Abstract
Discovering the structure and dynamics of transcriptional regulatory events in the genome with cellular and temporal resolution is crucial to understanding the regulatory underpinnings of development and disease. We determined the genomic distribution of binding sites for 92 transcription factors and regulatory proteins across multiple stages of Caenorhabditis elegans development by performing 241 ChIP-seq (chromatin immunoprecipitation followed by sequencing) experiments. Integration of regulatory binding and cellular-resolution expression data produced a spatiotemporally resolved metazoan transcription factor binding map. Using this map, we explore developmental regulatory circuits that encode combinatorial logic at the levels of co-binding and co-expression of transcription factors, characterizing the genomic coverage and clustering of regulatory binding, the binding preferences of, and biological processes regulated by, transcription factors, the global transcription factor co-associations and genomic subdomains that suggest shared patterns of regulation, and identifying key transcription factors and transcription factor co-associations for fate specification of individual lineages and cell types.
Similar content being viewed by others
Main
In multicellular organisms, transcription factors bind at cis-regulatory elements in the genome to mediate diverse gene expression programs with exquisite spatiotemporal control1,2,3. However, owing to the paucity of in vivo developmental stage transcription factor binding data and cellular transcription factor expression data, the integrated maps required to study transcriptional control of development with spatiotemporal resolution are lacking.
In this work, we analyse regulatory activity of a broad set of C. elegans transcription factors in one or more developmental stages. Exploiting recently developed methods4,5,6, we integrate transcription factor binding data with an initial cellular-resolution map of transcription factor expression in the embryo. Our integrated analyses support the discovery of many key transcription factors and candidate transcription factor co-associations for fate specification, providing insights into the temporal and spatial dynamics of regulatory interactions in development.
Large-scale survey of regulatory binding
As part of the modENCODE consortium, we performed 241 ChIP-seq experiments to identify in vivo binding sites for 92 (10%) C. elegans transcription factors and regulatory proteins (collectively termed factors) in one or more stages of development or treatments (Fig. 1a and Supplementary Table 1). To identify factor binding from the approximately 5.1 billion raw reads, we developed a uniform processing pipeline (Extended Data Fig. 1a–e and Methods) that enables comparison of orthologous transcription factor properties7, such as sequence preferences (Extended Data Fig. 1f–h). Eight previously reported8 experiments failed to pass our quality-control checks and were thus removed from consideration.
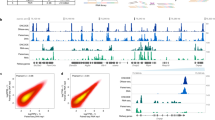
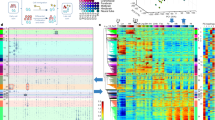
a, Transcription factors (TFs) and regulatory proteins assayed per developmental stage (or treatment) in 241 quality-filtered ChIP-seq experiments. Stages and treatments are early embryo (EE), late embryo (LE), embryo mixed (EM; EE and LE), larval L1 (L1), larval L2 (L2), larval L3 (L3), larval L4 (L4), young adult (YA), mixed larval and young adults (LY), day 4 adult (D4), and starved L1 (S1). Embryonic data sets were combined into a compiled embryonic stage (EX). Analyses in this report focus on embryonic (yellow) and larval (blue) experiments (N = 187). b, Genomic coverage (percent of genomic bases) of regulatory binding (excluding RNA polymerases) in 181 C. elegans (outer circle) and 339 H. sapiens (inner circle) ChIP-seq experiments. Genomic coverage of constitutive HOT (cHOT), HOT, and other regulatory binding (RGB) regions are highlighted in red, yellow and blue, respectively. Constitutive XOT (cXOT) and XOT percentages are shown in parenthesis. cHOT, HOT and RGB region coverage in the human genome are 0.17%, 1.4% and 6.1%, respectively7. c, Cut-off-normalized occupancy levels in 126 embryo-specific (yellow) and 91 larval L4-specific (blue) HOT regions. Bars indicate the 25th and 75th percentiles. d, Chromatin state, as determined in ref. 14, of L3 larvae binding regions by occupancy. RGB-, HOT- and XOT-region occupancy levels are indicated along the x axis as blue, yellow and red bars, respectively. Poly. Repress. and Heterochr. indicate Polycomb, repressed and heterochromatin states. e, f, Signal densities near enzymatically derived TSSs29 for BLMP-1 and ALY-2, and RNA Pol II. g, Functional (GO term) enrichment for gene targets of binding30. A subset of biological process terms (level ≥4) are shown for factors enriched (Benjamini–Hochberg-corrected, P < 0.01) in synaptic transmission; early MEP-1 and DPL-1 data sets are included for comparison. h, Example signal tracks near the UNC-104 locus.
We focused our analyses on embryonic and larval (L1–L4) stages, examining a total of 397,539 reproducible binding sites distributed across 33,833 binding regions in the genome. Collectively, factor binding (excluding RNA polymerases) is spread throughout 21.7% of the C. elegans genome (Fig. 1b), an upper-bound defined by ChIP-seq resolution9. We estimate that—within our ChIP-seq resolution and sensitivity—we have identified approximately 90% of the regulatory binding regions (albeit not the majority of binding events; Extended Data Fig. 1i). Consistent with this estimate, we observe binding within 2 kilobases (kb) upstream of a transcription start site (TSS) for 91.3% of genes (Extended Data Fig. 1j)10.
HOT regions are dynamic in development
Previous studies8,11,12,13 have revealed regions in metazoan genomes with heavily clustered transcription factor binding, termed high-occupancy target (HOT) regions. The availability of multiple data sets across stages allowed us to examine the dynamics of HOT regions through development. We identified HOT and extreme-occupancy target (XOT) regions for each developmental stage, where significant enrichments (false discovery rate (FDR) <5% and <1%, respectively) in transcription factor binding sites are observed (Extended Data Fig. 2a–c). We found a total of 9,142 HOT regions (spanning 2,948 genomic regions) in at least one developmental stage, and 858 constitutive HOT regions occurring across all stages assayed (Fig. 1b, Supplementary Table 2). Constitutive HOT regions are enriched in promoters of genes with house-keeping functions (Extended Data Fig. 2d and Supplementary Table 3). However, most HOT regions are dynamic across development: 31–56% of HOT regions change between sequential stages and occupancy at larval L4-specific HOT regions increases as development progresses (Fig. 1c).
Across developmental stages, 77–85% of HOT regions occur within 2 kb upstream of an annotated TSS (Extended Data Fig. 2e). Furthermore, 88.8% and 88.7% of constitutive HOT regions occur in promoter or enhancer states in embryos and L3 larvae, respectively (Extended Data Fig. 2f, g). These results indicate that HOT regions reside at important regulatory locations (Fig. 1d, Extended Data Fig. 3a–c) in the genome and are dynamic during development.
Preferences and roles of regulators
Factors displayed a range of chromatin state14 preferences, with a general bias towards promoter and enhancer states (Extended Data Fig. 3d, e). Although generally clustered near TSSs, many factors display enrichments for upstream or downstream binding (Extended Data Fig. 3f). Proximal and downstream binders include RNA Pol II (AMA-1) and other regulators of transcription initiation and elongation, respectively. Upstream binders may be enriched for chromatin remodellers and factors that recruit the transcriptional machinery. For example, binding of BLMP-1—the orthologue of the human repressor PRDM1 (refs 15, 16)—is tightly concentrated upstream of TSSs (Fig. 1e). Likewise, ALY-2, a human THOC4 messenger RNA export factor orthologue17, exhibits increased binding downstream of TSSs during development (Fig. 1e) and is increased at elongation chromatin states relative to other factors. Generally, transcription factors assayed in multiple stages retain their upstream and downstream binding preferences. Remarkably, RNA Pol II positioning shifts (Fig. 1f) from a strong elongating distribution in the early embryo to weaker elongation distributions in later stages, consistent with its previously observed continued presence at promoters that are down-regulated during development8,18.
Gene ontology (GO) analysis of the candidate protein-coding gene targets revealed 6,347 functional associations (BH-corrected, P < 0.05) for 75 factors (Extended Data Fig. 4a and Supplementary Table 4), suggesting biological roles for transcription factors of previously unknown function. The unstudied factors FKH-10 and C34F6.9 group with the established neuronal fate regulators SEM-4 (ZNF236), MAB-5 (HOXA2 and HOXB2), SAX-3 (ROBO1, ROBO2 and ROBO3), CES-1 and ZAG-1 in targeting neurotransmission genes, with C34F6.9 additionally regulating muscle development and sex differentiation (Fig. 1g). Most of these factors, including SEM-4 and C34F6.9 in L2 larvae, appear to regulate the neuronal kinesin UNC-104 (human KIF1A, KIF1C, Fig. 1h). Although expression of FKH-10 is restricted to six neuronal cells near the terminal bulb of the pharynx19, its specific molecular role in neuronal regulation and its regulatory targets were heretofore unknown.
Functional associations also demonstrate malleability of regulation. For example, UNC-62 transitions from regulating diverse muscle and neuronal development genes in embryos to regulating lipid metabolism processes in L4 larvae (Extended Data Fig. 4b). These changes are consistent with known diverse UNC-62 roles in motor neuron and vulval development, as well as locomotion, and ageing20,21. Similarly, SAX-3 transitions from targeting neuronal genes in L2 larvae to targeting carbohydrate and lipid metabolism genes in L4 larvae. The change in UNC-62 regulatory targets coincides with increased expression of the UNC-62 (7a) isoform in late larval and adult intestine20, which has been shown to affect lifespan22. Such early development regulators may often target metabolic regulation in later developmental stages23.
Global and subdomain transcription factor co-associations
Global analyses of pairwise transcription factor co-associations24 revealed a multitude of established and novel co-associations (Fig. 2a), many stage-specific clusters of co-association (Extended Data Fig. 5a), as well as differences in co-associations between expressed and repressed promoters (Extended Data Fig. 5b). FOS-1–JUN-1 as well as GEI-11–LIN-15B co-associations are readily apparent in L1 and L3 larvae, but not in L4 larvae. Likewise, ELT-3 and BLMP-1, which preferentially reside at molting and cuticle development gene promoters, co-localize in L1 larvae but not in embryos. The neuronal regulators CES-1 and FKH-10 co-associate across larval stages (L1, L3–L4) but their co-association is not apparent in late embryogenesis (Fig. 2a). Changes in co-association are often correlated with the presence of additional factors, for example, in the embryo to larval L1 transition, the increased ELT-3–BLMP-1 co-association is also accompanied by increased GEI-11 co-associations with these factors (Extended Data Fig. 5c–f). Other factors remain largely invariant through multiple stages, for example, ZTF-11, a human MTF1 orthologue.
a, Matrix of global pairwise (i, j) factor co-association strengths (N = 17,391) as defined by promoter interval statistics24. Co-association scores are scaled by the standard deviation (uncentred) for visualization purposes. Co-associations of interest and discussed in the text are highlighted. LX indicates larval stages L1–L4. A higher-resolution version is available in Extended Data Fig. 10. CES-1–FKH-10 co-associations are highlighted in the inset, top. Co-association strengths (unscaled) between early embryo and later stages are shown in the inset, bottom, for RNA Pol II-specific binding (blue), and for all factor-specific binding (light blue). b, Embryonic (EX) binding regions (N = 6,555) were clustered into a SOM describing 240 co-association patterns among 26 factors. c, Binding signatures (fraction of modules bound by each factor) of the learned co-association patterns are shown. The relative number of factors per co-association pattern, expression from overlapping promoters, distance to TSSs, and number of modules with each co-association pattern are indicated as a fraction of the maximum observed across co-association patterns. d, Functional enrichment for regions with UNC-62-bound co-association patterns of the embryo SOM. Molecular function (mf) and biological process (bp) terms are shown.
Functionally related factors were often co-associated. For example, FOS-1, NHR-77 and PQM-1 target promoters of genes in cellular lipid and ketone metabolic processes. Similarly, EFL-1 and LIN-35, the known interacting orthologues of human E2F and RB, show a strong co-association in L1 larvae, where they target membrane organization and endocytosis genes.
We observed strong similarities in RNA Pol II binding within embryonic (early and late embryo) and within larval L1–L4 stages, but larval RNA Pol II binding is only marginally and weakly co-associated with embryonic binding, reflecting the dynamic establishment of the transcriptional machinery through development (Fig. 2a).
To uncover higher-order co-associations (involving two or more factors), and their genomic subdomains, we applied self-organizing maps (SOMs), an unsupervised machine learning technique25. For each developmental stage, we trained SOMs to cluster genomic regions with shared transcription factor co-association patterns (Fig. 2b and Extended Data Fig. 6a–d), thereby concomitantly identifying transcription factor co-association patterns (Fig. 2c) and their target regulatory regions.
We performed GO analysis of the target regulatory regions for 240, 390, 439, 390 and 409 clusters in the embryo, larval L1, L2, L3 and L4 SOMs, respectively, revealing enrichments across 1,209 GO terms (BH-corrected, P < 0.05, Extended Data Fig. 6e and Supplementary Table 5). As illustrated in the embryo, higher-order co-association patterns show a richness of functional associations, with 137 clusters spanning 273 GO terms. A close examination of UNC-62 co-association patterns reveals how diverse patterns for individual factors can result in specialized functional targeting (Fig. 2d). Regions bound exclusively by UNC-62 and HLH-1 are highly-enriched at muscle development promoters. In contrast, genes targeted by more complex UNC-62 co-associations are enriched in synaptic transmission, regulation of cell death, and chromatin assembly functions. Higher-order co-associations are largely stage-specific (Fig. 3), a feature modulated by changes in the observed number of binding sites for individual factors between stages (Extended Data Fig. 7).
a, Embryonic (EX) and larval L1 binding SOM with raw binding sites. Binding data for factors (N = 15) assayed in embryos and L1 larvae was assigned to 25,261 stage-specific binding modules as shown in the inset. Stage-specific binding modules were clustered into an SOM describing 192 co-association patterns. The SOM is coloured by the embryonic (versus L1) stage specificity of the learned co-association patterns, measured as the fraction of binding modules that are embryonic. b, Histogram of preceding (T1) versus subsequent (T2) stage specificities.
Spatiotemporal transcription factor expression analysis
Although studies in C. elegans and D. melanogaster have led analyses of organismal-level regulatory binding circuits, such studies have generally lacked cell-type and tissue resolution. We sought to remedy this deficiency by tracking5,6 the expression of 180 diverse genes (mostly transcription factors) through early embryogenesis with cellular resolution (Extended Data Fig. 8a–d). Our expression data, from previously published5,6 and newly acquired series, includes 36 factors with genome-wide binding measurements (13 embryo, 23 larval).
We observed common and distinctive cellular expression patterns amongst a wide distribution of broadly- and narrowly-expressed genes (Extended Data Fig. 8e, f). For example, expression of DMD-4, an orthologue of the vertebrate spinal circuit configuration regulator DMRT3, is tightly limited to posterior regions of the pharynx. Similarly, F49E8.2 expression is exclusive to the Z2 and Z3 germ cells (Extended Data Fig. 8a). 95.7% of pairs of tracked cells show distinct gene expression signatures (R < 0.75).
Cellular expression mapped the regulatory activity of 16 assayed factors to specific tissues (Fig. 4a). As expected, the known regulators of pharynx and muscle development, PHA-4 and HLH-1, were respectively enriched in these tissues. The co-associated factors, MEP-1 and DPL-1 (human DP1 and DP2 orthologue), although broadly expressed, are enriched in neuronal lineages. This is consistent with the observed MEP-1 targeting of neuronal function genes in the larvae, and provides further support for the coordinate activities of MEP-1 and DPL-1 in targeting membrane organization, receptor-mediated endocytosis, and cell-cycle genes (Fig. 1f and Supplementary Table 4).
a, Tissue enrichment (−log10, P value) and depletion (log10, P value) scores for the expressing population of each gene are shown (Fisher’s exact, Bonferroni-corrected). Only genes with significant enrichments (or depletions) are shown. b, Co-association strength (Fig. 2a) versus cellular overlap coefficient for 13 focus factors. The Jaccard index for the cellular overlap is indicated for each gene pair by ring size and colour. c, Cellular-resolution regulatory binding SOM. Cellular-resolution binding modules were generated by annotating in each cell, the binding of focus factors expressed in the cell. Cellular-resolution binding modules (inset) were clustered into a SOM with 268 learned co-association patterns, 161 (68%) of which were discovered in the data. The SOM is colored by the number of factors in the learned co-association patterns. d, Tissue classes and co-association signatures are shown for 43 co-association patterns with significant enrichments. Tissue enrichments of interest are highlighted red. Circle size and colour indicate the Jaccard index (%) of cellular overlap. ‘Death’ and ‘Excretory’ represent tissue designations for cells with apoptotic and excretory cell fates.
More complex patterns of co-expression and co-association were observed in epidermal tissues, where CEH-16 (human Engrailed), and particularly ELT-1 and NHR-25 expression is concentrated. In both L2 and L3 larvae, ELT-1 and NHR-25 are modestly co-associated. ELT-1 targets transcriptional regulators, including NHR-25, and tail morphogenesis genes, whereas NHR-25 targets nuclear organization and genitalia development genes (Supplementary Table 4). However, larval L2 binding of ELT-1 and NHR-25 is co-associated with that of CEH-16, whose early embryonic expression is primarily concentrated in a subset of pharynx and epidermal cells.
In early embryo (Fig. 4a) and L1 larvae26, HLH-1, is primarily and broadly expressed in muscle tissues whereas posterior-specific HOX factors MAB-5, and EGL-5 are expressed in a small subset of posteriorly placed muscle, epidermal and neuronal precursors. We observed modest co-association signals between embryonic HLH-1 and MAB-5 binding, and larval L3 EGL-5 binding, perhaps reflecting the intersection of tissue-specific and positional regulatory programs. As expected, HLH-1 targets muscle differentiation genes (together with UNC-62); however in GO analysis, we only detect MAB-5 targeting of diverse neuronal functions (in mixed embryos and L2 larvae), consistent with its later role in neuron specification27. CO5D10.1, whose early embryonic expression is also restricted to muscle tissues is not co-associated with the above factors, and neither C05D10.1 nor EGL-5 showed specific functional associations. Thus, although co-associated factors were often expressed in the same tissue, this is not pervasive. Moreover, these co-expression patterns are dynamically established during embryogenesis (data not shown).
Refinement of embryonic co-associations
Despite extensive studies in metazoan regulatory networks, the relationship between regulator binding in overlapping genomic regions and co-expression in cell-types is not well studied. We examined this relationship among 13 ‘focus’ factors, for which both embryonic binding and cellular expression were assayed. This analysis is limited to the first half of embryogenesis, where expression was directly measured in 696 ‘focus’ cells. Later events may occur that would not be identified in our analysis. We found a poor correlation between transcription factor co-expression and co-association (R = 0.07, Fig. 4b), consistent with precise coordination of these separate processes underlying the differential establishment of cell- and lineage-specific regulatory circuits.
Integrated analysis shows that MEP-1 is co-associated and co-expressed with similarly broadly expressed factors (LIN-13, CEH-39) and narrowly expressed factors (CES-1, CEH-26), suggesting that MEP-1 often works in cis with these additional factors. MEP-1 binding is co-associated with CES-1 and CEH-26 in embryos, and expression of these factors is narrowly restricted within the MEP-1-expressing population. These MEP-1–CES-1 and MEP-1–CEH-26 co-associations are reminiscent of MEC-3–UNC-86 interactions in which the classic ‘terminal selector’ MEC-3 heterodimerizes with the broadly expressed UNC-86 exclusively in touch sensory neurons28. Thus, the co-association and co-expression of MEP-1–CEH-26 suggests CEH-26 may function as a terminal selector in head and tail neurons, and the excretory cell. The spatiotemporally-resolved co-association analyses demonstrate how broadly expressed factors, such as MEP-1 and LIN-13—which targets both neurotransmission functions and genitalia development—can have diversified functional roles during development through co-associations with narrowly expressed factors.
To determine how co-binding and co-expression co-ordinately define regulatory patterning in distinct cell-types and genomic regions, we intersected cellular expression and binding data by mapping focus factor binding to in silico genomes for cells where the factors are expressed. This procedure resulted in 4,779,810 binding sites distributed across 2,858,477 cell-resolved binding modules. We applied an SOM to cluster the cell-resolved binding modules by co-association patterns, uncovering 161 transcription factor co-association patterns and the genomic subdomains and specific cellular subsets of the embryo in which they may occur (Fig. 4c). The cellular distribution of transcription factor co-association patterns revealed co-associations shared among and unique to specific cell fates (Fig. 4d). For example, we found that distinct MEP-1, CEH-26 and NHR-2 co-associations were specific to neuronal tissues. Similarly, muscle cells were enriched in various HLH-1 co-associations.
We identified 39 co-association patterns whose cellular distribution coincides with the cellular expression of at least one of 124 target genes (non-focus factors; Bonferroni-corrected, P < 0.01). Focus factor binding allowed us to analyse co-association patterns at the promoters of 44 of these genes (where binding is observed). For 28 (63.6%) of these genes, co-association patterns were detected at the promoter and the gene’s cellular expression matched the cellular distribution. Moreover, the overlap between the expression cells for a gene and the co-association cells is higher in cases where the co-association occurs in the promoter of the gene (Wilcoxon, P = 5.1 × 10–6, Extended Data Fig. 8g). This result indicates that co-associations at promoters are correlated with cellular expression patterns for genes, and suggests a functional regulatory role for the discovered co-associations.
Discussion
We have generated a high-coverage transcription factor binding map of the C. elegans genome, revealing regulatory targets, co-associations, and dynamics across five developmental windows for 92 diverse factors. Gene targets suggest a multitude of functional associations for 75 factors, many previously unannotated and with clear mammalian homologues. Our work reveals extensive regulatory rewiring through development, with temporal differentiation of HOT regions in the genome, factor positioning preferences, regulatory targets, and co-associations.
A systematic analysis of transcription factor co-associations through development reveals sets of factors that assemble at genomic regions associated with more than 1,200 biological functions (GO terms), with probable spatiotemporal specificity. As illustrated with UNC-62, these higher-order co-associations reveal how individual transcription factors can participate in distinct transcription factor co-associations patterns at promoters of functionally diverse genes.
Lastly, cellular-resolution expression tracking allowed us to map the activity of 35 factors to precise cell and tissue types, demonstrating lineage-specific activities for 16 factors in the early embryo. Importantly, co-associations that are observed in whole-organism binding data are not always evident at the cellular level, highlighting the need to incorporate such information in our understanding of regulatory circuits. As additional expression patterns and transcription factor binding sites are determined, and methods to track transcription factor binding with cell-type specificity are developed, the broader and more precise regulatory logic of development should emerge.
Methods Summary
ChIP-seq assays of wild-type (N2) and transgenic nematodes were performed under controlled conditions (Extended Data Fig. 9) as described23. Experimental and computational methods are described in detail in the Methods.
Online Methods
This work builds on the goals of the modENCODE project31. Data from multiple stages of analysis in this work are available at http://encodeproject.org/comparative/regulation.
Strain construction
C. elegans strains were constructed essentially as described in ref. 32. In brief, each transgene fosmid constructed contains the entire transcription factor tagged at its carboxyl terminus with an in-frame green fluorescent protein (GFP):3 × FLAG tag. Transgenic strains were generated by microparticle bombardment of transgene fosmids. Twenty to fifty micrograms of fosmid DNA was used per transformation. The fosmid contained the unc-119 marker for selection of transgenic animals.
Strain growth and staging
Worms were grown on nematode growth medium (NGM) using standard growth protocols. Worms were synchronized by bleaching and L1 starvation, and grown to the desired developmental stage as determined by visual inspection33. In brief, animal populations consisting mostly of embryo-bearing adults were bleached and eggs were collected. Embryos were hatched in the absence of food to synchronize larval development, and then placed on food and grown for specified times to reach the appropriate larval or adult stage for collection and ChIP. To collect early stage embryos, young adult animals bearing relatively few embryos were collected and bleached. The subsequent embryos were mesh-purified and immediately fixed. To collect predominantly late stage embryos, the same procedure was used, except the embryos were incubated for six hours before fixing. All procedures result in a population synchronized within a 2-h developmental window. The vast majority of animals (>80%) are within this window.
Chromatin immunoprecipitation
C. elegans ChIP-seq assays were performed essentially as described in ref. 23. In brief, wild-type (N2) and transgenic worms expressing green-fluorescent protein (GFP)-tagged factors were grown to the desired developmental stage under controlled conditions (Extended Data Fig. 9) and cross-linked with 2% formaldehyde. Cell extracts were sonicated to yield predominantly DNA fragments in the range of 200–500 bp. For most experiments (∼93%), factor expression was driven by the endogenous promoters. With the exception of RNA Pol II (AMA-1), RNA Pol III (RPC-1), TBP-1, EOR-1 and EFL-1, where native antibodies were used, the sonicated lysates were immunoprecipitated using α-GFP antibody. Most immunoprecipitations were performed in 5% Triton, although a few were performed in 1% Triton. Direct comparison indicated that different concentrations of Triton had minimal effect on IP efficiency (data not shown). At least two biological replicates were performed for each ChIP, with parallel genomic DNA controls prepared from the same strain.
Library construction and sequencing
Sequencing libraries were prepared from independent biological replicates of immunoprecipitation-enriched and input DNA fragments. Libraries were multiplexed using four 4-bp barcodes34 and sequenced on an Illumina Genome Analyzer II.
Pre-processing of sequencing data
FASTQ files were aligned to the C. elegans ws220 genome with BWA35 and quality-filtered to retain only high-quality alignments (Q ≥ 30). As numerous ChIP and input DNA libraries were sequenced multiple times, we merge the sequencing files of re-sequenced libraries using the heuristics that follow. For each library with multiple re-sequencing files (instances), the following parameters are determined for each instance: aligned.reads = number of aligned reads; qc.reads = number of quality-filtered reads; qc.percent = percent of reads that pass quality filtering; qc.duplicates = fraction of quality-filtered reads that are duplicates (non-distinct).
For these libraries, these same metrics are calculated for all possible combinations of instances. Two additional metrics are calculated. Status is defined as ‘pass’ unless any of the constituent instances has <106 aligned reads or <20% quality-filtered reads (in which case the combination status is set to ‘fail’). In addition, we calculate the percent of effective alignments (qc.score) as a quality-control score for each combination. qc.score = qc.percent × (1 − qc.duplicate)
To select the best combination of instances, we choose the ‘passing’ combination that has ≥106 uniquely aligned reads. If no combination has status equal to ‘pass’, we choose the combination that has ≥106 uniquely aligned reads with the highest percent of effective alignments (qc.score). If no combination yields ≥106 uniquely aligned reads, all instances are used (that is, the combination with the highest number of reads is chosen). To perform uniform binding site identification on each data set (see below), we merge input DNA files from replicates into a single merged input DNA control.
Uniform binding site identification
All ChIP-seq experiments were scored against an appropriate input DNA control. For worm data sets, we used the SPP binding site caller to identify and score (rank) potential binding sites36. As described in ref. 7, we used the irreproducible discovery rate (IDR) framework for obtaining optimal thresholds and determine high confidence binding events by leveraging the reproducibility and rank consistency of binding site identifications across replicate experiments of each data set37. Code and detailed step-by-step instructions to call binding sites using the IDR framework are available at https://sites.google.com/site/anshulkundaje/projects/idr.
The SPP caller36 was used with a relaxed threshold (FDR = 0.9) to obtain a large number of binding sites (maximum of 30,000 for worm) that span true signal as well as noise (false identifications). Binding sites were ranked using the signal score output from SPP (which is a combination of enrichment over control with a penalty for binding site shape). The IDR method analyses a pair of replicates, and considers binding sites that are present in both replicates to belong to one of two populations: a reproducible signal group or an irreproducible noise group. Binding sites from the reproducible group are expected to show relatively higher ranks (ranked based on signal scores) and stronger rank-consistency across the replicates, relative to binding sites in the irreproducible groups. Based on these assumptions, a two-component probabilistic copula-mixture model is used to fit the bivariate binding site rank distributions from the pairs of replicates37.
The method adaptively learns the degree of binding site rank consistency in the signal component and the proportion of binding sites belonging to each component. The model can then be used to infer an IDR score for every binding site that is found in both replicates. The IDR score of a binding site represents the expected probability that the binding site belongs to the noise component, and is based on its ranks in the two replicates. Hence, low IDR scores represent high-confidence binding sites. An IDR score threshold of 5% for worm data sets was used to obtain an optimal binding site rank threshold on the replicate binding site sets (cross-replicate threshold). If a data set had more than two replicates, all pairs of replicates were analysed using the IDR method. The maximum binding site rank threshold across all pairwise analyses was used as the final cross-replicate binding site rank threshold.
Any thresholds based on reproducibility of binding site calling between biological replicates are bounded by the quality and enrichment of the worst replicate. Valuable signal is lost in cases for which a data set has one replicate that is significantly worse in data quality than another replicate. Hence, we used a rescue strategy to overcome this issue. To balance data quality between a set of replicates, mapped reads were pooled across all replicates of a data set, and then randomly sampled (without replacement) to generate two pseudo-replicates with equal numbers of reads. This sampling strategy tends to transfer signal from stronger replicates to the weaker replicates, thereby balancing cross-replicate data quality and sequencing depth. These pseudo-replicates were then processed using the same IDR pipeline as was used for the true biological replicates to learn a rescue threshold. For data sets with comparable replicates (based on independent measures of data quality), the rescue threshold and cross-replicate thresholds were found to be very similar. However, for data sets with replicates of differing data quality, the rescue thresholds were often higher than the cross-replicate thresholds, and were able to capture more binding sites that showed statistically significant and visually compelling ChIP-seq signal in one replicate but not in the other. Ultimately, for each data set, the best of the cross-replicate and rescue thresholds were used to obtain a final rank threshold. Reads from replicate data sets were then pooled and SPP was once again used to call binding sites on the pooled data with a relaxed FDR of 0.9. Pooled-data binding sites were once again ranked by signal score. The final rank threshold (best of cross-replicate and rescue threshold) was then used to threshold the ranked set of pooled-data binding sites.
All binding site sets were then screened against specially curated empirical blacklists for the worm genome. Briefly, these blacklist regions typically show the following characteristics: first, unstructured and extreme high signal in sequenced input DNA and control data sets as well as open chromatin data sets irrespective of developmental stage/treatment; second, an extreme ratio of multi-mapping to unique mapping reads from sequencing experiments.
The worm blacklist can be downloaded from https://www.encodeproject.org/comparative/regulation/#Wormset5.
ChIP-seq quality control
A number of quality metrics for all replicate experiments of each data set were computed38. In brief, these metrics measure ChIP enrichment and signal-to-noise ratios, sequencing depth and library complexity and reproducibility of binding site identification. These metrics are available through the ENCODE portal at http://encodeproject.org/comparative/regulation. We examined multiple quality-control thresholds, flagging data sets with low signal-to-noise ratios as determined by normalized strand cross-correlation scores (NSC < 1.03), low rank correlations between binding site scores across replicates (binding site rank correlation (RBS) < 0.3), or poor IDR models as indicated by a low correlation between binding site ranks and IDR ranks (binding site versus IDR rank correlation (RBI) < 0.3). A poor IDR model fit is a result of a pair of replicates having inseparable signal and noise components and abnormally low binding site rank consistency. Experiments that passed all quality-control thresholds were automatically scored as high-quality experiments. Experiments that passed most but not all quality-control thresholds where scored as medium-quality experiments. Experiments that did not pass multiple quality-control thresholds were discarded, excluded from further analyses with a few exceptions. As factors with genuinely few binding sites inherently have lower genome-wide signal-to-noise ratios, data sets with low NSC scores were rescued if the number of binding sites was low (<1,000). Analogously, high reproducibility scores (that is, low NP/NT ratios, see below) were occasionally allowed to rescue experiments where the IDR models appeared to have poor RBI values (<0.3) due to low numbers of binding sites. A summary of relevant quality metrics computed is provided below.
NP/NT ratio. This is the ratio of the number of binding sites passing 5% IDR thresholds based on comparison of pairs of pooled pseudo-replicates to pairs of biological replicates. The NP/NT ratio is a measure of reproducibility, computed as max(NP)/max(NT), where NP is the number of binding sites passing the 5% IDR threshold by comparing binding sites from a pair of pooled pseudo-replicates. The pair of pseudo-replicates is created by pooling reads from all replicates of a sample and randomly subsampling two equally sized sets of reads. NT is the number of binding sites passing the 5% IDR threshold by comparing binding sites from the best pair of biological replicates.
A high NP/NT ratio indicates that pooling replicates and subsampling substantially increased reproducibility in comparison to true replicates. This usually implies that at least one of the replicates has significantly higher enrichment as compared to others. The correlation between NP and NT across all experiments analysed is shown in Extended Data Fig. 1b.
Normalized strand cross-correlation (NSC). A genome-wide measure of ChIP enrichment or signal-to-noise ratio measure. A strand cross-correlation profile is computed as the Pearson correlation (y axis) between per-base read-start count vectors on the + and − strand over a wide range of strand shifts (x axis). The cross-correlation profile peaks at the predominant ChIP fragment length. The NSC is computed as the ratio of this maximal strand cross-correlation at the estimated fragment length (signal) to the minimum background cross-correlation over all shifts (noise). Samples for which both replicates had NSC < 1.03 are flagged as potential low signal-to-noise data sets. However, these can be rescued if the sample passes peak reproducibility criteria especially in cases in which the number of binding sites is low (<1,000).
Binding site rank correlation (RBS). Using the pre-IDR relaxed set of binding sites from the best pair of replicates, we find all binding sites that are present in both replicates. This set includes binding sites from the signal and noise components learned by the IDR model. We then compute the rank correlation of the binding site scores across the pair of replicates. Data sets with RBS < 0.3 are flagged as potentially low in binding site reproducibility.
Binding site versus IDR rank correlation (RBI). Using the pre-IDR relaxed set of binding sites from the best pair of replicates, we find all binding sites that are present in both replicates. These binding sites have scores from each of the replicates as well as an IDR score indicating the likelihood that the binding sites are not from the signal component. We rank the binding sites using the IDR scores and original binding site scores. For valid IDR models with good fits, the IDR scores and original binding site scores have a strong monotonic relationship and hence high rank correlation. Hence, we compute RBI as the rank correlation between the IDR scores and the original binding site scores as a measure of stability of the IDR models. Poor IDR model fits are usually a sign of abnormal rank consistency of binding sites and poor reproducibility. RBI is estimated as the primary data quality metric in that if a sample shows a poor IDR model fit no statements can be made about reproducibility. Data sets with RBI < 0.3 are considered to have poor IDR models. We make one exception for samples involving factors that bind few sites (<1,000) in the genome. In such cases, stable IDR models can obtain artificially low RBI scores. We perform additional tests of model stability for such samples, and allow for rescue if the models are deemed stable and if the NP/NT ratio is low.
ChIP-seq experiment selection
We uniformly processed approximately 5.1 billion raw reads from 323 worm ChIP-seq experiments, removing 82 (25%) low quality experiments that failed to meet our quality control standards (described above, Extended Data Fig. 1c). Examining approved experiments (Nr = 241), approximately 89% of the binding sites are shared between a pair of duplicate (redundant) experiments where binding was assayed for the same transcription factor and development stage (Nd = 22, Extended Data Fig. 1d). True biological duplicates—in which binding was assayed for the same developmental stage and factor, as driven by the same promoter, and assayed with the same ChIP protocol—share 77–92% of the binding sites. Thus, the identified binding sites have demonstrably reliable reproducibility rates.
We focused our analysis on a refined set of approved experiments (for 86 factors), selecting the highest-quality ChIP-seq data to produce a non-redundant set of embryo and larval experiments (N = 187) with unique factor and developmental stage combinations, prepared with the same ChIP protocol, and in which transcription factor expression is driven by the native promoter (Extended Data Fig. 1e). As such, the released collection corresponds to the top approximately 75% highest quality worm ChIP-seq experiments performed by the modENCODE consortium. Furthermore, the biological observations presented in this work stem from analysis of a top, non-redundant selection of embryo and larval experiments that collectively encompass approximately 58% of the worm ChIP-seq experiments performed.
Binding sites and reports for the released (Nr = 241) and analysed (N = 187) sets of ChIP-seq experiments are available online through the modENCODE data portal (http://encodeproject.org/comparative/regulation) and at http://tapanti.stanford.edu/cetrn.
Signal profiles
We generated signal track files for each ChIP-seq experiment using MACS2 (available at https://github.com/taoliu/MACS/) on pooled data (for ChIP and control), as follows:
macs2 callpeak -t ChIP.bam -c CONTROL.bam -B --nomodel --shiftsize round(FRAGLEN/2) --SPMR -g ce
where, --nomodel and --shiftsize round(FRAGLEN/2) tell MACS2 to use the estimated fragment length as fragment size (FRAGLEN, estimated in the uniform binding site identification pipeline) to pileup sequencing reads; -g ce lets MACS2 consider the C. elegans genome as background; and -B --SPMR indicate MACS2 to generate pileup signal files of ‘fragment pileup per million reads’ in bedGraph format.
To examine factor positioning preferences at high-resolution in each ChIP-seq experiment, we collected signal values per position (bp) within 1,000 bp of enzymatically enriched TSSs29 for protein coding genes. For visualization purposes (Fig. 1e, f and Extended Data Fig. 3f), we graph the scaled, mean signal density at each position, P(signal.density), calculated as:
P(signal.density) = (P(signal.mean) − min(signal.mean))/(max(signal.mean) − min(signal.mean))
where the average signal at any given position, P(signal.mean), is normalized to represent the fraction of the signal distance between the maximal average signal, max(signal.mean), and the minimal average signal, min(signal.mean). This normalization serves to correct signal:noise differences between ChIP-seq experiments.
For each factor and each ChIP-seq experiment, we calculated the log2-ratio of upstream to downstream binding in the windows >50 bp upstream and downstream from TSSs, respectively (Extended Data Fig. 3f).
Sequence preferences (motifs)
We examined C. elegans and H. sapiens binding sequence preferences7 among 21 transcription factor families, available from http://www.broadinstitute.org/~pouyak/motif-disc/integrate-cold/. In brief, these sequence preferences (motifs) were obtained by analysing sequence enrichment in the top 200 transcription factor binding sites from uniformly processed C. elegans (analysed here) and H. sapiens ChIP-seq experiments7. Sequence preferences were determined7 from transcription factor binding sites outside of HOT regions, un-mappable and blacklist regions, 3′ UTRs, and exons, and motif discovery was conducted using five discovery tools: AlignACE48 (v4.0 with default parameters), MDscan49 (v2004 with default parameters), MEME50 (v4.7.0 with -maxw 26 and -nmotifs 6), Weeder51 (v1.4.2 with option large), and Trawler52 (v1.2 with 200 random intergenic blocks for background). The top three motifs for each factor (and species) are selected after ranking by the enrichment in the data sets for the species and excluding motifs for which a similar motif was already selected (R > 0.7). The discovered motifs were augmented with known literature motifs in each gene family.
Among the 21 transcription factor families evaluated, C. elegans motifs were discovered for 15 transcription factor families (Extended Data Fig. 1f). We evaluated the prevalence of the discovered sequence preferences among binding sites from corresponding factors, scoring the fraction of binding sites with matches to the discovered motif for the top 200, 400, 600, 800 and 1,000 binding sites (Extended Data Fig. 1g). Motif matches in sequences were scored using the MAST module39 from MEME (v4.4), and applying an E-value cut-off equivalent to 10% of the input binding sites (FDR = 10%). For transcription factor families with multiple ChIP-seq experiments, we report the prevalence for the motif–ChIP-seq experiment combination with the highest correspondence. Across all binding site numbers evaluated, approximately 85% of the learned motifs have a prevalence exceeding 30% of the binding sites.
The C. elegans and H. sapiens motifs discovered for 12 transcription factor families in ref. 7 allow direct analysis of sequence preference conservation between these distant species (Extended Data Fig. 1f, h). We scored the similarity between the sequence preferences (motifs) of C. elegans and H. sapiens orthologous transcription factors within each family using the TOMTOM module40 from MEME (v4.4), qualifying significantly similar (P < 0.05) orthologous transcription factor sequence preferences as conserved (Extended Data Fig. 1f,h).
Chromatin states
Chromatin state and enhancer calls from C. elegans early embryos (EE) and stage 3 larvae (L3) were obtained from ref. 14. As recommended by the authors, we make use of the hierarchically-linked infinite hidden Markov model (hiHMM) segmentations reported14, examining 16 chromatin states derived from 8 histone marks.
Transcript expression analysis
The RNA-seq predicted transcripts per developmental stage, DCPM (depth of coverage per million reads) expression measurements for each gene or exon, TSS, transcription end site (TES), splice junctions, polyAs, and splice leader sites for C. elegans N2 early embryos (EE), late embryos (LE), and L1–L3 larvae were obtained as integrated transcript files from http://encodeproject.org/comparative/transcription.
HOT and XOT region determination
To identify regions with higher-than-expected binding occupancies, we first determined for each developmental stage the number and size distribution of observed binding sites for each factor assayed, as well as the total number and size distribution of binding regions in which these binding sites from all factors are clustered. For each developmental stage, we first analysed the number and size distribution of target binding regions (in which factor binding sites are concentrated). For each developmental stage simulation, we randomly selected an equivalent number of random binding regions with a matched size distribution. Next, for each factor assayed (in the target developmental stage), we evaluated the number and size of observed binding sites, and simulated an equivalent number and size distribution of target binding sites, restricting their placement to the simulated binding regions. We collapsed simulated binding sites from all factors into binding regions, verifying that these cluster into a similar number of simulated binding regions as the target binding regions. For each developmental stage simulation, the occupancy (number of binding sites), density (binding sites per kb), and complexity (diversity of factors) in the simulated binding regions are annotated. This procedure was repeated 1,000 times for each developmental stage. For each developmental stage, we constructed expected binding region occupancy (and density) distributions from the corresponding simulations (NS = 1,000). We determined the cut-offs at which fewer than 5% and 1% of the simulated binding regions have higher occupancies (Extended Data Fig. 2a). We classified observed binding regions with occupancies higher than the 5% and 1% cut-offs as high-occupancy target (HOT) and extreme-occupancy target (XOT) regions, respectively (Extended Data Fig. 2b, c). As such, HOT regions include XOT regions.
Recently, ref. 41 suggested regions with artefactual enrichment of ChIP-seq signals calling into question the validity of regions of high-occupancy where multiple ChIP-seq experiments produce enrichments. Using uniformly processed ChIP-seq binding sites7 from human cell-lines, we have established that our HOT regions are not an artefact of ‘hyper-ChIPable’ regions as described in ref. 41. In ref. 7, we have demonstrated that there is no correlation between our non-specific binding controls (IgG) and our measured transcription factor occupancy; that our HOT regions are not enriched in non-specific binding at any cut-off; and that non-specific binding can account at most for 0.5% of the binding signal as observed in RNA Pol II experiments. We note that the procedures used in ref. 41 are very different from ours and many others in the field. A brief discussion of these differences and their potential relevance to the results of ref. 41 follows.
The regions determined in ref. 41 have very low enrichment (twofold or less) of non-specific immunoprecipation in anti-GFP antibody controls over input DNA evaluated using a non-standard sliding-window approach. Importantly, immunoprecipitation/input ratios at this level are typically not considered enriched for binding in modern peak-calling procedures. For example, the median immunoprecipitation/input ratio for our human RNA Pol II experiments is 20-fold, and only 0.033% of human RNA Pol II peaks contain an immunoprecipitation/input ratio ≤ twofold. Thus, it is essential to note that the term ‘hyper-ChIPable’, coined by ref. 41, is quite misleading, as a correctly performed ChIP experiment will evaluate statistically enriched regions, with higher immunoprecipitation/input ratios. The so-called hyper-ChIPable regions in ref. 41 are not binding regions as determined under ChIP-seq best practices. Hence, when statistical peak-calling was performed in ref. 41 (using the established MACS peak-caller) to evaluate signals only at significantly enriched regions (Supplementary Table 1) only 17 (<7.5%) of the 238 claimed ‘hyper-ChIPable’ regions were called significant by all three Sir proteins. In fact, 68% of their 238 regions do not contain a binding site for any Sir protein as determined by MACS, despite even very liberal settings used (P < 10−5, no fold enrichment cut-off). Thus, the data of ref. 41 contradict its own major claim that all three Sir proteins showed enrichment at the 238 sites. Furthermore, as indicated in Supplementary Table 3 of ref. 41, the Sir2, Sir3 and Sir4 ChIP-seq experiments were performed only once each, which raises the question as to whether enrichment of Sir proteins at the 238 sites is reproducible. More rigorously, even for the remaining 17 genomic loci, their status as hyper-ChIPable is questionable as each region would first have to be established as a reproducible binding site in replicate experiments for each individual Sir protein. If you consider that Sir2, Sir3 and Sir4 ChIP-seq constitutes three replicates of Sir proteins, their data show that most of their claimed sites were not reproducibly enriched.
In addition to the analytical differences outlined above, other potential sources for the marked differences between our data and the Sir-enriched regions of ref. 41 are deviations from a typical ChIP protocol. In particular, ref. 41 employed a significantly longer cross-link time (1 h as opposed to the typical 10–20 min). This might contribute to formation of large non-specific protein–DNA complexes, which can in turn increase non-specific immunoprecipitation.
We believe that HOT regions, similar to other binding regions, are likely to reflect something other than a simple static model of transcription factor binding to DNA. Naturally, in the light of steric hindrances for large numbers of transcription factors in and the dynamic nature of molecular interactions, these high-occupancy regions may represent regions with diverse transient, or population-level diverse binding. Such a model is consistent with a known affinity for accessible DNA (as would be present in enhancer and promoter regions) and scanning mechanisms of transcription factor binding42. An alternative argument proposes HOT regions arise from multimeric transcription factor complexes that coordinately enrich genomic DNA from distinct loci. Thus, it is not clear that these regions are a meaningless artefact. In particular, these regions seem to segregate to enhancer and promoter regions with different chromatin architectures and different sets of transcription factors. Understanding how association and dissociation rates coordinately define residence time of transcription factor binding at individual sites, genome-wide and how chromosomal interactions relate to ChIP-seq signals will prove paramount to regulation but such analyses are outside the scope of this study.
Nevertheless, we have excluded HOT and XOT regions from sequence preference, functional, and global pairwise co-association analyses of factor binding. However, HOT (and XOT) regions were retained in self-organizing map (SOM) analyses since these analyses separate regions of high and lower occupancy.
Functional (GO term) enrichment analyses
To evaluate the functional role of regulators we performed GO enrichment analysis on the targets of binding of each ChIP-seq experiment. In brief, we applied ChIPpeakAnno30 to assign factor binding to genic targets as defined by binding within 1 kb of TSSs, and to evaluate the enrichment of genic targets for GO ontologies using standard procedures. We required a minimum of 20 binding sites per ChIP-seq experiment to evaluate enrichment and report Benjamini–Hochberg-corrected P values of enrichment (hypergeometric testing). We report GO terms in which at least one ChIP-seq experiment was significantly enriched (Benjamini–Hochberg-corrected, P < 0.05). The specific enrichments per HOT regions, per ChIP-seq experiment, and per stage-specific SOMs (see below) are provided in Supplementary Tables 3, 4 and 5, respectively. As high-occupancy can mask the biological significance of co-binding, sequence and target-gene specificity12, we focused our GO analysis on the 292,466 binding sites outside of XOT regions. Although we highlight GO terms in levels ≥4 in our figures, we report GO term enrichments in Supplementary Tables 3–5 without correcting for redundancy. As such, the raw GO term counts represent a serious overestimate, several-fold, of the number of distinct biological processes, molecular activities, or cellular components targeted by transcription factor binding but facilitate queries and analyses.
Global pairwise transcription factor co-associations
We determined the similarity in binding sites between ChIP-seq experiments applying recently developed interval statistics methods that allow calculation of exact P values for proximity between binding sites24. Using this method, we performed all pairwise, directional comparisons of ChIP-seq experiments evaluating binding similarity in 34,782 comparisons. To exclude the possibility of promiscuous binding regions and generate more conservative co-association estimates, we excluded binding sites from XOT regions in each developmental stage from these analyses (as above, see the previous section). We restrained interval analyses to the promoter domains by excluding binding intervals outside promoter regions, defined as 2,000 bp to 200 bp downstream of annotated TSSs. Focusing co-association analyses on the promoter domains serves to focus co-association evaluations on transcriptional regulatory interactions, and to account for the known biases in binding at TSSs, producing more conservative estimates of co-association significance. For each ChIP-seq experiment comparison (NC = 34,782), the intervals of the query ChIP-seq experiment are compared individually against all reference intervals of the reference ChIP-seq experiment, calculating the probability that a randomly located query interval of the same length would be at least as close to the reference set. For each ChIP-seq experiment comparison, we compute the fraction of proximal binding events in promoter domains that are significant (P < 0.05). As these comparisons are asymmetric—depending on the assignment of experiments as query or reference sets—we report the mean values of the complementary (inverted query and reference) comparisons and report this value as the ‘co-association strength’ (NT = 17,391) between ChIP-seq experiments. We refer to binding sites from pairs of ChIP-seq experiments as ‘co-associated’ if the co-association strength (unscaled) exceeds the 95th percentile of co-association strengths (CS95% = 0.4266, Extended Data Fig. 10) among comparisons of ChIP-seq experiments from distinct factors.
We examined co-association dynamics further by quantifying changes in co-associations (Δco-association) between factors assayed in sequential developmental stages. We were able to track 21 pairwise co-associations across all developmental stages and 78 across larval stages (Extended Data Fig. 5c–f). On average, 10% of the examined co-associations changed by more than 23.3% between sequential stages of development. Global co-association analysis was performed with an updated LIN-35 (L1) data set.
Stage-specific SOM analyses
Although global co-associations are useful surveys of factor co-binding, co-associations can have higher-order complexities involving three or more factors and vary between genomic subdomains. To uncover higher-order co-associations and the specific genomic subdomains in which they occur we applied SOMs, an unsupervised machine learning technique, in R using the kohonen package. Specifically, for each stage of development, we collapsed factor binding into developmental stage binding regions. For each binding region, we generate a binding module (for example, EX:I:10001174–10001734) with a binary signature indicating the presence or absence of binding (in the region) for each factor assayed in the developmental stage. For each stage, we generated a matrix of binding modules, and randomly seeded and trained 100 large, fine-grained SOMs to cluster binding modules by their binary signatures into coherent units (clusters) within a toroidal map. SOMs concomitantly discover common combinations of co-associated factors from the binary signatures (which we refer to as transcription factor co-association patterns) and assign binding modules (that is, the target regulatory regions) in which these combinations occur. Therefore, each cluster has a transcription factor co-association pattern (that is, a common set of co-associated factors) and a collection of putative target regulatory regions.
For each stage, we select the SOM with the lowest quantization error from the 100 trials for downstream analysis. Because we are interested in identifying transcription factor co-associations, we exclude binding modules from regions in which only one factor is bound from the matrix before SOM analyses. This approach generated maps with regulatory clusters that reveal how diverse transcription factor co-association patterns relate to target regulatory regions in the C. elegans genome at each stage (Fig. 2b, c and Extended Data Fig. 6a–d). For visualization and analysis of SOMs, we used a modified kohonen2 package25 and custom scripts.
Stage-comparison SOM analyses
To compare higher-order co-associations between sequential stages of development (T1 versus T2), we evaluated the relative representation of co-association patterns involving factors assayed in both stages of development. First, we collapsed binding across developmental stages into stage-independent binding regions. For each pair of stages to be compared (T1, T2), we generated a matrix combining stage-specific binding modules. Specifically, for each binding region we generated T1 and T2 binding modules (for example, EX:I:10001174–10001734 and L1:I:10001174–10001734) with the respective T1 and T2 binary signatures indicating the presence or absence of binding for each factor assayed in the two stages. We exclude binding modules from regions in which ≤ 1 factor is bound. We applied this approach to perform two types of comparative SOMs. In the first, we constructed such binding modules using all binding sites for each factor (that is, raw binding site model). In the second, we corrected for differences in binding site numbers for individual factors by sub-sampling binding sites from the stage with the higher binding site count (to those of the stage with lower binding site count). For this second approach (matched binding site model), we generated 100 such sub-sampled binding matrixes, and select the most representative matrix as that in which frequency of the individual binary signatures is best correlated with the frequency of binary signatures across the 100 sub-sampled matrixes (R > 0.9997). For both analyses, we then randomly seed and execute 100 SOMs to cluster binary signatures and select the SOM with the lowest quantization error for downstream analysis. To examine the stage-specificity of co-association patterns, we examined the relative abundance of T1 versus T2 binding modules per SOM cluster for each approach. Such stage-comparison SOMs were performed for sequential stages of development only (Fig. 3 and Extended Data Fig. 7e–g).
Cellular-resolution expression imaging and tracing
Embryonic lineage tracing and gene expression tracking were performed from both promoter reporter and protein fusion strains as previously described4,5,6,32,43. In brief, for target genes in promoter reporter strains, we cloned 2,250–5,750 bp upstream intergenic sequences (UIS) into pJIM20 (containing a cloning site followed by histone-mCherry and a permissive let-858 3′ UTR)43 using standard cloning methods. For each target promoter, we fixed the gene-proximal primer to the translation start site (including up to 6 amino acids of the endogenous protein). The resulting plasmids were used to generate transgenic C. elegans by microparticle bombardment of the strain CB4845[unc-119(ed3)] and histone::mCherry expression was tracked for at least three generations to verify stable inheritance. Promoter reporter strains were crossed with RW10029 to generate strains homozygous for the H3.3-GFP lineage tracing marker as well as for the histone-mCherry reporter. For protein fusion strains generated as part of the modENCODE project, we used strain RW10226 for the lineage tracing marker, and the colours were reversed for downstream analyses.
Strain imaging and lineage tracing was performed as previously described5,6,43, with lineages curated to at least the 350-cell stage. Expression values per cell were corrected for z-bias using a calculated attenuation level of 3.3% per plane5,43. Lineage data from each embryo was aligned to a reference lineage with standard cell cycle lengths44. We combined these data with previously published lineage data. The number of genes and image series from which expression data was derived is indicated below. The corresponding numbers of genes and image series previously published5,6 and recently acquired is as follows: compiled tracked genes = 180 (512 image series); previously published genes = 130 (324 image series); original report genes: = 50 (188 image series).
The cellular-resolution gene expression data are freely available for download through the Expression Patterns in C. elegans (EPIC) database (http://epic.gs.washington.edu) and via WormBase.
Cellular-resolution expression post-processing
For each gene, we obtained cellular-resolution expression measurements by assigning to each cell the average fluorescence signal from corresponding reporter experiments, and normalizing the signal in each cell by the maximum signal observed among imaged cells (Extended Data Fig. 8a and Supplementary Table 6).
We combed our imaging data to identify the set of cells tracked across all genes assayed (‘tracked’ cells), as well as the developmental time-point with the highest number of tracked cells. We directly measured expression of all 180 genes in a common set of 596 tracked cells, with maximal coverage of the embryo at 244 min of development, when 344 (98.3%) of the existing cells in the embryo have fluorescence measurements for all genes (Extended Data Fig. 8c, d). We refer to the set of factors (FF = 13) whose binding by ChIP-seq and expression by GFP reporters was measured in the embryo as the ‘focus’ factors. We identified the set of 696 cells for which expression of all 13 focus factors was directly measured and refer to this set of cells as the ‘focus’ cells.
As a heuristic to determine the population of cells in which a gene is expressed, referred to as the expressing population for the gene, we explored a range of expression cut-offs. We required a mean fluorescence signal ≥2,000 and chose 10% of maximal expression as the cellular expression cut-off on the basis of previous analysis5, as well as the strong and broad correlation in expression overlap with higher expression cut-offs, and its robust correlation with the quantitative expression of genes (Extended Data Fig. 8e). These expression calls revealed both distinctive and shared expressing populations for individual genes, and clusters of genes (such as a MEP-1-, CEH-39-, NHR-2-, NHR-28- and F23F12.9-containing cluster) with similar expressing populations (Extended Data Fig. 8f).
We derived gene expression values for the 671 terminal cells born during embryogenesis by ascribing to each cell its measured expression signal or that of its last measured ancestor. To examine lineage specificity of regulatory factors, we evaluated the enrichment of broad tissue classes in the expressing population of terminal cells of each gene.
Cellular-resolution expression data quality
For the vast majority of genes (approximately 80%), cellular expression signals were derived from multiple time-series (Extended Data Fig. 8b). Genes with multiple time-series have, on average, five time series recorded. Replicate time series (for 145 genes), allowed us to examine the correlation in cellular-resolution expression signals between N = 762 pairs of replicates (Extended Data Fig. 8b), revealing a median replicate signal correlation of R = 0.83. For genes with replicate time-series (N = 145), replicate time-series were strongly correlated (P < 10–11). The cellular overlap coefficient and Jaccard index between expressing populations of cells (A, B) as shown in Fig. 4b are calculated as:


As with the binding data, our embryonic, cellular expression data are unique in both resolution and scale. As such, homologous—quantitative, cellular-resolution, embryonic expression—measurements are not available (do not exist) for direct comparison. Nevertheless, we observe a high degree of correspondence between the cellular expression patterns of factors and previously published lineage involvements. Owing to our focus on integrating binding and expression data, only examples of correspondence for factors with both data types are highlighted in the main text. These include the previously known regulator of pharynx and muscle, PHA-4 and HLH-1, respectively. Our expression data show also shows consistencies between known, wide-spread roles of factors and cellular-expression breadth, as illustrated for MEP-1, an oocyte development zinc-finger protein required for maintenance of somatic versus germline differentiation45 that is broadly-expressed (Ncells = 379, 52% of examined cells). Undiscussed (but correlated) controls include the known regulators of intestine fate initiation and maintenance, ELT-2 and ELT-7 (ref. 46), the cell-body muscle-expressed helix–loop–helix factor, HND-1 (ref. 47), the pharyngeal-cell expression factor, CEH-34 (ref. 48), the human NeuroD homologue, CND-1 (ref. 49), and the hypodermally expressed molting factor, NHR-25 (ref. 50), among others.
Cellular-resolution SOM analyses
To integrate cellular-resolution expression and binding data, we simulated in silico genomes for each focus cell (FC = 696) and mapped (embryonic stage) focus factor (FF = 13) binding to the genome of cells in which factors are expressed in the early embryo. To examine physically plausible transcription factor co-associations and the cellular contexts wherein these may occur, we compiled the cellular-resolution binding data annotating binding modules per binding region, per cell. For each focus cell, we generate binding modules spanning each of the observed binding regions from the embryonic, organism-wide data, and annotate it with a binary signature describing which of the factors bound in the region (in the embryo) are expressed in the cell (in the early embryo). This approach resulted in 2,858,477 cellular-resolution binding modules (binding regions with cell identity; for example, ABalaa:I:10001174–10001734). We clustered cellular-resolution binding modules by their binding signatures in 100 separate SOMs, and selected the SOM with the lowest quantization error for downstream analysis (Fig. 4c, d). As before, we exclude binding modules from regions in which ≤ 1 factor is bound.
Lineage enrichment analyses
We constructed 3,915,749 cellular lineages in silico from the C. elegans embryogenesis cell-division tree. For each of the 696 focus cells, we generated up to 100,000 descendant lineages. We mined the cellular-resolution co-association map (Fig. 4c) for lineage-specific transcription factor co-association patterns by examining the enrichment (hyper-geometric) of cells in the co-association patterns discovered among the cells of each cellular lineage. We discovered significant overlaps involving eight transcription factor co-association patterns and 5 lineage nodes (Bonferroni-corrected, P < 0.01).
References
Davidson, E. H. Emerging properties of animal gene regulatory networks. Nature 468, 911–920 (2010)
Spitz, F. & Furlong, E. E. M. Transcription factors: from enhancer binding to developmental control. Nature Rev. Genet. 13, 613–626 (2012)
Lee, T. I. & Young, R. A. Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013)
Bao, Z. et al. Automated cell lineage tracing in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 103, 2707–2712 (2006)
Murray, J. I. et al. Multidimensional regulation of gene expression in the C. elegans embryo. Genome Res. 22, 1282–1294 (2012)
Mace, D. L., Weisdepp, P., Gevirtzman, L., Boyle, T. & Waterston, R. H. A high-fidelity cell lineage tracing method for obtaining systematic spatiotemporal gene expression patterns in Caenorhabditis elegans. G3 (Bethesda) 3, 851–863 (2013)
Boyle, A. P. et al. Comparative analysis of regulatory information and circuits across distant species. Nature http://dx.doi.org/10.1038/nature13668 (this issue)
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010)
Rhee, H. S. & Pugh, B. F. Comprehensive genome-wide protein-DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011)
Allen, M. A., Hillier, L. W., Waterston, R. H. & Blumenthal, T. A global analysis of C. elegans trans-splicing. Genome Res. 21, 255–264 (2011)
Moorman, C. et al. Hotspots of transcription factor colocalization in the genome of Drosophila melanogaster. Proc. Natl Acad. Sci. USA 103, 12027–12032 (2006)
Nègre, N. et al. A cis-regulatory map of the Drosophila genome. Nature 471, 527–531 (2011)
Yip, K. Y. et al. Classification of human genomic regions based on experimentally determined binding sites of more than 100 transcription-related factors. Genome Biol. 13, R48 (2012)
Ho, J. W. K. et al. Comparative analysis of metazoan chromatin organization. Nature http://dx.doi.org/10.1038/nature13415 (this issue)
Ohinata, Y. et al. Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436, 207–213 (2005)
Smith, M. A. et al. PRDM1/Blimp-1 controls effector cytokine production in human NK cells. J. Immunol. 185, 6058–6067 (2010)
Kuersten, S., Segal, S. P., Verheyden, J., LaMartina, S. M. & Goodwin, E. B. NXF-2, REF-1, and REF-2 affect the choice of nuclear export pathway for tra-2 mRNA in C. elegans. Mol. Cell 14, 599–610 (2004)
Baugh, L. R., Demodena, J. & Sternberg, P. W. RNA Pol II accumulates at promoters of growth genes during developmental arrest. Science 324, 92–94 (2009)
Hope, I. A., Mounsey, A., Bauer, P. & Aslam, S. The forkhead gene family of Caenorhabditis elegans. Gene 304, 43–55 (2003)
Van Auken, K. et al. Roles of the Homothorax/Meis/Prep homolog UNC-62 and the Exd/Pbx homologs CEH-20 and CEH-40 in C. elegans embryogenesis. Development 129, 5255–5268 (2002)
Curran, S. P. & Ruvkun, G. Lifespan regulation by evolutionarily conserved genes essential for viability. PLoS Genet. 3, e56 (2007)
Van Nostrand, E. L., Sánchez-Blanco, A., Wu, B., Nguyen, A. & Kim, S. K. Roles of the developmental regulator unc-62/Homothorax in limiting longevity in Caenorhabditis elegans. PLoS Genet. 9, e1003325 (2013)
Zhong, M. et al. Genome-wide identification of binding sites defines distinct functions for Caenorhabditis elegans PHA-4/FOXA in development and environmental response. PLoS Genet. 6, e1000848 (2010)
Chikina, M. D. & Troyanskaya, O. G. An effective statistical evaluation of ChIPseq dataset similarity. Bioinformatics 28, 607–613 (2012)
Xie, D. et al. Dynamic trans-acting factor colocalization in human cells. Cell 155, 713–724 (2013)
Liu, X. et al. Analysis of cell fate from single-cell gene expression profiles in C. elegans. Cell 139, 623–633 (2009)
Salser, S. J. & Kenyon, C. A. C. elegans Hox gene switches on, off, on and off again to regulate proliferation, differentiation and morphogenesis. Development 122, 1651–1661 (1996)
Hobert, O. Regulatory logic of neuronal diversity: terminal selector genes and selector motifs. Proc. Natl Acad. Sci. USA 105, 20067–20071 (2008)
Gu, W. et al. CapSeq and CIP-TAP identify Pol II start sites and reveal capped small RNAs as C. elegans piRNA precursors. Cell 151, 1488–1500 (2012)
Zhu, L. J., Gazin, C. & Green, M. R. ChIPpeakAnno: a bioconductor package to annotate ChIP-seq and ChIP-chip data. BMC Bioinformatics 11, 237–246 (2010)
Celniker, S. E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009)
Sarov, M. et al. A genome-scale resource for in vivo tag-based protein function exploration in C. elegans. Cell 150, 855–866 (2012)
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974)
Lefrançois, P. et al. Efficient yeast ChIP-Seq using multiplex short-read DNA sequencing. BMC Genomics 10, 37 (2009)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Kharchenko, P. V., Tolstorukov, M. Y. & Park, P. J. Design and analysis of ChIP-seq experiments for DNA-binding proteins. Nature Biotechnol. 26, 1351–1359 (2008)
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011)
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012)
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009)
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007)
Teytelman, L., Thurtle, D. M., Rine, J. & van Oudenaarden, A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc. Natl Acad. Sci. USA 110, 18602–18607 (2013)
Hammar, P. et al. Direct measurement of transcription factor dissociation excludes a simple operator occupancy model for gene regulation. Nature Genet. (2014)
Murray, J. I. et al. Automated analysis of embryonic gene expression with cellular resolution in C. elegans. Nature Methods 5, 703–709 (2008)
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983)
Unhavaithaya, Y. et al. MEP-1 and a homolog of the NURD complex component Mi-2 act together to maintain germline-soma distinctions in C. elegans. Cell 111, 991–1002 (2002)
Sommermann, E. M., Strohmaier, K. R., Maduro, M. F. & Rothman, J. H. Endoderm development in Caenorhabditis elegans: the synergistic action of ELT-2 and -7 mediates the specification→differentiation transition. Dev. Biol. 347, 154–166 (2010)
Mathies, L. D., Henderson, S. T. & Kimble, J. The C. elegans Hand gene controls embryogenesis and early gonadogenesis. Development 130, 2881–2892 (2003)
Hirose, T., Galvin, B. D. & Horvitz, H. R. Six and Eya promote apoptosis through direct transcriptional activation of the proapoptotic BH3-only gene egl-1 in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 107, 15479–15484 (2010)
Hallam, S., Singer, E., Waring, D. & Jin, Y. The C. elegans NeuroD homolog cnd-1 functions in multiple aspects of motor neuron fate specification. Development 127, 4239–4252 (2000)
Brooks, D. R., Appleford, P. J., Murray, L. & Isaac, R. E. An essential role in molting and morphogenesis of Caenorhabditis elegans for ACN-1, a novel member of the angiotensin-converting enzyme family that lacks a metallopeptidase active site. J. Biol. Chem. 278, 52340–52346 (2003)
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013)
Acknowledgements
We thank G. Euskirchen at the Stanford Center for Genomics and Personalized Medicine for sequencing ChIP libraries and members of the Reinke laboratory for contributing ChIP-seq data. We thank members of the Waterston laboratory, Sarov laboratory and Kim laboratory for tagged constructs and generating C. elegans strains. We thank J. Reuter, D. Phanstiel, H. Tilgner and D. Fowler for critical comments on the manuscript. This work is supported by the NHGRI as part of the modENCODE project (U01 HG004267).
Author information
Authors and Affiliations
Contributions
C.L.A., T.K., J.I.M., V.R., R.H.W. and M.P.S. designed experiments. T.K., L.J., L.M., W.N. and B.W. constructed strains and prepared ChIP libraries for sequencing. J.I.M., D.V., R.T., P.W., L.G. and D.M. constructed strains and performed reporter imaging experiments. A.K., C.L.A. and A.P.B. performed alignments, uniform binding site identification and quality filtering. C.L.A. organized data, pre-processed binding site and expression data, and executed all analyses. A.P.B., D.X. and C.L.A developed SOM analysis techniques. C.L.A., V.R., J.I.M., R.H.W. and M.P.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
ChIP-seq data reported here are available through the modENCODE DCC (http://www.modencode.org) and at http://tapanti.stanford.edu/cetrn.
Extended data figures and tables
Extended Data Figure 1 ChIP-seq uniform processing pipeline and quality controls.
a, ChIP-seq raw read data were processed using a uniform processing pipeline with identical alignment, filtering criteria, and standardized IDR binding site identification using SPP. b, Comparison of conservative (replicate) and pooled (pseudo-replicate) binding site calls from the cross-replicate and rescue thresholds, respectively. c, Distribution of NSC scores across 323 ChIP-seq experiments. Experiments are classified as high (blue, NHI = 181), medium (green, NMD = 60) and low quality (yellow, NLO = 82), and the relative fractions of each are indicated in the inset. High- and medium-quality experiments were approved for downstream analysis. d, The fraction of binding sites shared between duplicate, approved ChIP-seq experiments with (NU = 22) unique factor and stage combinations is shown. The fraction shared between the best-overlapping pairs of experiments with matched factor, stage combinations is shown in the light blue distribution. The fraction shared among all duplicates experiments (NP = 24) with matched factor, stage and promoter-driven transcription factor expression is shown in dark blue. The range of fractions shared between true biological duplicates (ND = 2) with matched factor, stage, promoter and ChIP protocol is indicated in dashed lines. For comparison, the fraction shared between randomly sampled pairs (NS = 500) of approved experiments from distinct factors is shown in grey. The median fractions for each distribution are shown. e, Binding site histogram for 187 embryo and larval ChIP-seq experiments with unique factor-stage combinations, and a common ChIP protocol, selected for analysis in this work. The fraction of high- (blue, NHI = 138) and medium-quality (green, NMD = 49) ChIP-seq experiments selected is indicated (inset). f, Analysis of sequence preferences for 21 C. elegans factors (NO) with human orthologue binding data7. The fraction of C. elegans factors for which sequence preferences could be determined (NM = 15, 71.4%) is shown (left). The fraction of factors with conserved sequence preferences (NC = 8, 66.7%, P < 0.05) from NX = 12 human–worm orthologues with determined sequence preferences is shown (right). g, The distribution in the fraction of binding sites with matches to the discovered preferred sequence (motif) is shown for 15 factors. The prevalence of the preferred sequence is evaluated among the top 200, 400, 600, 800 and 1,000 binding sites for each factor (see Methods). h, Discovered sequence preferences for 12 human or worm orthologues. Factors with similar (P < 0.05) and distinct sequence preferences are indicated in dark blue and light blue, respectively. The consensus sequence preference for the ONECUT3 homeobox factor was obtained from ref. 51. i, Saturation analysis of regulatory binding data. Using either binding data from embryonic and larval (L1–L4) stages or L2 larvae only (inset), k ChIP-seq experiments were randomly sampled (50 times each), collapsing overlapping binding sites into binding regions. For each k ChIP-seq experiment, the number of binding regions from 50 iterations is plotted (red points, ± 1 s.d.). For each series, an exponential curve (blue, dashed line) was fit to the data and used to estimate the total number of binding regions. The percentage of binding regions (CBP) observed in the acquired data are reported for each series. j, Amongst genes with annotated TSSs, the fraction of genes with binding observed within the specified window upstream of a TSS is shown. Promoter regions examined correspond to the windows (1) 1,000/100 bp, z(1) 2,000/200 bp, (3) 3,000/300 bp, (4) 4,000/400 bp and (5) 5,000/500 bp upstream or downstream of the TSS, respectively.
Extended Data Figure 2 Stage-dependent determination and analysis of HOT and XOT regions.
a, Correlations in occupancy (number of binding sites, x axis) and density (number of binding sites per kb, y axis) in embryo and larval L1–L4 binding regions. Quantiles for occupancy and density derived from binding site simulations are indicated on each axis. The fraction of binding regions (b) and the fraction of binding sites in regions (c) exceeding the significance cutoffs (quantiles from simulations) is indicated for both occupancy (yellow) and density (blue). Fractions exceeding cut-offs for both metrics are shown in red. Specific occupancy and density cut-offs for each significance level are indicated above each point. HOT (5% significance) and XOT (1% significance) regions exceed the specific occupancy thresholds indicated with arrows. d, GO enrichment analysis of constitutive HOT (cHOT), embryo, and larval L1–L4 HOT regions. For each stage, the non-cHOT-stage-derived HOT regions were analysed. GO enrichments in stage-specific HOT regions are available in Supplementary Table 3. e, The distribution of HOT region distances from annotated TSS in the C. elegans genome (ws220) is indicated for cHOT regions, non-constitutive HOT regions (non-cHOT), and stage-specific HOT regions. With the exception of larval L1-specific HOT regions, stage-specific HOT regions tend to be more distal. The overlap of HOT regions with embryonic (f) and larval L3 (g) chromatin states14 is indicated for cHOT, stage-derived HOT regions, and stage-specific HOT regions. With the exception of larval L1-specific HOT regions, cHOT regions show stronger promoter-associated chromatin states than non-constitutive HOT regions.
Extended Data Figure 3 Chromatin state distribution and positional preferences of regulatory binding.
a, Chromatin state distribution (y axis) of embryonic binding regions as a function of binding region occupancy (x axis). Embryonic binding regions with occupancies spanning 1–20 were mapped to 16 hierarchically linked infinite HMM (hiHMM) chromatin states14 discovered in embryos. Regulator binding regions (RGB)-, HOT-region and XOT-region occupancy levels are indicated along the x axis as blue, yellow and red bars, respectively. Chromatin state identities are indicated underneath. b, c, Fold change in frequency of chromatin states as a function of occupancy in embryos (b) and in L3 larvae (c). HOT and XOT cut-offs for each stage are indicated in dashed lines. d, e, Chromatin state distribution of factor binding in embryonic and larval L3 stages. Embryonic (d) and larval L3 (e) binding sites from individual ChIP-seq experiments were mapped to chromatin states derived from embryos and L3 larvae, respectively14. f, Signal densities near enzymatically-derived TSSs29. The log2 ratio of upstream (red) versus downstream (blue) binding is colour-coded below. Factors discussed in the text are highlighted.
Extended Data Figure 4 Functional enrichment analysis for gene targets of TF binding.
a, Gene ontology (GO) enrichment matrix for 150 binding experiments (75 factors) spanning 6,347 significant GO enrichments (Benjamini–Hochberg-corrected, P < 0.05) across 713 GO terms (level ≥4). For each experiment, GO-term enrichment was performed on gene targets as defined by binding within 1 kb of TSSs (ChipPeakAnno)30. Enrichments for biological process (bp) and molecular function (mf) ontologies are shown, with distinct sets of enrichments highlighted (i–viii). b, GO term enrichments among targets of UNC-62 binding show dramatical changes in the functional role of UNC-62 regulatory activity through development. Biological process terms (level ≥4) enriched in UNC-62 libraries are shown. The number of UNC-62 binding sites identified per stage is indicated in parenthesis. Although changes in targets between mid-larval and adult stages have been suggested previously22, our analyses (performed with uniformly called binding sites) and expanded data indicate that the most dramatic changes occur between embryo and L4 larval stages. (+) MEP-1 indicates experiments performed in strain OP102.
Extended Data Figure 5 Structure of global co-associations and changes between stages and domains.
a, Clustering patterns in pairwise transcription factor co-associations. Clustered libraries from shared factors are coloured blue. Clustered embryonic libraries are coloured yellow. ChIP-seq libraries that cluster in embryonic groups and with distinct stages for the same factor are coloured green. BLMP-1 and ELT-3 libraries are colored purple. FOS-1 and JUN-1 libraries are coloured red. All other libraries are colored grey in the dendrogram. The clustering dendrogram is derived from Fig. 2a. b, Difference in pairwise transcription factor co-associations at expressed and repressed promoter domains. For embryonic and larval L1 stages, we computed co-association strength 2 kb upstream and 200 bp downstream domains of TSSs associated with expressed and repressed genes, from stage-specific binding experiments with IntervalStats24. For each comparison (and each domain), the difference in the strength of co-associations between the expressed and repressed domains is shown for embryo (bottom left) and larval L1 stages (top right). Positive values indicate stronger co-associations in the expressed domain whereas negative values indicate stronger co-associations in the domain of repressed promoters. c–f, Change in pairwise transcription factor co-associations across sequential developmental stages. For factors assayed in sequential developmental stages, the difference in the co-association strengths for pairs of factors is shown. The change in co-association strengths are shown for the embryo to larval L1 (c), larval L1 to L2 (d), larval L2 to L3 (e), and larval L3 to L4 transitions (f). Co-association strengths for pairs of factors at each stage are derived from Fig. 2a.
Extended Data Figure 6 Stage-specific analysis of higher-order co-associations in the larvae.
For each larval stage of development, binding regions were annotated with binary signatures indicating the presence or absence of factor binding and clustered into SOMs describing the co-association patterns amongst factors assayed in each stage. a–d, SOMs are coloured by the number of factors per co-association pattern with respective patterns in each cluster are indicated underneath. e, For each co-association pattern discovered in stage-specific SOMs, GO enrichment analysis was performed on genes associated by binding within 1 kb of TSSs (ChipPeakAnno)30. GO terms are arranged along the circumference of the graph, and their enrichment is indicated in each stage. The inner-most layer contains the gene ontology colour key as indicated and subsequent layers (from the centre) indicate embryonic (EX), L1, L2, L3 and L4 enrichment of each GO term. For visualization purposes, only GO terms with 5 ≤ annotated genes ≤ 25 (NGO = 419) are shown.
Extended Data Figure 7 Stage-comparison SOMs highlight patterns in the specificity of higher-order transcription factor co-associations.
a, Abundance of co-association patterns is graphed as function of the number of factors in each co-association in stage-comparison SOMs for the embryo versus larval L1 stage comparison. Similar patterns are observed in all stage-comparisons SOMs. b, Difference in binding sites between embryos and L1 larvae for each factor (grey dots). The fractional difference, calculated as fraction of the larger set of binding sites represented by the difference in binding sites, is shown. Factors are rank-ordered by their difference in binding sites. The fraction of co-association patterns that are stage-specific (≥90% embryonic or larval L1) in SOMs is indicated for the raw binding sites with all factors (Fig. 3a, dashed line), in SOMs with individual factors removed (blue), and in SOMs with factors sequentially removed (red). c, Embryonic and larval L1 binding SOM with matched numbers of binding sites. Briefly, binding data for the 15 factors assayed in the embryo and L1 larvae was sub-sampled to generate stage-specific binding modules with equal numbers of binding sites for each factor (see Methods). Stage-specific binding modules with matched binding sites were clustered in an SOM describing 140 co-association patterns. SOM is coloured as in Fig. 3a. d, Binding signatures (fraction of modules bound by each factor) are shown for each co-association pattern from c. Sidebar indicates the embryonic (versus L1) stage-specificity of each co-association pattern as in c. Stage-comparison SOMs with raw and matched binding sites are presented for the larval L1 versus L2 comparison (e), larval L2 versus L3 comparison (f), and larval L3 versus L4 comparison (g). Binding region comparisons are performed as in Fig. 3. Briefly, binding data for factors assayed in sequential stages are assigned to stage-resolved binding modules (that is, L1:I:10001174–10001734). Stage-resolved binding modules are clustered into SOMs describing shared and stage-specific co-association patterns. SOMs are colored by the T1 versus T2 (for example, L1 versus L2) stage-specificity of the learned co-association patterns, measured as the fraction of binding modules that are T1. T1- and T2-specific co-association patterns are shown in red and blue, respectively. Sidebars indicate the T1 (versus T2) stage-specificity of each co-association pattern. As in Fig. 3, SOMs with matched binding sites were generated by sub-sampling binding sites to generate stage-resolved binding modules with equal numbers of binding sites for each factor. For each comparison, the most representative sampling (from 100 iterations) was selected to seed SOM analyses. For each of the stage-comparison SOMs with matched binding sites (e–g), the matrix of learned co-association patterns (fraction of modules bound by each factor) are shown below each SOM. h–j, The fraction of co-association patterns that are stage-specific (≥90% either stage) in SOMs is indicated for the raw binding sites with all factors assayed in both stages (dashed line), in SOMs with individual factors removed (blue), and in SOMs with factors sequentially removed (red) are shown for the larval L1 and L2 stage (h), larval L2 and L3 stage (i), and larval L3 and L4 stage (j) comparisons.
Extended Data Figure 8 Cellular resolution tracking of protein expression in the C. elegans embryo.
a, Cellular-resolution, protein expression levels for 180 genes (x axis) in terminal embryo cells (N = 671, y axis). For each gene, the normalized expression signal in each cell is shown (see Methods). For each gene, expression signals in cells not measured directly correspond to the expression signal of the last measured ancestor. Focus factors (FF = 13) whose binding was assayed in embryonic stages are labelled red. Factors whose binding was assayed only in larval stages are labelled blue (FL = 23). The broad tissue class of each cell is indicated in the sidebar. b, Embryonic, cellular-resolution expression data quality controls. The number of time-series recorded per gene (x axis) is shown. For genes with multiple time-series (NGR = 145), the Pearson correlation coefficient (R) in the fluorescence signals of cells recorded was calculated between NPR = 762 pairs of time-series (replicates). The distribution of correlation coefficients is shown. The median correlation co-efficient among replicate experiments is shown (R = 0.8310). The number (c) and percentage (d) of embryonic cells with expression measurements across any of the assayed genes (assayed cells, grey), all of the assayed genes (tracked cells), and all of the 13 genes (focus factors) for which both embryonic binding data and cellular-resolution expression data was acquired (focused cells) are plotted as a function of developmental time (Sulston minutes). The specific developmental times with the maximum coverage of the cells in the embryo are indicated for the tracked (TT) and focused cells (TF). e, Previously, Murray et al.5 suggested that a robust heuristic to identify cells in which individual genes are expressed can be obtained by requiring a fluorescence signal ≥2000 and a fluorescence signal that is ≥10% of the maximum signal observed for each reporter (gene). To confirm these recommendations, we calculated the overlap in the expressing cell populations for pairs of genes at 10% (ƒ = 0.1) and 20% (ƒ = 0.2) of the maximal signal for each gene, and computed the correlation between calculated overlaps per gene-pair between the two thresholds (R = 0.94). This analysis was extended to compare a wide range of expression cut-offs (ƒ) in e, where we observed robust correlations for the 10% cut-off (ƒ = 0.1). f, Cellular expression overlap matrix for 180 genes in the early embryo. For each pairwise gene comparison, we calculated the significance of the overlap between the population of cells expressing each gene. The overlap enrichment and depletion P values between gene pairs were determined using directional Fisher’s exact tests and were Benjamini–Hochberg corrected. To generate a final overlap score, we select the most significant of the enrichment and depletion scores, reporting either the -log10(P value of enrichment) or the log10(P value of depletion) to obtain positive and negative values for enrichment and depletion, respectively. g, Overlap between co-association cells and the gene-expressing cells (the expressing population) for non-focus factors (NNF = 168). For each cellular-resolution co-association pattern discovered (Fig. 4c), the set of co-association cells is defined as the population of cells in which the co-association is observed in the SOM. For 39 co-association patterns, co-association cells significantly overlap (hypergeometric test, Bonferroni-corrected, P < 0.01) the gene-expression cells of at least one of 124 non-focus factor target genes. Co-association patterns and target gene pairs with significant overlaps between the co-association cells and gene-expression cells were classified as ‘co-association in promoter’ if the co-association pattern with the significant enrichment was observed at the promoter at the target gene, and as ‘co-association not in promoter’ if this was not the case. The distribution of overlap significance values for the two classes and the respective Wilcoxon test P value for similarity between the two distributions is shown. MEP-1 (+) indicates experiments performed with strain OP102.
Extended Data Figure 9 Representative samples of staged, transgenic C. elegans embryos and larvae expressing GFP-tagged fusion proteins.
GFP fluorescence images, differential interference contrast (DIC) images, and merged (GFP/DIC) images are labelled with green, white and blue dots, respectively. The 10-µm scale bar is shown in GFP fluorescence images. Images were selected independent of binding experiment results. Approved binding experiments include: MEP-1 (mixed embryo, L2 larvae), DPL-1 (L1 larvae), C27D6.4 (L2 larvae), NHR-23 (L3 larvae) and CEH-16 (L4 larvae) experiments.
Extended Data Figure 10 Full-resolution view of global pairwise transcription factor co-association matrix.
As outlined in Fig. 2a, the significance of co-binding (co-association strength) 2 kb upstream and 200 bp downstream of TSSs was measured reciprocally between all binding experiments (IntervalStats24, see Methods). For each comparison (NC = 34,782), the fraction of significant (P < 0.05) co-binding events was computed and the mean fraction of reciprocal tests is reported (NT = 17,391). Co-association scores are scaled by the standard deviation (uncentred) for visualization purposes. Co-associations were examined among 292,466 binding sites outside of XOT regions. Inset (i) shows the distribution of global transcription factor co-association strengths from pairwise comparisons of 187 ChIP-seq experiments. The distribution of co-association strengths is shown from comparisons of all (distinct) ChIP-seq experiments (NDE = 17,391, light blue) and from comparisons of ChIP-seq experiments from distinct factors (NDF = 17,197, dark blue). The 75th, 90th and 95th percentiles from comparisons between distinct factors (CS75% = 0.2437, CS90% = 0.3589 and CS95% = 0.4266) are indicated as light red, red and dark red dashed lines, respectively. Co-association strengths between FOS-1–JUN-1 in L1, L3 and L4 larvae are indicated with arrows. Inset (ii) highlights the similarity (Wilcoxon test, P = 0.4913) between distributions from distinct factors and distinct experiments.
Supplementary information
Supplementary Data
This zipped file contains Supplementary Tables 1-6. (ZIP 48559 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Araya, C., Kawli, T., Kundaje, A. et al. Regulatory analysis of the C. elegans genome with spatiotemporal resolution. Nature 512, 400–405 (2014). https://doi.org/10.1038/nature13497
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13497
This article is cited by
-
The impact of species-wide gene expression variation on Caenorhabditis elegans complex traits
Nature Communications (2022)
-
A 4D single-cell protein atlas of transcription factors delineates spatiotemporal patterning during embryogenesis
Nature Methods (2021)
-
Reverse engineering highlights potential principles of large gene regulatory network design and learning
npj Systems Biology and Applications (2017)
-
Genome-wide identification and characterisation of HOT regions in the human genome
BMC Genomics (2016)
-
A quantitative genome-wide RNAi screen in C. elegans for antifungal innate immunity genes
BMC Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.