News & Views |
Featured
-
-
Letter |
 Sequences enriched in Alu repeats drive nuclear localization of long RNAs in human cells
Sequences enriched in Alu repeats drive nuclear localization of long RNAs in human cells
A sequence that is frequently found in Alu elements drives the localization of some long RNAs to the nucleus in human cells.
- Yoav Lubelsky
- & Igor Ulitsky
-
Toolbox |
 A test drive of a DNA-analysis toolkit in the cloud
A test drive of a DNA-analysis toolkit in the cloud
The Bioconductor project gathers genomics tools and data into a handy package that can run in the cloud.
- W. Wayt Gibbs
-
Article |
 A single-cell survey of the small intestinal epithelium
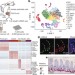
A single-cell survey of the small intestinal epithelium
Profiling of 53,193 individual epithelial cells from the mouse small intestine identifies previously unknown cell subtypes and corresponding gene markers, providing insight into gut homeostasis and response to pathogens.
- Adam L. Haber
- , Moshe Biton
- & Aviv Regev
-
Article
| Open Access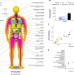 Genetic effects on gene expression across human tissues
Genetic effects on gene expression across human tissues
Samples of different body regions from hundreds of human donors are used to study how genetic variation influences gene expression levels in 44 disease-relevant tissues.
- François Aguet
- , Andrew A. Brown
- & Jingchun Zhu
-
Letter |
 Dynamic landscape and regulation of RNA editing in mammals
Dynamic landscape and regulation of RNA editing in mammals
Using the GTEx data and others, a comprehensive analysis of adenosine-to-inosine RNA editing in mammals is presented; targets of the various ADAR enzymes are identified, as are several potential regulators of editing, such as AIMP2.
- Meng How Tan
- , Qin Li
- & Jin Billy Li
-
Letter |
 Single-cell RNA sequencing reveals a signature of sexual commitment in malaria parasites
Single-cell RNA sequencing reveals a signature of sexual commitment in malaria parasites
Highly parallel single-cell transcriptome profiling of Plasmodium falciparum blood stages provides insight into the role AP2-G plays in early sexual development of this eukaryotic pathogen.
- Asaf Poran
- , Christopher Nötzel
- & Björn F. C. Kafsack
-
Article |
 The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation
The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation
Neuromedin receptor NMUR1 is specifically expressed by a subpopulation of type 2 innate lymphoid cells and promotes the inflammatory response of these cells in response to allergens, indicating the importance of neuro-immune crosstalk in allergic responses.
- Antonia Wallrapp
- , Samantha J. Riesenfeld
- & Vijay K. Kuchroo
-
Toolbox |
 Single-cell sequencing made simple
Single-cell sequencing made simple
Data from thousands of single cells can be tricky to analyse, but software advances are making it easier.
- Jeffrey M. Perkel
-
Letter |
 Multilineage communication regulates human liver bud development from pluripotency
Multilineage communication regulates human liver bud development from pluripotency
Single-cell RNA sequencing analysis of two- and three-dimensional hepatic differentiation reveals that both systems recapitulate certain transcriptomic features of human hepatogenesis.
- J. Gray Camp
- , Keisuke Sekine
- & Barbara Treutlein
-
Technology Feature |
 The RNA code comes into focus
The RNA code comes into focus
As researchers open up to the reality of RNA modification, an expanded epitranscriptomics toolbox takes shape.
- Kelly Rae Chi
-
Article |
 Intragenic DNA methylation prevents spurious transcription initiation
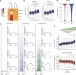
Intragenic DNA methylation prevents spurious transcription initiation
Intragenic DNA methylation, dependent on Dnmt3b, protects the gene body from spurious entry of RNA Polymerase II and aberrant transcription initiation events.
- Francesco Neri
- , Stefania Rapelli
- & Salvatore Oliviero
-
Letter |
 Single-cell spatial reconstruction reveals global division of labour in the mammalian liver
Single-cell spatial reconstruction reveals global division of labour in the mammalian liver
Single-molecule fluorescence in situ hybridization is performed to identify several landmark genes in the liver and their level of expression in single-cell RNA sequencing is used to spatially reconstruct the zonation of all liver genes.
- Keren Bahar Halpern
- , Rom Shenhav
- & Shalev Itzkovitz
-
News & Views |
 A new methyl mark on messengers
A new methyl mark on messengers
The presence of an N1 methyl group on adenine bases in DNA and RNA was thought to be a form of damage. Results now show that it also occurs at specific sites in messenger RNAs, where it affects protein expression. See Article p.441
- Anna M. Kietrys
- & Eric T. Kool
-
News & Views |
 Small RNA with a large impact
Small RNA with a large impact
A simultaneous comparison of the RNA molecules expressed by Salmonella bacteria and human cells during infection reveals how a bacterial small RNA alters the transcript profiles of both the bacteria and the host cells. See Article p.496
- Matthias P. Machner
- & Gisela Storz
-
Letter |
 Dynamic m6A mRNA methylation directs translational control of heat shock response
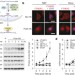
Dynamic m6A mRNA methylation directs translational control of heat shock response
Under stress, such as heat shock, the N6-methyladenosine (m6A) modification is shown to accumulate primarily in the 5′ untranslated region of induced mRNAs owing to the translocation of an m6A interacting protein, YTHDF2, into the nucleus, resulting in increased cap-independent translation of these mRNAs, indicating one possible mechanism by which stress-responsive genes can be preferentially expressed.
- Jun Zhou
- , Ji Wan
- & Shu-Bing Qian
-
News & Views |
 How to catch rare cell types
How to catch rare cell types
The development of an algorithm called RaceID enables the identification of rare cell types by single-cell RNA sequencing, even when they are part of a complex mixture of similar cells. See Letter p.251
- Lu Wen
- & Fuchou Tang
-
Letter |
 Structural imprints in vivo decode RNA regulatory mechanisms
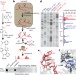
Structural imprints in vivo decode RNA regulatory mechanisms
The single-stranded nature of RNAs synthesized in the cell gives them great scope to form different structures, but current methods to measure RNA structure in vivo are limited; now, a new methodology allows researchers to examine all four nucleotides in mouse embryonic stem cells.
- Robert C. Spitale
- , Ryan A. Flynn
- & Howard Y. Chang
-
News & Views |
 RNA modification does a regulatory two-step
RNA modification does a regulatory two-step
The m6A structural modification of RNA regulates gene expression. It has now been found to mediate an unusual control mechanism: by altering the structure of RNA, m6A allows a regulatory protein to bind to that RNA. See Letter p.560
- Dominik Theler
- & Frédéric H.-T. Allain
-
Letter |
 N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions
N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions
The binding motifs for many RNA-binding proteins are normally buried within structured regions; now, the N6-methyladenosine modification is shown to act as a switch to remodel these regions, expose the motif, and thereby facilitate binding of RNA-binding proteins.
- Nian Liu
- , Qing Dai
- & Tao Pan
-
News & Views |
 Seeing the wood and the trees
Seeing the wood and the trees
The identification of the gene regulatory network that controls the formation of xylem — the major component of wood — opens up new avenues for manipulating plant biomass. See Article p.571
- Anthony Bishopp
- & Malcolm J. Bennett
-
Letter |
 A relative shift in cloacal location repositions external genitalia in amniote evolution
A relative shift in cloacal location repositions external genitalia in amniote evolution
It has been known for some time that limbs share at least some of their molecular patterning mechanism with external genitalia; here, this connection is examined in a variety of species, revealing that once-shared developmental trajectories could help to explain the observed patterning similarities.
- Patrick Tschopp
- , Emma Sherratt
- & Clifford J. Tabin
-
News & Views |
 Leaf veins share the time of day
Leaf veins share the time of day
Techniques for isolating and analysing leaf cell types have now been developed, leading to the discovery that circadian clocks in the plant vasculature communicate with and regulate clocks in neighbouring cells. See Letter p.419
- María C. Martí
- & Alex A. R. Webb
-
News & Views |
 A compass for stem-cell differentiation
A compass for stem-cell differentiation
The development of CellNet — network-biology software that determines how cell types generated in vitro relate to their naturally occurring counterparts — could improve our ability to produce desirable cells in culture.
- Franz-Josef Müller
- & Jeanne F. Loring
-
News & Views |
 Radiating genomes
Radiating genomes
Genome sequences and gene-expression data from representatives of five distinct lineages of African cichlid fish reveal signatures of the genomic changes that underlie the astounding cichlid diversity seen today. See Article p.375
- Chris D. Jiggins
-
Letter |
 Saturation editing of genomic regions by multiplex homology-directed repair
Saturation editing of genomic regions by multiplex homology-directed repair
The authors perform saturation mutagenesis of genomic regions in their native endogenous chromosomal context by using CRISPR/Cas9 RNA-guided cleavage and multiplex homology-directed repair; its utility is demonstrated by measuring the effects of hundreds to thousands of genomic edits to BRCA1 and DBR1 on splicing and cellular fitness, respectively.
- Gregory M. Findlay
- , Evan A. Boyle
- & Jay Shendure
-
-
Letter
| Open Access Comparative analysis of the transcriptome across distant species
Comparative analysis of the transcriptome across distant species
Uniform processing and detailed annotation of human, worm and fly RNA-sequencing data reveal ancient, conserved features of the transcriptome, shared co-expression modules (many enriched in developmental genes), matched expression patterns across development and similar extent of non-canonical, non-coding transcription; furthermore, the data are used to create a single, universal model to predict gene-expression levels for all three organisms from chromatin features at the promoter.
- Mark B. Gerstein
- , Joel Rozowsky
- & Robert Waterston
-
News & Views |
 Directions for the drivers
Directions for the drivers
A comparison of colorectal cancer and normal cells from 103 patients identifies dozens of genes that are differently expressed in tumour cells as a result of altered regulation of transcription. See Letter p.87
- Greg Gibson
-
News & Views |
 Fine-tuned amplification in cells
Fine-tuned amplification in cells
The transcription factor Myc has been posited to cause a cell-wide increase in gene expression. But two studies show that Myc, when modulated by other transcription factors, can amplify select targets. See Letters p.483 and p.488
- Chi V. Dang
-
News |
 Two crows kept separate by their looks
Two crows kept separate by their looks
Despite their differing colourings, carrion crows and hooded crows are almost identical genetically.
- Karen Ravn
-
Article |
 An atlas of active enhancers across human cell types and tissues
An atlas of active enhancers across human cell types and tissues
Using the FANTOM5 CAGE expression atlas, the authors show that bidirectional capped RNAs are a signature feature of active enhancers and identify over 40,000 enhancer candidates from over 800 human cell and tissue samples across the whole human body.
- Robin Andersson
- , Claudia Gebhard
- & Albin Sandelin
-
Article |
 A promoter-level mammalian expression atlas
A promoter-level mammalian expression atlas
A study from the FANTOM consortium using single-molecule cDNA sequencing of transcription start sites and their usage in human and mouse primary cells, cell lines and tissues reveals insights into the specificity and diversity of transcription patterns across different mammalian cell types.
- Alistair R. R. Forrest
- , Hideya Kawaji
- & Yoshihide Hayashizaki
-
Article
| Open Access Diversity and dynamics of the Drosophila transcriptome
Diversity and dynamics of the Drosophila transcriptome
A large-scale transcriptome analysis in Drosophila melanogaster, across tissues, cell types and conditions, provides insights into global patterns and diversity of transcription initiation, splicing, polyadenylation and non-coding RNA expression.
- James B. Brown
- , Nathan Boley
- & Susan E. Celniker
-
News |
 RNA activity mapped across cells
RNA activity mapped across cells
Technique adds spatial dimension to studies of gene expression.
- Brendan Borrell
-
Letter |
 Two independent transcription initiation codes overlap on vertebrate core promoters
Two independent transcription initiation codes overlap on vertebrate core promoters
The transcription start sites used during the maternal to zygotic transition in zebrafish are mapped, revealing that the transition is characterized by a switch between two different sequence signs to guide transcription initiation, which often co-exist in core promoters.
- Vanja Haberle
- , Nan Li
- & Boris Lenhard
-
Letter |
 Landscape and variation of RNA secondary structure across the human transcriptome
Landscape and variation of RNA secondary structure across the human transcriptome
An RNA secondary structure (RSS) map of coding and noncoding RNA from a human family (two parents and their child) is produced; this reveals that approximately 15% of all transcribed single nucleotide variants (SNVs) alter local RNA structure, and these SNVs are depleted in certain locations, suggesting that particular RNA structures are important at those sites.
- Yue Wan
- , Kun Qu
- & Howard Y. Chang
-
Research Highlights |
Tracking genes in myriad single cells
-
Article |
 Effect of natural genetic variation on enhancer selection and function
Effect of natural genetic variation on enhancer selection and function
Naturally occurring genetic variation between inbred mouse strains is used as a mutagenesis strategy to investigate mechanisms responsible for the selection and function of cis-regulatory elements in macrophages; lineage-determining transcription factors are proposed to select enhancer-like regions in the genome in a collaborative fashion and facilitate the binding of signal-dependent factors.
- S. Heinz
- , C. E. Romanoski
- & C. K. Glass
-
Article |
 Transcriptome and genome sequencing uncovers functional variation in humans
Transcriptome and genome sequencing uncovers functional variation in humans
Sequencing and deep analysis of mRNA and miRNA from lymphoblastoid cell lines of 462 individuals from the 1000 Genomes Project reveal widespread genetic variation affecting the regulation of most genes, with transcript structure and expression level variation being equally common but genetically largely independent, and the analyses point to putative causal variants for dozens of disease-associated loci.
- Tuuli Lappalainen
- , Michael Sammeth
- & Emmanouil T. Dermitzakis
-
Letter |
 MBNL proteins repress ES-cell-specific alternative splicing and reprogramming
MBNL proteins repress ES-cell-specific alternative splicing and reprogramming
This study identifies MBNL proteins as negative regulators of alternative splicing events that are differentially regulated between ES cells and other cell types; several lines of evidence show that these proteins repress an ES cell alternative splicing program and the reprogramming of somatic cells to induced pluripotent stem cells.
- Hong Han
- , Manuel Irimia
- & Benjamin J. Blencowe
-
Letter |
 Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell RNA sequencing is used to investigate the transcriptional response of 18 mouse bone-marrow-derived dendritic cells after lipopolysaccharide stimulation; many highly expressed genes, such as key immune genes and cytokines, show bimodal variation in both transcript abundance and splicing patterns. This variation reflects differences in both cell state and usage of an interferon-driven pathway involving Stat2 and Irf7.
- Alex K. Shalek
- , Rahul Satija
- & Aviv Regev
-
Letter |
 Extensive transcriptional heterogeneity revealed by isoform profiling
Extensive transcriptional heterogeneity revealed by isoform profiling
Variation among RNA transcript isoforms can be generated from alternative start and polyadenylation sites, and results in RNAs and proteins with different properties being generated from the same genomic sequence; here a new method termed transcript isoform sequencing is described in yeast, and the method allows a fuller exploration of transcriptome diversity across the compact yeast genome.
- Vicent Pelechano
- , Wu Wei
- & Lars M. Steinmetz
-
Letter |
 Control of somatic tissue differentiation by the long non-coding RNA TINCR
Control of somatic tissue differentiation by the long non-coding RNA TINCR
The human long non-coding RNA TINCR binds to STAU1 and controls epidermal differentiation by stabilizing key differentiation mRNAs, by means of a TINCR-binding motif found enriched in epidermal differentiation genes.
- Markus Kretz
- , Zurab Siprashvili
- & Paul A. Khavari
-
Article |
 An anatomically comprehensive atlas of the adult human brain transcriptome
An anatomically comprehensive atlas of the adult human brain transcriptome
Laser microdissection and microarrays are used to assess 900 precise subdivisions of the brains from three healthy men with 60,000 gene expression probes; the resulting atlas allows comparisons between humans and other animals, and will facilitate studies of human neurological and psychiatric diseases.
- Michael J. Hawrylycz
- , Ed S. Lein
- & Allan R. Jones
-
Letter |
 A transcriptomic hourglass in plant embryogenesis
A transcriptomic hourglass in plant embryogenesis
As it develops from a single-celled zygote to a mature plant embryo, the thale cress Arabidopsis thaliana passes through a stage during which phylogenetically very ancient genes are preferentially expressed, showing that animals and plants have independently acquired the developmental hourglass as a similar way of managing gene expression as they pass through embryogenesis, even though their morphological development is very different.
- Marcel Quint
- , Hajk-Georg Drost
- & Ivo Grosse
-
Article
| Open Access Landscape of transcription in human cells
Landscape of transcription in human cells
A description is given of the ENCODE effort to provide a complete catalogue of primary and processed RNAs found either in specific subcellular compartments or throughout the cell, revealing that three-quarters of the human genome can be transcribed, and providing a wealth of information on the range and levels of expression, localization, processing fates and modifications of known and previously unannotated RNAs.
- Sarah Djebali
- , Carrie A. Davis
- & Thomas R. Gingeras
-
Letter |
 Autoregulation of microRNA biogenesis by let-7 and Argonaute
Autoregulation of microRNA biogenesis by let-7 and Argonaute
MicroRNA in worms is shown to target non-coding primary microRNA transcripts through interaction with the Argonaute protein, promoting the production of further microRNA and thus generating a positive-feedback loop.
- Dimitrios G. Zisoulis
- , Zoya S. Kai
- & Amy E. Pasquinelli
-
Article |
 Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq
Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq
N6-methyladenosine (m6A) is the most prevalent internal modification in messenger RNA; here the human and mouse m6A modification landscape is presented in a transcriptome-wide manner, providing insights into this epigenetic modification.
- Dan Dominissini
- , Sharon Moshitch-Moshkovitz
- & Gideon Rechavi
-
Article |
 The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups
The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups
Integrative analysis of copy number and gene expression in 2,000 primary breast tumours with long-term clinical follow-up revealed putative cis-acting driver genes, novel subgroups and trans-acting aberration hotspots that modulate subgroup-specific gene networks.
- Christina Curtis
- , Sohrab P. Shah
- & Samuel Aparicio

 An inflammatory transcriptional switch
An inflammatory transcriptional switch Hiding in plain sight
Hiding in plain sight