Featured
-
-
Article
| Open Access Translational regulation shapes the molecular landscape of complex disease phenotypes
Translational regulation shapes the molecular landscape of complex disease phenotypes
To what extent translational control can contribute to global gene expression patterns in the disease state is poorly defined. Here the authors conduct genome-wide RNA-seq and ribosome profiling in a rat model of hypertension and uncover altered translation patterns in disease associated genes.
- Sebastian Schafer
- , Eleonora Adami
- & Norbert Hubner
-
Article
| Open Access Visualization and tracking of tumour extracellular vesicle delivery and RNA translation using multiplexed reporters
Visualization and tracking of tumour extracellular vesicle delivery and RNA translation using multiplexed reporters
Extracellular vesicles (EVs) act as a conduit for intercellular communication through the exchange of cellular materials without direct cell-to-cell contacts. Here the authors develop a multiplexed reporter system that allows monitoring of EV exchange, cargo delivery and protein translation between different cell populations.
- Charles P. Lai
- , Edward Y. Kim
- & Xandra O. Breakefield
-
Article |
 Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation
Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation
Post-translational import of nuclear-encoded proteins shapes the proteome of organelles. Here, Kim et al.show that AKR2A, a critical targeting factor for chloroplast outer membrane proteins, binds to client proteins co-translationally as they exit the ribosome.
- Dae Heon Kim
- , Jae-Eun Lee
- & Inhwan Hwang
-
Article
| Open Access Structural basis for full-spectrum inhibition of translational functions on a tRNA synthetase
Structural basis for full-spectrum inhibition of translational functions on a tRNA synthetase
Borrelidin is an antibiotic with antimicrobial, antifungal, antimalarial and immunosuppressive activity that targets threonyl-tRNA synthetase. Here the authors show that borrelidin functions by preventing binding of all three ThrRS substrates and inducing a distinct, non-productive, conformation of the enzyme.
- Pengfei Fang
- , Xue Yu
- & Min Guo
-
Article |
 FAAH genetic variation enhances fronto-amygdala function in mouse and human
FAAH genetic variation enhances fronto-amygdala function in mouse and human
Fatty acid amide hydrolase (FAAH) is a key regulator of endocannabinoid signalling. Here, the authors develop a knock-in mouse that recapitulates a common human mutation in the FAAH gene and demonstrate parallel neural and behavioural alterations across species, suggesting a gain-of-function in fear regulation.
- Iva Dincheva
- , Andrew T. Drysdale
- & Francis S. Lee
-
Article
| Open Access Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan
Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan
Cellular pathways modulating longevity and stress resistance are known to affect protein translation. Here the authors show that the RNA methyltransferase, Nsun5, or its yeast homologue Rcm1, regulates lifespan of three different model organisms by modifying ribosomal RNA at a specific cytosine residue.
- Markus Schosserer
- , Nadege Minois
- & Johannes Grillari
-
Article
| Open Access Double-sieving-defective aminoacyl-tRNA synthetase causes protein mistranslation and affects cellular physiology and development
Double-sieving-defective aminoacyl-tRNA synthetase causes protein mistranslation and affects cellular physiology and development
Accurate loading of amino acids to their cognate tRNA is essential to avoid mistranslation during protein synthesis, which has been linked to human diseases. Here, Lu et al. present a Drosophilamodel that demonstrates the necessity of two distinct ‘sieves’ to ensure accurate amino acid loading for proper development.
- Jiongming Lu
- , Martin Bergert
- & Beat Suter
-
Article
| Open Access MNKs act as a regulatory switch for eIF4E1 and eIF4E3 driven mRNA translation in DLBCL
MNKs act as a regulatory switch for eIF4E1 and eIF4E3 driven mRNA translation in DLBCL
Diffuse large B-cell lymphoma (DLBCL) is a highly aggressive and heterogeneous type of non-Hodgkin’s lymphoma. Here the authors demonstrate that the differential regulation of eIF4E1 and eIF4E3 by the MAPK-interacting kinases is involved in DLBCL aetiology through modification of the cellular translatome.
- Ari L. Landon
- , Parameswary A. Muniandy
- & Ronald B. Gartenhaus
-
Article |
 Unravelling the mechanism of non-ribosomal peptide synthesis by cyclodipeptide synthases
Unravelling the mechanism of non-ribosomal peptide synthesis by cyclodipeptide synthases
Cyclodipeptide synthases hijack aminoacyl tRNAs to produce various cyclic dipeptides—the biosynthetic precursors of several secondary metabolites. Here, the authors solved the crystal structure of a cyclodipeptide synthase bound to a reaction intermediate analogue and provide novel insights into the mechanism of synthesis.
- Mireille Moutiez
- , Emmanuelle Schmitt
- & Muriel Gondry
-
Article |
 Out-of-frame start codons prevent translation of truncated nucleo-cytosolic cathepsin L in vivo
Out-of-frame start codons prevent translation of truncated nucleo-cytosolic cathepsin L in vivo
The lysosomal protease cathepsin L has been observed in compartments other than endosomes and lysosomes. Here the authors show using knock-in mice that nuclear localization of cathepsin L cannot be caused by N-terminal truncation of procathepsin L as previously hypothesized.
- Martina Tholen
- , Larissa E. Hillebrand
- & Thomas Reinheckel
-
Article
| Open Access 4E-BPs require non-canonical 4E-binding motifs and a lateral surface of eIF4E to repress translation
4E-BPs require non-canonical 4E-binding motifs and a lateral surface of eIF4E to repress translation
eIF4E-binding proteins (4E-BPs) are a conserved class of translational repressors that play essential roles in the regulation of protein expression. Here, Igreja et al.indentify non-canonical interactions between 4E-BPs and eIF4E that are required to effectively displace eIF4G and inhibit translation.
- Cátia Igreja
- , Daniel Peter
- & Elisa Izaurralde
-
Article |
 Functionally diverse dendritic mRNAs rapidly associate with ribosomes following a novel experience
Functionally diverse dendritic mRNAs rapidly associate with ribosomes following a novel experience
Dendritic protein synthesis is implicated in synaptic plasticity and memory storage. Ainsley et al., develop a method for collecting ribosome-bound mRNAs from mouse brain dendrites, and use RNA sequencing to characterize dendritic mRNAs that bind to ribosomes after mice experience a novel environment.
- Joshua A. Ainsley
- , Laurel Drane
- & Leon G. Reijmers
-
Article |
 High-efficiency translational bypassing of non-coding nucleotides specified by mRNA structure and nascent peptide
High-efficiency translational bypassing of non-coding nucleotides specified by mRNA structure and nascent peptide
Translational bypassing occurs when the ribosome skips nucleotides downstream of a take-off codon and re-initiates protein synthesis at a location further down on the mRNA. Here, Samatova et al.show that translational bypassing of bacteriophage T4 gene60 mRNA is specified both by the mRNA sequence and the nascent peptide.
- Ekaterina Samatova
- , Andrey L. Konevega
- & Marina V. Rodnina
-
Article |
 Interplay between trigger factor and other protein biogenesis factors on the ribosome
Interplay between trigger factor and other protein biogenesis factors on the ribosome
Ribosome-associated protein biogenesis factors act during protein synthesis to facilitate modification, targeting and folding of the nascent polypeptide. Here, Bornemann et al.establish the dynamic interplay between these factors, thus providing new insight into the early steps of protein biogenesis.
- Thomas Bornemann
- , Wolf Holtkamp
- & Wolfgang Wintermeyer
-
Article |
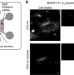 Local translation of TC10 is required for membrane expansion during axon outgrowth
Local translation of TC10 is required for membrane expansion during axon outgrowth
Axon growth requires exocyst-dependent membrane expansion, however it is unclear how this process is spatially regulated. Gracias et al.show that axonal translation of the exocyst regulator TC10 is necessary for stimulated membrane growth, and propose that local translation coordinates membrane and cytoskeletal enlargement.
- Neilia G. Gracias
- , Nicole J. Shirkey-Son
- & Ulrich Hengst
-
Article |
 60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle
60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle
Our understanding of ribosome biogenesis is limited by a lack of structural knowledge of assembly intermediates. Here, Leidig et al.report a high-resolution cryo-EM structure of a pre-60S particle that suggests that substantial rearrangements of the 5S RNP are required during ribosome maturation.
- Christoph Leidig
- , Matthias Thoms
- & Roland Beckmann
-
Article |
 Molecular basis for erythromycin-dependent ribosome stalling during translation of the ErmBL leader peptide
Molecular basis for erythromycin-dependent ribosome stalling during translation of the ErmBL leader peptide
In bacteria, the ribosomal stalling during translation of leader peptides is a mechanism of antibiotic resistance that has not been well understood. Here, the structure of a drug-dependent stalled ribosome complex has allowed the authors to propose a detailed mechanism for this translational arrest.
- Stefan Arenz
- , Haripriya Ramu
- & Daniel N. Wilson
-
Article |
 Kinetic modelling indicates that fast-translating codons can coordinate cotranslational protein folding by avoiding misfolded intermediates
Kinetic modelling indicates that fast-translating codons can coordinate cotranslational protein folding by avoiding misfolded intermediates
The speed of codon translation at the ribosome has a large bearing on the structure of the final protein, with faster rates thought to promote misfolding. Here the authors present a theoretical analysis suggesting that in some cases fast-translating codons may instead improve cotranslational folding.
- Edward P. O’Brien
- , Michele Vendruscolo
- & Christopher M. Dobson
-
Article
| Open Access Ribosome profiling reveals features of normal and disease-associated mitochondrial translation
Ribosome profiling reveals features of normal and disease-associated mitochondrial translation
Mitochondrial ribosomes are uniquely affected by mutations in the mitochondrial genome. By mapping the position of ribosomes on transcripts, the authors here reveal functional differences between mitochondrial and cytosolic ribosomes, and show that mutations in mitochondrial tRNAs induce ribosome stalling.
- Koos Rooijers
- , Fabricio Loayza-Puch
- & Reuven Agami
-
Article |
 VapC20 of Mycobacterium tuberculosis cleaves the Sarcin–Ricin loop of 23S rRNA
VapC20 of Mycobacterium tuberculosis cleaves the Sarcin–Ricin loop of 23S rRNA
Toxin–antitoxin systems have been implicated in the pathogenicity of Mycobacterium tuberculosis. Here, the authors study the function of the M. tuberculosistoxin VapC20 and show that it can impair protein translation and inhibit bacterial growth by cleaving the Sarcin–Ricin loop of 23S rRNA
- Kristoffer S. Winther
- , Ditlev E. Brodersen
- & Kenn Gerdes
-
Article
| Open Access A versatile cis-acting inverter module for synthetic translational switches
A versatile cis-acting inverter module for synthetic translational switches
Artificial genetic circuits have been designed to enable precise control of cellular behaviour and phenotypes. Saito and colleagues present a new RNA module that can invert the function of a translational OFF to an ON switch and demonstrate its utility in mammalian cells.
- Kei Endo
- , Karin Hayashi
- & Hirohide Saito
-
Article |
 Genome-wide search for exonic variants affecting translational efficiency
Genome-wide search for exonic variants affecting translational efficiency
Genetic effects on gene expression by variants at expression quantitative trait loci (eQTLs), can contribute to human genetic diseases. Here, Liet al. present a method to study eQTLs with effects on protein translation on a transcriptome-wide scale.
- Quan Li
- , Angeliki Makri
- & Hui-Qi Qu
-
Article |
 Deregulation of translation due to post-transcriptional modification of rRNA explains why erm genes are inducible
Deregulation of translation due to post-transcriptional modification of rRNA explains why erm genes are inducible
Erm methyltransferases confer antimicrobial drug resistance and their expression is induced by macrolides. Gupta et al.show that Erm-catalysed modification of rRNA affects synthesis of some proteins and reduces cell fitness, explaining why expression of Erm is deleterious in the absence of antibiotics.
- Pulkit Gupta
- , Shanmugapriya Sothiselvam
- & Alexander S. Mankin
-
Article |
 Translation of HTT mRNA with expanded CAG repeats is regulated by the MID1–PP2A protein complex
Translation of HTT mRNA with expanded CAG repeats is regulated by the MID1–PP2A protein complex
Expansion of CAG repeats in messenger RNAs is a common feature of various neurodegenerative disorders, including Huntington’s disease. Krauß et al.show that messenger RNAs with expanded CAG repeats bind to a protein complex that regulates translation and promotes overproduction of such aberrant proteins.
- Sybille Krauß
- , Nadine Griesche
- & Susann Schweiger
-
Article |
 Plant tumour biocontrol agent employs a tRNA-dependent mechanism to inhibit leucyl-tRNA synthetase
Plant tumour biocontrol agent employs a tRNA-dependent mechanism to inhibit leucyl-tRNA synthetase
Agrobacterium radiobacter strain K84 generates an antibiotic targeting pathogenic strains of Agrobacterium tumefaciens, enabling its use as a biocontrol to prevent infection of crops. Here the authors show that this antibiotic inhibits leucyl-tRNA synthetases via an unusual mechanism that depends on binding of tRNALeu.
- Shaileja Chopra
- , Andrés Palencia
- & John S. Reader
-
Article |
 Evolution of the protein stoichiometry in the L12 stalk of bacterial and organellar ribosomes
Evolution of the protein stoichiometry in the L12 stalk of bacterial and organellar ribosomes
The ribosomal stalk L12 is the only multi-copy protein in the ribosome and is essential for translation. Here Davydov et al.use a bioinformatics and mass spectrometry approach to study the evolution of L12 in bacterial ribosomes and predict its stoichiometry in a wide range of species.
- Iakov I. Davydov
- , Ingo Wohlgemuth
- & Marina V. Rodnina
-
Article
| Open Access Structural insights into protein-only RNase P complexed with tRNA
Structural insights into protein-only RNase P complexed with tRNA
RNase P is a key enzyme implicated in transfer RNA maturation that removes the 5′-leader sequences from transfer RNA precursors. In this study, a biophysical characterization of a novel protein-only variant of RNase P, known as PRORP (PROteinaceous RNase P), reveals that transfer RNA recognition by PRORP is similar to that by ribonucleoprotein RNase P.
- Anthony Gobert
- , Franziska Pinker
- & Philippe Giegé
-
Article |
 A structural basis for streptomycin-induced misreading of the genetic code
A structural basis for streptomycin-induced misreading of the genetic code
The antibiotic streptomycin increases errors in protein translation, but it is unclear how streptomycin exerts its effect on the ribosome. Demirci et al. present X-ray crystal structures that reveal conformational changes induced by streptomycin, which may inspire future efforts in antibiotics design.
- Hasan Demirci
- , Frank Murphy IV
- & Gerwald Jogl
-
Article |
 Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
In response to stress, yeast cells selectively translate proteins that can enhance cell survival. In this study, the authors find that tRNALEU(CAA)in yeast cells is modified in response to oxidative stress by a methyltransferase and this results in the selective translation of the mRNA for these genes.
- Clement T.Y. Chan
- , Yan Ling Joy Pang
- & Peter C. Dedon
-
Article |
 Prediction of variable translation rate effects on cotranslational protein folding
Prediction of variable translation rate effects on cotranslational protein folding
Proteins can undergo folding while being translated by the ribosome, and the extent of this folding is influenced by the rate at which amino acids are added to the nascent chain. This study provides a framework for predicting domain folding probabilities as a function of the kinetics of amino-acid addition.
- Edward P. O'Brien
- , Michele Vendruscolo
- & Christopher M. Dobson
-
Article
| Open Access A shift of the TOR adaptor from Rictor towards Raptor by semaphorin in C. elegans
A shift of the TOR adaptor from Rictor towards Raptor by semaphorin in C. elegans
What controls the binding partner selection of the target of rapamycin protein, TOR, is unknown. Using theCaenorhabditis elegans tail as a model, Nukazuka et al. determine that signals of semaphorin through plexin control the binding partner selection of TOR and are required for the correct organization of rays in the tail.
- Akira Nukazuka
- , Shusaku Tamaki
- & Shin Takagi
-
Article |
 Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Aminoacyl-transfer RNA synthetases are conserved between bacteria and eukaryotes; however, bacterial enzymes cannot acylate eukaryote tRNAs. Now, fusion of a human and bacterial enzyme is shown to overcome the species barrier and confer tRNA specificity during both codon selection and proofreading on the ribosome.
- Cuiping Liu
- , Howard Gamper
- & Ya-Ming Hou
-
Article |
 ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
Uridines at the wobble position of transfer RNA anticodons are usually modified to allow efficient decoding of messenger RNA codons. In this study, ALKBH8 is shown to be a bifunctional transfer RNA modification enzyme required for the formation of a novel diastereomeric pair of modified wobble uridines.
- Erwin van den Born
- , Cathrine B. Vågbø
- & Pål Ø. Falnes
-
Article
| Open Access Synthetic human cell fate regulation by protein-driven RNA switches
Synthetic human cell fate regulation by protein-driven RNA switches
The control of cell fate and apoptosis is a continuing challenge in synthetic biology. In this study, systems are developed in which an intracellularly expressed genome-encoded protein simultaneously achieves up- and downregulation of two distinct apoptosis pathways.
- Hirohide Saito
- , Yoshihiko Fujita
- & Tan Inoue

 Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA
Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA