 50 cells per condition per experiment, For (b) and (d), error bars indicate mean S.E. In (b), **P<0.01 versus FBS; in (d), **P<0.01 as indicated; and #P<0.01 versus FBS
We hypothesized that the mechanism whereby mibolerone blocks autophagy induction could be by the re-activation of mTOR signalling in serum-starved cells. Interestingly, mibolerone supplementation did not increase phosphorylation of mTOR and its effectors compared with cells serum-starved alone (Figure 2a and b). Similarly, the mTOR inhibitor rapamycin almost completely blocked phosphorylation of mTOR and its downstream effectors in serum-starved cells and the addition of mibolerone did not over-ride this inhibition (Figure 2a and b). These results suggest that mibolerone may act upon the autophagy pathway in a mechanism independent of mTOR.
To investigate directly the role of the mTOR pathway in autophagy, we treated cells with rapamycin to promote autophagy and then determined whether the addition of mibolerone could circumvent this effect. GFP-LC3-expressing cells were treated with FBS, SF or SF+mibolerone with or without rapamycin. The addition of rapamycin, as anticipated, significantly induced autophagy in all conditions (Figure 2c and d). However, as observed with controls, serum starvation increased the number of cells that were positive for GFP-LC3 punctae above those maintained in FBS, but the presence of mibolerone blocked this enhancement (Figure 2c and d). This is supported by the observation that endogenous LC3 lipidation, determined by western blot, was enhanced in rapamycin-treated cells maintained in serum-free medium compared with those in mibolerone, an effect that was pronounced in the presence of bafilomycin A (S3). Taken together, these results show that the mechanism by which mibolerone blocks autophagy induction is largely mTOR independent, as mibolerone attenuated rapamycin-induced autophagy without impacting upon activation of the mTOR pathway.
Effect of androgen receptor activation on serum starvation-induced ER stress
As mTOR activation could be excluded as the mechanism by which mibolerone inhibited autophagy, we next investigated whether a reduction in ER stress induced by serum withdrawal could underlie this blockade. Initially, protein and mRNA levels of the ER chaperone Grp78/BiP were assessed, as this protein has been implicated as a key mediator of ER stress-induced autophagy.17 In serum-starved cells, Grp78/BiP protein levels remained unchanged despite significant increases in mRNA (Figure 3a and b). In contrast, both Grp78/BiP protein and mRNA were significantly upregulated in mibolerone-treated cells (Figure 3a and b). This result raises the possibility that AR may indeed reduce the ER stress response by enhancing Grp78/BiP translation or protein stability and in turn stabilizing the ER.17
Effect of androgen on serum starvation-induced ER stress. For all panels, cells were maintained in FBS, SF or SF+mib for the indicated time points. For (a), (c) and (e), cells were lysed and western blotted with the indicated antibodies. Graphs depict densitometric analysis of band intensities of indicated proteins, normalized to total protein levels and expressed as fold change from untreated control. For (b), (d) and (f), total RNA was extracted and mRNA was quantified by RT-PCR and normalized to actin levels (b and d) or subjected to 2% agarose gel separation (f). (a and b) Expression of Grp78/BiP protein and mRNA. (c) Activation of eIF2α and CHOP. (d) Levels of ATF4 mRNA. (e) ATF6 protein expression and cleavage. Full-length (90 kDa) and cleaved (50 kDa) forms of ATF6 are indicated by arrows. n/s=non-specific band. (f) XBP-1 mRNA splicing. Full length and spliced XBP-1 are indicated by arrows. Cells treated for 6 h with 2 μM thapsigargin (TG) were used as a positive control. All panels are representative of at least three independent experiments. Error bars indicate mean S.E., *P<0.05, **P<0.01 versus FBS or as indicated. For (d), *P<0.05 SF+mib versus all groups
To explore further, we investigated the effect of mibolerone on activation of the three UPR signalling pathways that lie downstream of Grp78/BiP. First, we examined the activation of the PERK-eIF2α-ATF4 axis. In serum-starved cells, eIF2α phosphorylation was markedly enhanced at 2 h and was maintained significantly above basal levels up to 24 h (Figure 3c). ATF4 could not be detected at the protein level, presumably owing to its short half-life35 and no changes in mRNA were observed (Figure 3d). However, protein levels of CHOP, an ATF4 target gene, were upregulated by 24 h (Figure 3c). Taken together, these results show the activation of the PERK-eIF2α-ATF4 axis upon serum starvation. In contrast, cells supplemented with mibolerone exhibited initial eIF2α phosphorylation; however, this was not maintained at 24 h (Figure 3c). In accordance, ATF4 mRNA levels declined in these cells and CHOP protein expression was not induced. These results suggest that the presence of mibolerone attenuates activation of this pathway.
Next, the second arm of the UPR – ATF6 expression and cleavage – was investigated. Expression of the full-length 90-kDa protein was significantly increased by 24 h in cells serum starved with or without mibolerone, and similarly, comparable levels of the cleaved 50 kDa form were detected (Figure 3e). This shows that serum starvation-induced activation of ATF6 was not affected by mibolerone treatment. Finally, assessment of the third arm of the UPR, IRE-1-mediated splicing of XBP-1 mRNA, revealed that this pathway is not activated by serum starvation in these cells (Figure 3f).
Thus, while serum starvation activates both the PERK-eIF2α-ATF4 and ATF6 pathways, the presence of mibolerone prevents activation of the former and crucially leads to the marked upregulation of Grp78/BiP. Taken together, these results raise the possibility that AR inhibition of autophagy may be mediated by alterations to the ER stress response.
Knockdown of Grp78/BiP attenuates androgen-mediated inhibition of autophagy
As Grp78/BiP is a critical mediator of ER stability, we postulated that increases in its expression could underlie the suppression of autophagy by mibolerone. We addressed this by targeted knockdown of Grp78/BiP by siRNA. It has been reported that almost complete knockdown of Grp78/BiP results in spontaneous activation of the UPR and blocks stress-induced autophagy.10, 36 We therefore aimed to use siRNA concentrations that would attenuate mibolerone-mediated protein upregulation, yet would not alter the basal level of autophagy. Cells stably expressing mRFP-GFP-LC3 were pre-treated with control or Grp78/BiP siRNA, and after 24 h post-transfection, they were serum starved with or without mibolerone. Significant knockdown of Grp78/BiP was obtained in FBS cells compared with corresponding siRNA control cells (Figure 4a and b), and assessment of puncta formation confirmed that Grp78/BiP protein was decreased to a level that did not induce autophagy (Figure 4c). Importantly, in Grp78/BiP siRNA-treated cells, the mibolerone-mediated upregulation of this protein was significantly reduced compared with SF+mibolerone controls (Figure 4a and b).
Knockdown of Grp78/BiP prevents androgen-mediated inhibition of autophagy. (a) Control siRNA- and Grp78/BiP-siRNA-treated cells were maintained for 24 h in FBS, SF and SF+mib before lysis. Lysates were analysed by western blot, with GAPDH as loading control. (b) Graphs depict densitometric analysis of band intensities of indicated proteins, normalized to total protein levels and expressed as fold change from untreated control. (c) Representative confocal images of cells stably expressing mRFP-GFP-LC3 treated as described in (a). Merged images depict overlay of GFP (green) and mRFP (red) images. Scale bar=20 nm. (d) Quantification of mRFP+ (right panel) and mRFP+/GFP+ (left panel) punctae in cells shown in (c). n=3, 25 cells per condition per experiment. For (b) and (d), error bars indicate mean S.E., *P<0.05 and **P<0.01 versus FBS or as indicated
Next, the effect of Grp78/BiP knockdown on AR suppression of autophagy was determined. Cells treated with control siRNA exhibited serum starvation-induced autophagy that was blocked by mibolerone (Figure 4c and d). After knockdown of Grp78/BiP, autophagy induced by serum starvation was comparable to that observed in control siRNA cells (Figure 4c and d). However, mibolerone was unable to inhibit autophagy in Grp78/BiP knockdown cells and quantification of mRFP+/GFP and mRFP+ punctae revealed that autophagosomes were induced and trafficked to lysosomes to a comparable extent as cells serum starved only (Figure 4c and d). This shows that Grp78/BiP knockdown is sufficient to revert mibolerone-mediated inhibition of autophagy and confirms that the upregulation of this protein by androgens underlies autophagy modulation.
Androgen inhibition of autophagy is not mediated by eIF2α dephosphorylation
As we had observed attenuated signalling of the p-eIF2α-ATF4-CHOP pathway in mibolerone-treated cells (Figure 3c and d), we hypothesized that this may be a direct result of upregulated Grp78/BiP, thus preventing PERK dimerization and activation. To investigate further, we examined whether Grp78/BiP knockdown would abolish eIF2α dephosphorylation observed after mibolerone treatment. In FBS and SF cells, Grp78/BiP knockdown significantly increased p-eIF2α levels, as expected (Figure 4a and b). However, surprisingly, we observed significantly reduced phosphorylation of eIF2α in mibolerone-treated cells compared with serum starvation only, even upon Grp78/BiP knockdown (Figure 4a and b). This dephosphorylation could be owing to residual Grp78/BiP upregulation by mibolerone in knockdown cells or alternatively could represent a Grp78/BiP-independent mechanism of eIF2α modulation by AR. However, as autophagy was significantly induced by mibolerone in Grp78/BiP knockdown cells despite reduced eIF2α phosphorylation, this suggests that attenuated signalling of this UPR axis does not underlie AR-mediated autophagy regulation.
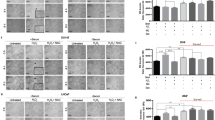
In order to investigate specifically the role of eIF2α phosphorylation in autophagy inhibition by androgen, we co-treated cells with 50 μM salubrinal, an inhibitor of the CReP/GADD34/PP1 complex responsible for eIF2α dephosphorylation.37 Treatment with salubrinal significantly increased p-eIF2α levels in all groups, whereas control cells exhibited a serum starvation-induced activation only (Figure 5a and b). Importantly, salubrinal treatment did not impinge upon mibolerone-mediated Grp78/BiP upregulation, allowing for an independent assessment of the effects of eIF2α manipulation (Figure 5a and b).
Blocking eIF2α dephosphorylation with salubrinal does not impair androgen-mediated autophagy inhibition. (a) Cells treated for 24 h in FBS, SF or SF+mib with 50 μM salubrinal or DMSO vehicle control were lysed and western blotted with the indicated antibodies. Actin was used as loading control. (b) Graphs depict densitometric analysis of band intensities of indicated proteins, normalized to total protein levels and expressed as fold change from untreated control. (c) Representative confocal images of cells treated as described in (a). Scale bar=20 nm. (d) Quantification of GFP-LC3 puncta-positive cells shown in (c). A positive cell contained five or more punctae. n=3, 50 cells per condition per experiment, For (b) and (d), error bars indicate mean S.E., *P<0.05 and **P<0.01 as indicated; #P<0.01 versus FBS
Interestingly, salubrinal treatment induced autophagy in all groups, as evidenced by a significant increase in the number of GFP-LC3 puncta-positive cells compared with control (Figure 5c and d). However, although the basal level of autophagy was enhanced with salubrinal, we observed that serum starvation further increased the number of positive cells and that this effect was absent in the presence of mibolerone (Figure 5c and d). Thus, maintained eIF2α phosphorylation is insufficient to block autophagy inhibition by mibolerone.
Taken together, these results suggest that while AR activation may modulate phosphorylation of eIF2α and the subsequent activation of its effectors (Figure 3c and d), this pathway is not involved in AR-mediated inhibition of autophagy.
Chronic serum starvation in the presence of androgen delays, but does not suppress autophagy induction
As it had been established that mibolerone could block serum starvation-induced autophagy up to 24 h, we investigated whether this effect was maintained upon chronic serum starvation. This was assessed by the quantification of mRFP+/GFP+ and mRFP+ punctae in mRFP-GFP-LC3-expressing cells up to 96 h of serum starvation with or without mibolerone (Figure 6a–c). Interestingly, in serum-starved cells, the average number of punctae increased marginally over time, suggesting that the rate of autophagosomal induction and turnover increases slowly and stably. In contrast, cells serum starved in the presence of mibolerone exhibited a rapid increase in the rate of autophagy induction and trafficking, with significant increases in both mRFP+/GFP and mRFP+ punctae observed between each time point up to 72 h (Figure 6a–c). In addition, endogenous LC3 levels and lipidation were upregulated in both conditions over 96 h (Figure 6d). This result shows that the inhibition of autophagy by mibolerone at 24 h represents a delay in the onset, rather than an absolute suppression, of this process, and by 96 h, these cells exhibit comparable autophagic activity as cells serum starved alone.
Androgen delays chronic serum starvation-induced autophagy. (a and b). Time course of autophagy induction and trafficking in mRFP-GFP-LC3 stable cells. Quantification of mRFP+/GFP+ (a) and mRFP+ (b) punctae in cells maintained in SF or SF+mib for 24, 48, 72 and 98 h. Twenty-five cells were analysed per condition per experiment, n=3, error bars indicate mean S.E., *P<0.05. (c). Representative confocal images of cells maintained in FBS medium or after 96 h of SF or SF+mib. Merged images depict overlay of GFP (green) and mRFP (red) images. Scale bar=20 nm. (d) Cells were treated as described in (a), lysed and subjected to western blot analysis with the indicated antibodies. All panels are representative of three independent experiments
To confirm that the onset of autophagy in mibolerone-treated cells was not owing to reduction in the efficacy of this non-metabolizable drug, the levels of AR and PSA were investigated by western blot (Figure 6d). AR and PSA expression were comparable to levels observed in FBS cells and were markedly greater than those of cells serum starved alone. Thus, a reduction in drug efficacy was excluded as an explanation for autophagy induction.
As it had been established that knockdown of Grp78/BiP was sufficient to permit autophagy in mibolerone-treated cells, we reasoned that changes in the expression of this protein may underlie the onset of autophagy observed at later time points. Indeed, Grp78/BiP levels declined over time and were comparable to those seen in SF cells by 72 h (Figure 6d). This correlates with the induction and upregulation of autophagy in mibolerone-treated cells (Figure 6a–c), supporting the hypothesis that autophagy modulation by mibolerone is mediated by Grp78/BiP.
Grp78/BiP knockdown blocks androgen-mediated delay in cell death
In order to determine the consequence of long-term serum starvation, cells were maintained as in Figure 6a and subjected to cell death analysis by propidium iodide (PI) positivity. Although serum starvation did not initially induce cell death, by 72 h these cells exhibited a significant increase in the percentage of PI-positive cells compared with basal levels, an effect not observed in mibolerone-treated cells (Figure 7a). However, by 96 h, the amount of cell death seen in both conditions was comparable (Figure 7a). This shows that chronic serum starvation induces cell death and this induction is delayed, but not suppressed, by mibolerone.
Androgen delays chronic serum starvation-induced cell death. (a) Time course of cell death of cells maintained in FBS, SF or SF+mib for 24, 48, 72 or 96 h. Cell death was assessed by staining live cells with PI before FACS analysis. (b) Cells were transfected with control or Grp78/BiP siRNA for 24 h, and then treated for a further 72 h with FBS, SF or SF+mib and analysed as in (a). n=3, error bars indicate mean S.E. *P<0.05 and **P<0.01 as indicated; #P<0.05 versus corresponding control
As we had established that Grp78/BiP mediated autophagy inhibition in mibolerone-treated cells (Figure 4c and d), we considered whether this protein also mediated the delay in cell death. We investigated death at 72 h, as at this time point a significant difference was observed between cells serum starved with or without mibolerone (Figure 7a). As expected, assessment of PI positivity in control siRNA cells showed that serum starvation-induced death was decreased by mibolerone. However, Grp78/BiP knockdown abrogated this inhibitory effect of AR activation (Figure 7b). This result shows that the delay in cell death induced by mibolerone is mediated by the initial upregulation of Grp78/BiP. Taken together, these findings highlight that this chaperone protein is essential for affecting two critical biological processes – autophagy and survival – downstream of the activated AR.
Discussion
In this study, we have established that AR activation abolishes autophagy induced by acute serum starvation. We report the novel finding that in this context, androgen treatment does not inhibit autophagy by initiating mTOR signalling, but rather acts via upregulation of Grp78/BiP, a key mediator of ER stability. Crucially, we show that the adaptation to serum starvation by androgens is temporary, as upon prolonged serum withdrawal androgen-treated cells exhibit a delay, rather than a suppression, of both autophagy and cell death, events which correlate with a reduction in Grp78/BiP protein levels. Finally, knockdown of this protein from the onset of serum starvation abrogates the survival advantage provided by androgen supplementation, establishing a definitive role for Grp78/BiP as a key effector of AR signalling.
Oligonucleotide arrays and proteomic analysis of androgen-treated prostate cancer cells have revealed the regulation of several ER chaperones and resident proteins, including Grp78/BiP.38, 39, 40, 41, 42, 43, 44 Immunohistochemical studies of human prostate cancer samples show that increased expression of Grp78/BiP is associated with aggressive forms of the disease and prostate-specific knockout of this protein blocks tumorigenesis in a PTEN-null mouse model, suggesting a key role for Grp78/BiP in prostate cancer development.45, 46, 47 However, a direct functional relationship between AR and Grp78/BiP had not been investigated, and despite the mounting evidence of a role for androgens in the regulation of the ER, this is the first report that describes an effect on UPR signalling.
Interestingly, of the two proximal UPR pathways activated by serum starvation, only PERK-eIF2α-ATF4 activation was diminished by androgen, whereas ATF6 cleavage was unaffected. The mechanisms underlying this difference are unclear; however, our evidence suggests that the inhibitory effect of androgen on the PERK-eIF2α-ATF4 pathway may occur independently of Grp78/BiP upregulation (Figure 4a and b). This raises the possibility that an additional level of input into this pathway by AR may exist, for example, by changing the expression profile of proteins that reside outside the ER lumen, which regulate PERK or eIF2α. However, it is clear from our results that modulation of this pathway is not necessary for AR to exert its inhibitory effects on autophagy induction, in contrast to that of its upstream regulator, Grp78/BiP. This highlights a critical question with regard to the role of Grp78/BiP upregulation. If this increase in protein expression does not act to suppress UPR signalling, then by what mechanism does it regulate autophagy? Several reports have identified Grp78/BiP localization on the plasma membrane or cytosol, suggesting multiple roles beyond ER stability and UPR modulation.48, 49, 50 Further experiments are currently underway to elucidate potential signalling mechanisms by which this protein may feed into the autophagy pathway.
The observation that androgen inhibited autophagy via Grp78/BiP upregulation, rather than by activation of mTOR signalling, is a novel finding given that previous reports have shown AR activation can positively affect this key signalling node.10, 51 It appears, however, that the influence of AR upon mTOR activity is dependent on the energy and nutrient status of the cell. For example, DHT stimulation led to the activation of mTOR signalling in LNCaP cells maintained in medium with steroid-depleted serum.51 In serum-starved conditions, protein synthesis is blocked, evidenced in our model by the reduction in mTOR signalling and phosphorylation of eIF2α, thus inhibiting ribosomal biogenesis and translation initiation.52 That the AR target genes PSA and AR itself are expressed throughout mibolerone treatment illustrates the ability of AR to drive protein synthesis of specific genes even in nutrient-poor conditions, and our results suggest that this may occur by attenuating translational inhibition through the modulation of eIF2α phosphorylation, rather than by reactivating mTOR. Similarly, we observe that mibolerone reduces rapamycin-induced autophagy while not influencing mTOR activation, highlighting that the opposing effects on autophagy driven by these two compounds are mediated by different mechanisms. This observation is supported by Wang et al.,53 who investigated the apoptotic effects of rapamycin and the AR antagonist bicalutamide and revealed that mTOR and AR impact upon survival by parallel, not overlapping, pathways. Taken together, we have clearly shown that AR activation can exert mTOR-independent cellular effects.
One consequence of adaptation to starvation-induced stress by androgens was to inhibit autophagy; however, this adaptation only provided temporary respite. Autophagy is a catabolic process, whereas androgens directly promote several well-characterized anabolic pathways, such as lipogenesis and enhanced glucose utilization.54, 55 In our model, the net effect of these two opposing stimuli – serum starvation and AR activation – was a delay in autophagy induction. Given that the outcome of AR signalling is anabolic, the initial increase in Grp78/BiP levels may reflect adaptation to increased protein synthesis to avoid ER overloading. However, upon chronic serum starvation, we observed both a reduction in Grp78/BiP levels and an increase in cell death. The switch between stress adaptation and death appears to lie in the selective stability of pro-survival mRNA and proteins (i.e. Grp78/BiP and other ER chaperones) compared with those that promote apoptosis (i.e. CHOP).21, 22, 35 This is reiterated in our model as preventing androgen-mediated upregulation of Grp78/BiP from the onset of serum starvation with siRNA negated any survival advantage of androgen supplementation.
These results have critical implications with regard to the mechanisms by which AR signalling contributes to the evasion of cell death in vivo. Extrapolation of our results to model the nutrient-poor tumour microenvironment suggests that maintained AR signalling, be it via local or systemic androgen production, mutation, cross-talk or by other mechanisms, promotes acute adaptation to nutrient starvation, thus providing an initial survival advantage.1, 56 We can attribute this to AR-stimulated upregulation of Grp78/BiP, providing a novel mechanistic hypothesis for the observed correlation of high protein levels with aggressive forms of the disease. According to our results, maintained AR signalling will not overcome the deleterious effects of chronic starvation, arguing for the requirement for multiple genetic changes to promote androgen-independent prostate cancer. However, our results highlight a novel mechanism whereby AR may contribute to the progression to this phenotype by allowing for temporary adaptation to cellular stress.
Materials and Methods
Cell culture protocols, transfections and reagents
The human prostate cancer cell line LNCaP was obtained by American Type Culture Collection (ATCC). Cells were maintained in RPMI 1640 (Gibco, Invitrogen, Paisley, UK) supplemented with 10% FBS (Harlan, Indianapolis, IN, USA) and 2 mM L-glutamine (Gibco) at 37°C and 5% CO2. For serum starvation experiments, cells were washed twice in PBS, and then maintained in phenol red-free RPMI 1640 supplemented with 2 mM L-glutamine. For GFP-LC3 expression, subconfluent LNCaP cells were infected with an adenoviral vector encoding GFP-LC3 fusion, a kind gift of Aviva Tolkovsky (University of Cambridge, Cambridge, UK). At 24 h post-infection, cells were washed with fresh medium before treatment. For construction of the pWZL-mRFP-GFP-LC3 expression vector, mRFP-GFP-LC3 fragment33 was isolated by Nhee1–Sal1 digest and inserted into SnaB1/Sal1-digested pWZL-HYGRO plasmid. For the establishment of stable cell lines, subconfluent cells were harvested and subjected to nucleofection with Nucleofector Kit R (Lonza, Basel, Switzerland), according to the manufacturer's instructions. Cells were selected according to hygromycin B resistance (100 μg/ml). Thapsigargin, bafilomycin A, rapamycin, hygromycin and PI were all purchased from Sigma (St Louis, MO, USA); mibolerone from PerkinElmer (Boston, MA, USA); and salubrinal from Tocris (Ellisville, MI, USA).
Western blotting
Cells were washed in ice-cold PBS before lysis with 2 × Laemmli buffer containing 10% β-mercaptoethanol. Lysates were boiled for 10 min at 95°C, proteins were separated using 15% or 4–12% SDS-PAGE (Invitrogen) and transferred to polyvinylidene difluoride (Immobilon-P, Millipore, Bedford, MA, USA). Membranes were blocked in 5% skim milk powder/TBS-Tween-20 and incubated in primary antibody overnight at 4°C. After washing, membranes were incubated in secondary antibody for 1 h at room temperature and visualized with chemiluminescence detection (GE Healthcare, Buckinghamshire, UK) on autoradiographic film (Fuji, Bedfordshire, UK). Densitometric analysis was performed on scanned autoradiographs using the Image J software (version 1.40g).
Primary antibodies used in this study were purchased from Sigma (actin, clone AC-40; GAPDH-peroxidase, clone GAPDH-71.1); Cell Signalling Technology, Danvers, MA, USA (CHOP, clone L63F7; phospho-mTOR (S2448); phospho-eIF2α (S51), clone 119A11; phospho-4EBP1 (S65), clone 174A9; phospho-p70 S6 kinase (T421/S424); phospho-PERK (T980), clone 16F8; Grp78/BiP, clone C50B12; phospho-S6 (S235/6), clone 2F9; p70 S6 kinase; mTOR; S6); Imgenex, San Diego, CA, USA (ATF6); Novus, Littleton, CO, USA (LC3); and Santa Cruz (AR; PSA, clone C-19). Secondary antibodies used were anti-mouse, anti-rabbit and anti-sheep IgG-HRP conjugate from Cell Signalling Technology.
siRNA
siRNA oliogonucleotides used in this study were purchased from Thermo Scientific, Waltham, MA, USA. For siRNA uptake, cells were nucleofected using Nucleofector Kit R (Lonza) according to the manufacturer's instructions. At 24 h post-nucleofection, the medium was changed to treatment conditions for the indicated time.
Confocal microscopy
To visualize punctae in GFP-LC3- or mRFP-GFP-LC3-expressing cells, cultures were fixed in 4% paraformaldehyde, washed in PBS and mounted in Vectashield (Vector Laboratories, Burlingame, CA, USA). Cells were visualized using an Olympus FV-1000 confocal microscope under × 63 objective and analysed using the Image J software (version 1.40g).
Q-RT-PCR
Subconfluent cells were harvested and RNA extracted using RNeasy Mini Kit (Qiagen, Hilden, Germany). Synthesis of cDNA and PCR amplification from total RNA was performed using DyNAmo SYBR Green 2-step qRT-PCR kit (Finnzymes, Espoo, Finland) and MJ Research Peltier Thermal Cycler apparatus (Bio-Rad, Hercules, CA, USA). Quantification was determined using the MJ Opticon Monitor 3 analysis software, version 3.1 (Bio-Rad). The following primers were used for qRT-PCR: ATF4 – GGGACAGATTGGATGTTGGAG A (sense), ACCCAACAGGGCATCCAAGT (antisense); Grp78/BiP – CATGGTTCTCACTAAAATGAAAG (sense), GCTGGTACAGTAACAACTG (antisense); XBP1 – CTGGAACAGCAAGTGGTAGA (sense), CTGGGTCCTTCTGGGTAGAC (antisense); and β-actin – AAGATCAAGATCATTGCTCCTCCT (sense), TCATAGTCCGCCTAGAAGCA (antisense). Primers for ATF4 and Grp78/BiP were purchased from Invitrogen and XBP-1 and β-actin from VHBio (Newcastle, UK).
Cell death assays
For determination of cell death, adherent and floating cells were collected, pelleted and resuspended in 0.1 mg/ml PI. Cells were analysed according to fluorescence intensity using the FACSCaliber flow cytometer and Cell Quest software, version 5.2.1 (BD Bioscience, San Jose, CA, USA).
Statistical analysis
Statistical analysis was performed using the unpaired Student's t-test, with differences of P<0.05 considered to be statistically significant.
Abbreviations
- Grp78/BiP:
-
glucose-regulated protein 78/BiP
- ER:
-
endoplasmic reticulum
- mTOR:
-
mammalian target of rapamycin
- eIF2α:
-
eukaryotic initiation factor 2α
- PERK:
-
PRK-like ER kinase
- IRE-1:
-
inositol-requiring transmembrane kinase/endonuclease
- ATF4,6:
-
activating transcription factor 4, 6
- XBP-1:
-
X-box-binding protein-1
- PI:
-
propidium iodide
References
Feldman BJ, Feldman D . The development of androgen-independent prostate cancer. Nat Rev Cancer 2001; 1: 34–45.
Cohen MB, Rokhlin OW . Mechanisms of prostate cancer cell survival after inhibition of AR expression. J Cell Biochem 2009; 106: 363–371.
Huggins C . Endocrine-induced regression of cancers. Cancer Res 1967; 27: 1925–1930.
Kaarbo M, Klokk TI, Saatcioglu F . Androgen signaling and its interactions with other signaling pathways in prostate cancer. Bioessays 2007; 29: 1227–1238.
Lin Y, Kokontis J, Tang F, Godfrey B, Liao S, Lin A et al. Androgen and its receptor promote Bax-mediated apoptosis. Mol Cell Biol 2006; 26: 1908–1916.
Tang F, Kokontis J, Lin Y, Liao S, Lin A, Xiang J . Androgen via p21 inhibits TNF-alpha-induced JNK activation and apoptosis. J Biol Chem 2009; 284: 32353–32358.
Coto-Montes A, Tomas-Zapico C, Martinez-Fraga J, Vega-Naredo I, Sierra V, Caballero B et al. Sexual autophagic differences in the androgen-dependent flank organ of Syrian hamsters. J Androl 2009; 30: 113–121.
Du L, Hickey RW, Bayir H, Watkins SC, Tyurin VA, Guo F et al. Starving neurons show sex difference in autophagy. J Biol Chem 2009; 284: 2383–2396.
Kwong J, Choi HL, Huang Y, Chan FL . Ultrastructural and biochemical observations on the early changes in apoptotic epithelial cells of the rat prostate induced by castration. Cell Tissue Res 1999; 298: 123–136.
Li M, Jiang X, Liu D, Na Y, Gao GF, Xi Z . Autophagy protects LNCaP cells under androgen deprivation conditions. Autophagy 2008b; 4: 54–60.
Levine B . Cell biology: autophagy and cancer. Nature 2007; 446: 745–747.
White E, DiPaola RS . The double-edged sword of autophagy modulation in cancer. Clin Cancer Res 2009; 15: 5308–5316.
Xie Z, Klionsky DJ . Autophagosome formation: core machinery and adaptations. Nat Cell Biol 2007; 9: 1102–1109.
Eisenberg-Lerner A, Bialik S, Simon HU, Kimchi A . Life and death partners: apoptosis, autophagy and the cross-talk between them. Cell Death Differ 2009; 16: 966–975.
Kroemer G, Levine B . Autophagic cell death: the story of a misnomer. Nat Rev Mol Cell Biol 2008; 9: 1004–1010.
Scarlatti F, Granata R, Meijer AJ, Codogno P . Does autophagy have a license to kill mammalian cells? Cell Death Differ 2009; 16: 12–20.
Li J, Ni M, Lee B, Barron E, Hinton DR, Lee AS . The unfolded protein response regulator GRP78/BiP is required for endoplasmic reticulum integrity and stress-induced autophagy in mammalian cells. Cell Death Differ 2008a; 15: 1460–1471.
Milani M, Rzymski T, Mellor HR, Pike L, Bottini A, Generali D et al. The role of ATF4 stabilization and autophagy in resistance of breast cancer cells treated with Bortezomib. Cancer Res 2009; 69: 4415–4423.
Ogata M, Hino S, Saito A, Morikawa K, Kondo S, Kanemoto S et al. Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol Cell Biol 2006; 26: 9220–9231.
Sakaki K, Wu J, Kaufman RJ . Protein kinase Ctheta is required for autophagy in response to stress in the endoplasmic reticulum. J Biol Chem 2008; 283: 15370–15380.
Heath-Engel HM, Chang NC, Shore GC . The endoplasmic reticulum in apoptosis and autophagy: role of the BCL-2 protein family. Oncogene 2008; 27: 6419–6433.
Schonthal AH . Endoplasmic reticulum stress and autophagy as targets for cancer therapy. Cancer Lett 2009; 275: 163–169.
Schroder M . Endoplasmic reticulum stress responses. Cell Mol Life Sci 2008; 65: 862–894.
Ron D, Walter P . Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol 2007; 8: 519–529.
Shen J, Chen X, Hendershot L, Prywes R . ER stress regulation of ATF6 localization by dissociation of BiP/GRP78 binding and unmasking of Golgi localization signals. Dev Cell 2002; 3: 99–111.
Hetz C, Glimcher LH . Fine-tuning of the unfolded protein response: assembling the IRE1alpha interactome. Mol Cell 2009; 35: 551–561.
Gozuacik D, Bialik S, Raveh T, Mitou G, Shohat G, Sabanay H et al. DAP-kinase is a mediator of endoplasmic reticulum stress-induced caspase activation and autophagic cell death. Cell Death Differ 2008; 15: 1875–1886.
Murthy LR, Johnson MP, Rowley DR, Young CY, Scardino PT, Tindall DJ . Characterization of steroid receptors in human prostate using mibolerone. Prostate 1986; 8: 241–253.
Krongrad A, Wilson CM, Wilson JD, Allman DR, McPhaul MJ . Androgen increases androgen receptor protein while decreasing receptor mRNA in LNCaP cells. Mol Cell Endocrinol 1991; 76: 79–88.
Wolf DA, Herzinger T, Hermeking H, Blaschke D, Horz W . Transcriptional and posttranscriptional regulation of human androgen receptor expression by androgen. Mol Endocrinol 1993; 7: 924–936.
Klionsky DJ, Abeliovich H, Agostinis P, Agrawal DK, Aliev G, Askew DS et al. Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy 2008; 4: 151–175.
Mizushima N, Yoshimori T . How to interpret LC3 immunoblotting. Autophagy 2007; 3: 542–545.
Kimura S, Noda T, Yoshimori T . Dissection of the autophagosome maturation process by a novel reporter protein, tandem fluorescent-tagged LC3. Autophagy 2007; 3: 452–460.
Diaz-Troya S, Perez-Perez ME, Florencio FJ, Crespo JL . The role of TOR in autophagy regulation from yeast to plants and mammals. Autophagy 2008; 4: 851–865.
Rutkowski DT, Arnold SM, Miller CN, Wu J, Li J, Gunnison KM et al. Adaptation to ER stress is mediated by differential stabilities of pro-survival and pro-apoptotic mRNAs and proteins. PLoS Biol 2006; 4: e374.
Suzuki T, Lu J, Zahed M, Kita K, Suzuki N . Reduction of GRP78 expression with siRNA activates unfolded protein response leading to apoptosis in HeLa cells. Arch Biochem Biophys 2007; 468: 1–14.
Boyce M, Bryant KF, Jousse C, Long K, Harding HP, Scheuner D et al. A selective inhibitor of eIF2alpha dephosphorylation protects cells from ER stress. Science 2005; 307: 935–939.
Labrie C, Lessard J, Ben Aicha S, Savard MP, Pelletier M, Fournier A et al. Androgen-regulated transcription factor AIbZIP in prostate cancer. J Steroid Biochem Mol Biol 2008; 108: 237–244.
Ma C, Yoshioka M, Boivin A, Gan L, Takase Y, Labrie F et al. Atlas of dihydrotestosterone actions on the transcriptome of prostate in vivo. Prostate 2009; 69: 293–316.
Nantermet PV, Xu J, Yu Y, Hodor P, Holder D, Adamski S et al. Identification of genetic pathways activated by the androgen receptor during the induction of proliferation in the ventral prostate gland. J Biol Chem 2004; 279: 1310–1322.
Rosen MB, Wilson VS, Schmid JE, Gray LE . Gene expression analysis in the ventral prostate of rats exposed to vinclozolin or procymidone. Reprod Toxicol 2005; 19: 367–379.
Rowland JG, Robson JL, Simon WJ, Leung HY, Slabas AR . Evaluation of an in vitro model of androgen ablation and identification of the androgen responsive proteome in LNCaP cells. Proteomics 2007; 7: 47–63.
Segawa T, Nau ME, Xu LL, Chilukuri RN, Makarem M, Zhang W et al. Androgen-induced expression of endoplasmic reticulum (ER) stress response genes in prostate cancer cells. Oncogene 2002; 21: 8749–8758.
Velasco AM, Gillis KA, Li Y, Brown EL, Sadler TM, Achilleos M et al. Identification and validation of novel androgen-regulated genes in prostate cancer. Endocrinology 2004; 145: 3913–3924.
Daneshmand S, Quek ML, Lin E, Lee C, Cote RJ, Hawes D et al. Glucose-regulated protein GRP78 is up-regulated in prostate cancer and correlates with recurrence and survival. Hum Pathol 2007; 38: 1547–1552.
Fu Y, Wey S, Wang M, Ye R, Liao CP, Roy-Burman P et al. Pten null prostate tumorigenesis and AKT activation are blocked by targeted knockout of ER chaperone GRP78/BiP in prostate epithelium. Proc Natl Acad Sci USA 2008; 105: 19444–19449.
Pootrakul L, Datar RH, Shi SR, Cai J, Hawes D, Groshen SG et al. Expression of stress response protein Grp78 is associated with the development of castration-resistant prostate cancer. Clin Cancer Res 2006; 12: 5987–5993.
Ni M, Zhou H, Wey S, Baumeister P, Lee AS . Regulation of PERK signaling and leukemic cell survival by a novel cytosolic isoform of the UPR regulator GRP78/BiP. PLoS One 2009; 4: e6868.
Whitaker HC, Stanbury DP, Brinham C, Girling J, Hanrahan S, Totty N et al. Labeling and identification of LNCaP cell surface proteins: a pilot study. Prostate 2007; 67: 943–954.
Zhang Y, Liu R, Ni M, Gill P, Lee AS . Cell surface relocalization of the endoplasmic reticulum chaperone and unfolded protein response regulator GRP78/BiP. J Biol Chem 2010; 285: 15065–15075.
Xu Y, Chen SY, Ross KN, Balk SP . Androgens induce prostate cancer cell proliferation through mammalian target of rapamycin activation and post-transcriptional increases in cyclin D proteins. Cancer Res 2006; 66: 7783–7792.
DuRose JB, Scheuner D, Kaufman RJ, Rothblum LI, Niwa M . Phosphorylation of eukaryotic translation initiation factor 2alpha coordinates rRNA transcription and translation inhibition during endoplasmic reticulum stress. Mol Cell Biol 2009; 29: 4295–4307.
Wang Y, Mikhailova M, Bose S, Pan CX, deVere White RW, Ghosh PM . Regulation of androgen receptor transcriptional activity by rapamycin in prostate cancer cell proliferation and survival. Oncogene 2008; 27: 7106–7117.
Heemers HV, Verhoeven G, Swinnen JV . Androgen activation of the sterol regulatory element-binding protein pathway: current insights. Mol Endocrinol 2006; 20: 2265–2277.
Schnier JB, Nishi K, Gumerlock PH, Gorin FA, Bradbury EM . Glycogen synthesis correlates with androgen-dependent growth arrest in prostate cancer. BMC Urol 2005; 5: 6.
Mostaghel EA, Montgomery B, Nelson PS . Castration-resistant prostate cancer: targeting androgen metabolic pathways in recurrent disease. Urol Oncol 2009; 27: 251–257.
Acknowledgements
We would like to thank Aviva Tolkovsky for adenovirus-expressing GFP-LC3, Tanatsu Yoshimori for mRFP-GFP-LC3 fragment and Paul Timpson for the critical reading of this manuscript. This work was supported by Cancer Research UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Piacentini
Supplementary Information accompanies the paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Bennett, H., Fleming, J., O'Prey, J. et al. Androgens modulate autophagy and cell death via regulation of the endoplasmic reticulum chaperone glucose-regulated protein 78/BiP in prostate cancer cells. Cell Death Dis 1, e72 (2010). https://doi.org/10.1038/cddis.2010.50
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2010.50
Keywords
This article is cited by
-
Celastrol-mediated autophagy regulation in cancer
Applied Biological Chemistry (2020)
-
Titanium dioxide nanoparticles induced the apoptosis of RAW264.7 macrophages through miR-29b-3p/NFAT5 pathway
Environmental Science and Pollution Research (2020)
-
Serum starvation enhances nonsense mutation readthrough
Journal of Molecular Medicine (2019)
-
Deregulation of ATG9A by impaired AR signaling induces autophagy in prostate stromal fibroblasts and promotes BPH progression
Cell Death & Disease (2018)
-
Food components and contaminants as (anti)androgenic molecules
Genes & Nutrition (2017)