Featured
-
-
Article |
 Crystal structure of the 14-subunit RNA polymerase I
Crystal structure of the 14-subunit RNA polymerase I
RNA polymerase (Pol) I transcribes ribosomal RNA that is critically required for ribosome assembly, and the enzyme is a major determinant of protein biosynthesis and cell growth; here the crystal structure of the complete 14-subunit Pol I from yeast is determined, providing insights into its unique architecture and the possible functional roles of its components.
- Carlos Fernández-Tornero
- , María Moreno-Morcillo
- & Christoph W. Müller
-
Letter |
 A high-resolution map of the three-dimensional chromatin interactome in human cells
A high-resolution map of the three-dimensional chromatin interactome in human cells
A novel approach to analyse high-depth Hi-C data provides a comprehensive chromatin interaction map at approximately 5–10 kb resolution in human fibroblasts; this reveals that TNF-α-responsive enhancers are already in contact with target promoters before signalling and that this chromatin looping is a strong predictor of gene induction.
- Fulai Jin
- , Yan Li
- & Bing Ren
-
Article |
 Genomic organization of human transcription initiation complexes
Genomic organization of human transcription initiation complexes
The ChIP-exo technique is used to map the organization of transcription initiation complexes across the human genome at near-base-pair resolution; most of the transcription initiation complexes give rise to non-coding, non-polyadenylated RNA, indicating that pervasive non-coding transcription arise from specific promoters and is regulated.
- Bryan J. Venters
- & B. Franklin Pugh
-
Letter |
 lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs
lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs
A study of prostate cancer cells reveals a transcriptional activation role for long non-coding RNAs (PRNCR1 and PCGEM1) that bind to the androgen receptor, and is also observed for the truncated androgen receptor characteristic of many aggressive prostate cancers.
- Liuqing Yang
- , Chunru Lin
- & Michael G. Rosenfeld
-
Letter |
 Activity-dependent phosphorylation of MeCP2 threonine 308 regulates interaction with NCoR
Activity-dependent phosphorylation of MeCP2 threonine 308 regulates interaction with NCoR
Rett syndrome is caused by mutations in MeCP2, and this study identifies a site on MeCP2, T308, whose phosphorylation is regulated by neuronal activity: phosphorylation of T308 blocks the interaction of MeCP2 with the NCoR co-repressor complex, suppressing MeCP2's ability to repress transcription, and mice carrying mutations of MeCP2 T308 show Rett-syndrome-related symptoms.
- Daniel H. Ebert
- , Harrison W. Gabel
- & Michael E. Greenberg
-
Letter |
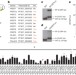 PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
The malaria parasite Plasmodium falciparum escapes immune detection by expressing one of 60 antigenically distinct var genes at any one time during the course of infection: here it is shown that the P. falciparum protein PfSETvs has a key role in var gene silencing through the trimethylation of histone H3K36.
- Lubin Jiang
- , Jianbing Mu
- & Louis H. Miller
-
Letter |
 A stable transcription factor complex nucleated by oligomeric AML1–ETO controls leukaemogenesis
A stable transcription factor complex nucleated by oligomeric AML1–ETO controls leukaemogenesis
A multiprotein complex containing AML1–ETO, the most common fusion protein found in acute myeloid leukaemia, is revealed and analysed in leukaemic cells, and a novel, functionally important protein-binding interface is identified.
- Xiao-Jian Sun
- , Zhanxin Wang
- & Robert G. Roeder
-
Letter |
 Promoter directionality is controlled by U1 snRNP and polyadenylation signals
Promoter directionality is controlled by U1 snRNP and polyadenylation signals
Asymmetric sequence determinants flanking gene transcription start sites are shown to control directionality of transcription elongation in mammalian cells by regulating promoter-proximal cleavage and polyadenylation.
- Albert E. Almada
- , Xuebing Wu
- & Phillip A. Sharp
-
Letter |
 Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription
Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription
It is unclear whether bidirectional non-coding RNAs transcribed from enhancer elements (eRNAs) have any functional role; here, the repressive functions of Rev-Erb nuclear receptors in macrophages are shown to be linked to their ability to inhibit the transcription of eRNAs.
- Michael T. Y. Lam
- , Han Cho
- & Christopher K. Glass
-
Article |
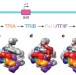 Structural visualization of key steps in human transcription initiation
Structural visualization of key steps in human transcription initiation
Cryo-electron microscopy structures of key intermediates during the sequential assembly of the pre-initiation complex are presented; structures of the closed and open-promoter complexes allow insights into the process of promoter melting.
- Yuan He
- , Jie Fang
- & Eva Nogales
-
Letter |
 The architecture of human general transcription factor TFIID core complex
The architecture of human general transcription factor TFIID core complex
The structures of three distinct human transcription factor IID (TFIID) protein assemblies are solved using cryo-electron microscopy; by incorporating TAF8 and TAF10, the key structural changes that remodel TFIID during assembly are determined, particularly the transition from a symmetric core-TFIID to an asymmetric holo-complex.
- Christoph Bieniossek
- , Gabor Papai
- & Imre Berger
-
Letter |
 Coordinated control of replication and transcription by a SAPK protects genomic integrity
Coordinated control of replication and transcription by a SAPK protects genomic integrity
Upregulation of gene transcription in stressed cells can lead to clashes between the transcription and repair machineries; here, a stress-activated protein kinase (SAPK), Hog1, is shown to coordinate these two processes in yeast.
- Alba Duch
- , Irene Felipe-Abrio
- & Francesc Posas
-
Letter |
 Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Crystal structures of the Pol II–TFIIB complex in free form and bound by the DNA template and a short RNA product are reported; the latter complex represents an initially transcribing complex, a critical transient state in the pathway from transcription initiation to elongation.
- Sarah Sainsbury
- , Jürgen Niesser
- & Patrick Cramer
-
Letter |
 BATF–JUN is critical for IRF4-mediated transcription in T cells
BATF–JUN is critical for IRF4-mediated transcription in T cells
The pleiotropic transcription factor IRF4 is shown to regulate CD4+ T-cell differentiation and TH17 function through cooperative binding interactions with BATF and JUN family proteins via AP1–IRF4 composite elements (AICEs).
- Peng Li
- , Rosanne Spolski
- & Warren J. Leonard
-
Letter |
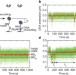 Initiation of transcription-coupled repair characterized at single-molecule resolution
Initiation of transcription-coupled repair characterized at single-molecule resolution
The early stages of transcription-coupled DNA repair are observed at single-molecule resolution; the Escherichia coli DNA translocase molecule Mfd is shown to promote RNA polymerase dissociation by catalysing two irreversible, ATP-dependent transitions.
- Kévin Howan
- , Abigail J. Smith
- & Terence R. Strick
-
Article
| Open Access An expansive human regulatory lexicon encoded in transcription factor footprints
An expansive human regulatory lexicon encoded in transcription factor footprints
DNase I footprinting in 41 cell and tissue types reveals millions of short sequence elements encoding an expansive repertoire of conserved recognition sequences for DNA-binding proteins.
- Shane Neph
- , Jeff Vierstra
- & John A. Stamatoyannopoulos
-
Article
| Open Access Landscape of transcription in human cells
Landscape of transcription in human cells
A description is given of the ENCODE effort to provide a complete catalogue of primary and processed RNAs found either in specific subcellular compartments or throughout the cell, revealing that three-quarters of the human genome can be transcribed, and providing a wealth of information on the range and levels of expression, localization, processing fates and modifications of known and previously unannotated RNAs.
- Sarah Djebali
- , Carrie A. Davis
- & Thomas R. Gingeras
-
Letter |
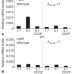 Control of a Salmonella virulence locus by an ATP-sensing leader messenger RNA
Control of a Salmonella virulence locus by an ATP-sensing leader messenger RNA
The Salmonella virulence gene mgtC is regulated by the levels of ATP through an ATP-sensing leader messenger RNA.
- Eun-Jin Lee
- & Eduardo A. Groisman
-
Letter |
 An inverse relationship to germline transcription defines centromeric chromatin in C. elegans
An inverse relationship to germline transcription defines centromeric chromatin in C. elegans
Centromere identity is thought to be epigenetically propagated by stable inheritance of nucleosomes containing the histone variant CENP-A; the authors propose a different model here in which germline transcription defines the genomic regions that exclude CENP-A incorporation during embryogenesis in the holocentric worm Caenorhabditis elegans.
- Reto Gassmann
- , Andreas Rechtsteiner
- & Arshad Desai
-
Article |
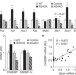 A novel ChREBP isoform in adipose tissue regulates systemic glucose metabolism
A novel ChREBP isoform in adipose tissue regulates systemic glucose metabolism
Downregulation of the glucose transporter GLUT4 in adipose tissue occurs early in the development of type 2 diabetes; here GLUT4-mediated glucose uptake is shown to induce a novel form of the transcription factor ChREBP, which regulates de novo lipogenesis and systemic glucose metabolism.
- Mark A. Herman
- , Odile D. Peroni
- & Barbara B. Kahn
-
News & Views |
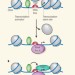 Transcription initiation unwrapped
Transcription initiation unwrapped
A genome-wide, high-resolution study of DNA-binding sites for proteins that transcribe DNA into RNA reveals details about how this process occurs in vivo. See Article p.295
- Stephen Buratowski
-
Letter |
 Circadian rhythms govern cardiac repolarization and arrhythmogenesis
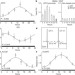
Circadian rhythms govern cardiac repolarization and arrhythmogenesis
Circadian rhythmicity of cardiac ion-channel expression and of an index of myocardial repolarization is under the control of Klf15, a clock-dependent oscillator that is required for generating transient outward potassium current, and deficiencies or excesses of which cause loss of rhythmic variation in myocardial and abnormal repolarization, and an enhanced susceptibility to ventricular arrhythmias.
- Darwin Jeyaraj
- , Saptarsi M. Haldar
- & Mukesh K. Jain
-
Letter |
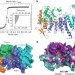 The same pocket in menin binds both MLL and JUND but has opposite effects on transcription
The same pocket in menin binds both MLL and JUND but has opposite effects on transcription
Crystal structures of menin in its free form and in complexes with MLL1 or with JUND, or with an MLL1–LEDGF heterodimer, show that menin contains a deep pocket that binds short peptides of MLL1 or JUND in the same manner, but produces opposite effects on transcription.
- Jing Huang
- , Buddha Gurung
- & Ming Lei
-
Letter |
 Maternal and paternal genomes contribute equally to the transcriptome of early plant embryos
Maternal and paternal genomes contribute equally to the transcriptome of early plant embryos
Transcriptome sequencing and analysis of hybrid embryos show that in contrast to early animal embryogenesis, early plant embryogenesis is mostly under zygotic control.
- Michael D. Nodine
- & David P. Bartel
-
Article |
 Genome-wide structure and organization of eukaryotic pre-initiation complexes
Genome-wide structure and organization of eukaryotic pre-initiation complexes
Ultra-high-resolution mapping of the eukaryotic transcription machinery across the yeast genome reveals several unifying principles of pre-initiation complexes at coding and non-coding genes.
- Ho Sung Rhee
- & B. Franklin Pugh
-
Letter |
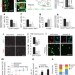 Ars2 maintains neural stem-cell identity through direct transcriptional activation of Sox2
Ars2 maintains neural stem-cell identity through direct transcriptional activation of Sox2
Mammalian zinc finger protein Ars2 is revealed as a sequence-specific transcription factor that promotes the self-renewal of postnatal and adult neural stem cells by directly activating transcription of the pluripotency factor Sox2.
- Celia Andreu-Agullo
- , Thomas Maurin
- & Eric C. Lai
-
Letter |
 Regulatory evolution through divergence of a phosphoswitch in the transcription factor CEBPB
Regulatory evolution through divergence of a phosphoswitch in the transcription factor CEBPB
- Vincent J. Lynch
- , Gemma May
- & Günter P. Wagner
-
Letter |
 Chromatin-associated RNA interference components contribute to transcriptional regulation in Drosophila
Chromatin-associated RNA interference components contribute to transcriptional regulation in Drosophila
- Filippo M. Cernilogar
- , Maria Cristina Onorati
- & Valerio Orlando
-
Letter |
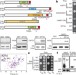 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen
14-3-3 proteins act as intracellular receptors for rice Hd3a florigen
- Ken-ichiro Taoka
- , Izuru Ohki
- & Ko Shimamoto
-
Letter |
 Architecture of the Mediator head module
Architecture of the Mediator head module
- Tsuyoshi Imasaki
- , Guillermo Calero
- & Yuichiro Takagi
-
Letter |
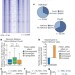 Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA
Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA
- Dong Wang
- , Ivan Garcia-Bassets
- & Xiang-Dong Fu
-
Article |
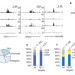 TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity
TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity
- Kristine Williams
- , Jesper Christensen
- & Kristian Helin
-
Letter |
 X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila
X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila
Different organisms use a variety of mechanisms to compensate for X chromosome dosage imbalance between the sexes. In Drosophila, the MSL complex increases transcription on the single X chromosome of males and is thought to regulate transcription elongation, although mechanistic details have been unclear. Here, a global run-on sequencing technique is used to reveal that the MSL complex seems to enhance transcription by facilitating the progression of RNA polymerase II across the bodies of active X linked genes. In this way, MSL can impose dosage compensation on diverse genes with a wide range of transcription levels along the X chromosome.
- Erica Larschan
- , Eric P. Bishop
- & Mitzi I. Kuroda
-
Letter |
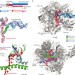 Structural basis of RNA polymerase II backtracking, arrest and reactivation
Structural basis of RNA polymerase II backtracking, arrest and reactivation
During gene transcription, RNA polymerase (Pol) II moves forward along DNA and synthesizes mRNA. However, Pol II can also move backwards and stall, which is important for regulatory purposes or when the polymerase hits an obstacle such as a nucleosome. This arrested state is reactivated by the transcription factor TFIIS. Here, a crystal structure is presented of a backtracked yeast Pol II complex in which the backtracked RNA can be observed, plus a structure of a backtracked complex that contains TFIIS. A model is presented for Pol II backtracking, arrest and reactivation during transcription elongation.
- Alan C. M. Cheung
- & Patrick Cramer
-
News & Views |
 Redox redux
Redox redux
Oscillations in gene transcription that occur in response to biological daily clocks coordinate the physiological workings of living organisms. But turnover in cellular energy may be sufficient to make the clock tick. See Article p.498 & Letter p.554
- Joseph Bass
- & Joseph S. Takahashi
-
Article |
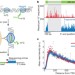 Nascent transcript sequencing visualizes transcription at nucleotide resolution
Nascent transcript sequencing visualizes transcription at nucleotide resolution
A novel technique called native elongating transcript sequencing (NET-seq) is described, which can quantify transcription with single nucleotide resolution. It is based on sequencing nascent transcripts associated with RNA polymerase II that are captured directly from live cells, and is used to gain insights into polymerase pausing and backtracking and the directionality of transcription.
- L. Stirling Churchman
- & Jonathan S. Weissman
-
Letter |
 Transcriptional activation of polycomb-repressed genes by ZRF1
Transcriptional activation of polycomb-repressed genes by ZRF1
Ubiquitination of histone H2A has been implicated in polycomb-mediated transcriptional silencing, but its precise functions are unclear. Here, ZRF1 is shown to be recruited to ubiquitinated H2A and to function in displacing polycomb-repressive complex 1 (PRC1) from chromatin to facilitate transcriptional activation.
- Holger Richly
- , Luciana Rocha-Viegas
- & Luciano Di Croce
-
News & Views |
 The coherent Mediator
The coherent Mediator
Enhancer sequences increase gene transcription with the help of a co-activator complex, the Mediator. Another protein complex — cohesin — seems to work with Mediator to bring together enhancers and promoters. See Article p. 430
- Rolf Ohlsson
-
Letter |
 New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism
New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism
In the course of characterizing short RNAs from human cells using single-molecule high-throughput sequencing, these authors identify a new short RNA species. The presence of non-genomically encoded poly(U) residues at their 5' ends implies the existence of an unknown RNA copying mechanism in human cells.
- Philipp Kapranov
- , Fatih Ozsolak
- & Patrice M. Milos
-
Letter |
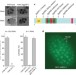 Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
Small regulatory RNAs function both in the cytoplasm, inhibiting expression from messenger RNAs, and in the nucleus, silencing heterochromatin and preventing genome rearrangement. Now a new protein involved in RNA interference in the nucleus has been characterized. This protein, NRDE-2, associates with NRDE-3 and short interfering RNAs on nascent transcripts. This association prevents elongation of the transcripts by RNA polymerase II, making this a co-transcriptional form of gene silencing.
- Shouhong Guang
- , Aaron F. Bochner
- & Scott Kennedy
-
Letter |
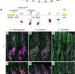 Phenotypic robustness conferred by apparently redundant transcriptional enhancers
Phenotypic robustness conferred by apparently redundant transcriptional enhancers
Transcriptional enhancers are segments of regulatory DNA located some distance from the coding region of a gene, and several of them may sometimes serve apparently redundant functions. These authors demonstrate in Drosophila that such 'redundant' enhancers, by contributing higher overall levels of transcription, ensure robustness of phenotypes against both genetic and environmental perturbations, for example mutations in other genes or temperature changes that would otherwise lead to aberrant development.
- Nicolás Frankel
- , Gregory K. Davis
- & David L. Stern
-
Research Highlights |
Genomics: Not-so-dark genome
-
News |
 Existence of RNA 'dark matter' in doubt
Existence of RNA 'dark matter' in doubt
The abundance of transcripts from the genome may have been overestimated.
- Melissa Lee Phillips
-
News & Views |
 Enhancers make non-coding RNA
Enhancers make non-coding RNA
Genomes don't just encode protein-coding RNAs. They also give rise to various groups of RNAs that can regulate gene expression. Short RNAs that form from enhancer sequences might be one such class of regulatory RNA.
- Bing Ren
-
Article |
 Widespread transcription at neuronal activity-regulated enhancers
Widespread transcription at neuronal activity-regulated enhancers
Regulatory proteins bind non-coding DNA either at promoters (near to a gene's transcription start site) or at enhancers (far away). Binding at enhancers helps to bring the transcription enzyme RNA polymerase to promoters. Here, studies of some 12,000 enhancers that respond to electrical activity in neurons show that binding to enhancers also brings the polymerase to the enhancers themselves, where it transcribes a novel class of non-coding RNAs. Enhancers may thus be more similar to promoters than hitherto appreciated.
- Tae-Kyung Kim
- , Martin Hemberg
- & Michael E. Greenberg
-
Letter |
 Transcription-independent ARF regulation in oncogenic stress-mediated p53 responses
Transcription-independent ARF regulation in oncogenic stress-mediated p53 responses
In response to oncogenic stress, the tumour suppressor ARF activates the p53 protein. ARF protein is highly stable in most human cell lines, so it has been thought that ARF activation occurs mainly at the level of transcription. Here, however, ARF is shown to be unstable in normal human cells but stable in cancer cells, through a transcription-independent mechanism. A ubiquitin ligase for ARF is identified and shown to promote ARF degradation in normal cells. This activity is prevented in cancer cells, stabilizing ARF.
- Delin Chen
- , Jing Shan
- & Wei Gu
-
Letter |
 Transcriptional control of preadipocyte determination by Zfp423
Transcriptional control of preadipocyte determination by Zfp423
An understanding of how fat cells (adipocytes) develop will contribute to our understanding of obesity. The differentiation of committed preadipocytes into adipocytes is known to be controlled by PPARγ and several other transcription factors. But what turns a cell into a preadipocyte? Here, the zinc-finger protein Zfp423 is identified as a transcriptional regulator of preadipocyte determination.
- Rana K. Gupta
- , Zoltan Arany
- & Bruce M. Spiegelman
-
Letter |
 Transcriptional role of cyclin D1 in development revealed by a genetic–proteomic screen
Transcriptional role of cyclin D1 in development revealed by a genetic–proteomic screen
Although cyclin D1 is frequently overexpressed in human cancers, the full range of its functions in normal development and oncogenesis is unclear. Here, tagged cyclin D1 knock-in mouse strains are developed to allow a search for cyclin D1-binding proteins in different mouse organs using high-throughput mass spectrometry. The results show that, in addition to its established cell cycle roles, cyclin D1 has an in vivo transcriptional function in mouse development.
- Frédéric Bienvenu
- , Siwanon Jirawatnotai
- & Piotr Sicinski
-
Letter |
 An allosteric mechanism of Rho-dependent transcription termination
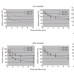
An allosteric mechanism of Rho-dependent transcription termination
Rho is a general transcription termination factor in bacteria, but the mechanism by which it disrupts the RNA polymerase (RNAP) elongation complex is unknown. Here, Rho is shown to bind tightly to the RNAP throughout the transcription cycle, with the formation of the RNAP–Rho complex being crucial for termination. Furthermore, RNAP is proposed to have an active role in Rho termination through an allosteric mechanism.
- Vitaly Epshtein
- , Dipak Dutta
- & Evgeny Nudler

 The nuclear receptor Rev-erbα controls circadian thermogenic plasticity
The nuclear receptor Rev-erbα controls circadian thermogenic plasticity