Featured
-
-
Article
| Open Access Alignment of single-cell trajectory trees with CAPITAL
Alignment of single-cell trajectory trees with CAPITAL
Global alignment of complex cell state trajectories between single-cell datasets remains challenging. Here, the authors present a computational method called CAPITAL to compare branching trajectories, and demonstrate that this method achieves accurate and robust alignments.
- Reiichi Sugihara
- , Yuki Kato
- & Yukio Kawahara
-
Article
| Open Access GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
Plant genotype alone appears to be insufficient to explain trait variations. This study integrates GWAS, MWAS and mGWAS in 827 foxtail millet cultivars, revealing that root-associated microbiota affect plant phenotypes in a host genotype-dependent manner.
- Yayu Wang
- , Xiaolin Wang
- & Huan Liu
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Single cell and spatial transcriptomic analyses reveal microglia-plasma cell crosstalk in the brain during Trypanosoma brucei infection
Single cell and spatial transcriptomic analyses reveal microglia-plasma cell crosstalk in the brain during Trypanosoma brucei infection
Detailed insight into how the brain responds to Trypanosoma brucei infection is lacking. Here, single cell and spatial transcriptomics are integrated to characterise this response, identifying a unique crosstalk between microglia and plasma cells.
- Juan F. Quintana
- , Praveena Chandrasegaran
- & Annette MacLeod
-
Article
| Open Access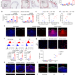 Single-cell transcriptomics reveal cellular diversity of aortic valve and the immunomodulation by PPARγ during hyperlipidemia
Single-cell transcriptomics reveal cellular diversity of aortic valve and the immunomodulation by PPARγ during hyperlipidemia
Identifying the mechanisms underlying the early inflammatory phase of aortic valve disease is crucial for disease prevention. Here the authors perform single-cell RNA sequencing to show the immunomodulatory role of PPARγ in valvular endothelial cells during hyperlipidemia.
- Seung Hyun Lee
- , Nayoung Kim
- & Jae-Hoon Choi
-
Article
| Open Access Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Diabetic kidney disease leads to changes in glucose metabolism and inflammation. Here the authors use multimodal single cell sequencing to show that this disease leads to reduced accessibility of glucocorticoid receptor binding sites in the proximal tubule and increased gluconeogenesis.
- Parker C. Wilson
- , Yoshiharu Muto
- & Benjamin D. Humphreys
-
Article
| Open Access PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
Protein kinase-mediated phosphorylation plays a critical role in many biological processes. Here the authors develop a trans-omics-based algorithm called Central Kinase Inference to integrate quantitative transcriptomic and phosphoproteomic data, finding that PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest.
- Yangyang Yuan
- , Chenwei Wang
- & Pengyu Huang
-
Article
| Open Access Heterogeneity and transcriptome changes of human CD8+ T cells across nine decades of life
Heterogeneity and transcriptome changes of human CD8+ T cells across nine decades of life
The characterisation of T cells during aging is important to predict functional outcomes in vaccination or infection. Here the authors use flow cytometry and scRNA sequencing to transcriptionally age CD8 T cells and then use a machine learning model to interpret cell age from transcriptional profiles.
- Jian Lu
- , Raheel Ahmad
- & Nan-ping Weng
-
Article
| Open Access Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Development of methods for simultaneous single cell analysis of transcription and translation is still underway. Here, Hu et al. develop single-cell transcriptome and translatome dual-omics on human oocytes, which enables them to identify OOSP2 as an induction factor during human oocyte maturation.
- Wenqi Hu
- , Haitao Zeng
- & Kehkooi Kee
-
Article
| Open Access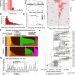 Deconvoluting complex correlates of COVID-19 severity with a multi-omic pandemic tracking strategy
Deconvoluting complex correlates of COVID-19 severity with a multi-omic pandemic tracking strategy
There is a genetic component to the risk of severe COVID-19, but the genetic effects are difficult to separate from social constructs that covary with genetic ancestry. To address this, the authors identify determinants of COVID-19 severity using admixture mapping, viral phylodynamics, and host immune and metagenomic sequencing.
- Victoria N. Parikh
- , Alexander G. Ioannidis
- & Euan A. Ashley
-
Article
| Open Access Integrated multi-omics reveals cellular and molecular interactions governing the invasive niche of basal cell carcinoma
Integrated multi-omics reveals cellular and molecular interactions governing the invasive niche of basal cell carcinoma
The role of reciprocal tumour-stroma interactions in tumour invasion remains poorly characterised. Here, single-cell and spatial transcriptomics identifies the cell populations and their transcriptional reprogramming contributing to the spatial organization of the basal cell carcinoma invasive niche.
- Laura Yerly
- , Christine Pich-Bavastro
- & François Kuonen
-
Article
| Open Access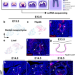 Spatiotemporal single-cell regulatory atlas reveals neural crest lineage diversification and cellular function during tooth morphogenesis
Spatiotemporal single-cell regulatory atlas reveals neural crest lineage diversification and cellular function during tooth morphogenesis
The mechanisms that govern cell fate decisions of postmigratory cranial neural crest cells remain largely unknown. Here the authors present a spatiotemporal single-cell regulatory atlas tracking these cells’ dental lineage diversification.
- Junjun Jing
- , Jifan Feng
- & Yang Chai
-
Article
| Open Access Clonal diversification and histogenesis of malignant germ cell tumours
Clonal diversification and histogenesis of malignant germ cell tumours
The molecular characterisation of germ cell tumours (GCT) is necessary to understand their development and histological diversification. Here, the authors use whole-genome and transcriptome sequencing of GCTs across distinct histologies to reveal their somatic evolution and clonal diversification, as well as identify several putative biomarkers for treatment stratification.
- Thomas R. W. Oliver
- , Lia Chappell
- & Sam Behjati
-
Article
| Open Access Cell landscape of larval and adult Xenopus laevis at single-cell resolution
Cell landscape of larval and adult Xenopus laevis at single-cell resolution
Single-cell RNA sequencing technology offers a unique opportunity to dissect cell heterogeneity of animals. Here, the authors construct a Xenopus cell landscape including larval and adult organs to dissect cell heterogeneity of the amphibian.
- Yuan Liao
- , Lifeng Ma
- & Xiaoping Han
-
Article
| Open Access Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
The Mexican axolotl is a well-established tetrapod model for regeneration and development. Here the authors report a scRNA-seq method to profile neotenic, metamorphic and limb development stages, highlighting unique perturbation patterns of cell type-related gene expression throughout metamorphosis.
- Fang Ye
- , Guodong Zhang
- & Guoji Guo
-
Article
| Open Access Cutaneous and acral melanoma cross-OMICs reveals prognostic cancer drivers associated with pathobiology and ultraviolet exposure
Cutaneous and acral melanoma cross-OMICs reveals prognostic cancer drivers associated with pathobiology and ultraviolet exposure
While cutaneous melanoma is linked to UV radiation, acral melanoma is not. Epigenetic mechanisms function as sensors to exposures and determinants of cell identity. Here, the authors use DNA methylation data to identify dysregulated pathways associated with UV radiation and pathobiology in cutaneous and acral melanomas.
- Anna Luiza Silva Almeida Vicente
- , Alexei Novoloaca
- & Akram Ghantous
-
Article
| Open Access Identifying multicellular spatiotemporal organization of cells with SpaceFlow
Identifying multicellular spatiotemporal organization of cells with SpaceFlow
A critical task in spatial transcriptomics analysis is to understand inherently spatial relationships among cells. Here, the authors present a deep learning framework to integrate spatial and transcriptional information, spatially extending pseudotime and revealing spatiotemporal organization of cells.
- Honglei Ren
- , Benjamin L. Walker
- & Qing Nie
-
Article
| Open Access A reference single-cell regulomic and transcriptomic map of cynomolgus monkeys
A reference single-cell regulomic and transcriptomic map of cynomolgus monkeys
Non-human primates are attractive laboratory animal models that can accurately reflect some developmental and pathological features of humans. Here the authors chart a reference cell map of cynomolgus monkeys using both scATAC-seq and scRNA-seq data across multiple organs, providing insights into the molecular dynamics and cellular heterogeneity of this organism.
- Jiao Qu
- , Fa Yang
- & Dijun Chen
-
Article
| Open Access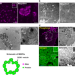 DDX1 vesicles control calcium-dependent mitochondrial activity in mouse embryos
DDX1 vesicles control calcium-dependent mitochondrial activity in mouse embryos
The DEAD box protein DDX1 is known to form large aggregates in the cytoplasm of early mouse embryos. Here the authors identify DDX1-containing vesicles and show that loss of Ddx1 affects their integrity, compromising mitochondria function and causing embryonic lethality.
- Yixiong Wang
- , Lubna Yasmin
- & Roseline Godbout
-
Article
| Open Access Association of mitochondrial DNA content, heteroplasmies and inter-generational transmission with autism
Association of mitochondrial DNA content, heteroplasmies and inter-generational transmission with autism
Most genetic studies of autism spectrum disorder (ASD) have focused on the nuclear genome. Here, the authors show that variations in mitochondrial DNA, detectable at birth, are also associated with risk of ASD.
- Yiqin Wang
- , Xiaoxian Guo
- & Zhenglong Gu
-
Article
| Open Access Context-aware deconvolution of cell–cell communication with Tensor-cell2cell
Context-aware deconvolution of cell–cell communication with Tensor-cell2cell
Cellular contexts such as disease state, organismal life stage and tissue microenvironment, shape intercellular communication, and ultimately affect an organism’s phenotypes. Here, the authors present Tensor-cell2cell, an unsupervised method for deciphering context-driven intercellular communication.
- Erick Armingol
- , Hratch M. Baghdassarian
- & Nathan E. Lewis
-
Article
| Open Access Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level
Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level
Pigs are important large animal models for biomedical research. Here, the authors construct a single-cell landscape of pig tissues, unravelling the phenotypic heterogeneity of blood endothelial cells in adipose tissues and the evolutionally conserved regulons of microglia in brains.
- Fei Wang
- , Peiwen Ding
- & Yonglun Luo
-
Article
| Open Access A hexa-species transcriptome atlas of mammalian embryogenesis delineates metabolic regulation across three different implantation modes
A hexa-species transcriptome atlas of mammalian embryogenesis delineates metabolic regulation across three different implantation modes
Mammalian embryogenesis relies on glycolysis and oxidative phosphorylation, but understanding of the dynamics of metabolic regulation in the postimplantation embryo in vivo remains elusive. Here the authors compile single-cell embryo profiling data in six mammalian species and reveal a conserved metabolic programme despite different implantation modes.
- Anna Malkowska
- , Christopher Penfold
- & Thorsten E. Boroviak
-
Article
| Open Access Phasing analysis of lung cancer genomes using a long read sequencer
Phasing analysis of lung cancer genomes using a long read sequencer
Long-read sequencing technologies are useful for the multifaceted task of characterising somatic mutations, including structural variants, in cancers. Here, the authors combine short and long read sequencing for the phasing analysis, which enables them to resolve the chromosomal backgrounds of somatic mutations in Japanese non-small cell lung cancers.
- Yoshitaka Sakamoto
- , Shuhei Miyake
- & Ayako Suzuki
-
Article
| Open Access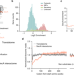 Ribosome profiling reveals multiple roles of SecA in cotranslational protein export
Ribosome profiling reveals multiple roles of SecA in cotranslational protein export
Using a combination of ribosome profiling methods, Zhu et al. investigate the principles governing the cotranslational interaction of SecA with nascent proteins and reveal a hierarchical organization of protein export pathways in bacteria.
- Zikun Zhu
- , Shuai Wang
- & Shu-ou Shan
-
Article
| Open Access Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing
Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing
Studies of circular RNAs have often been limited to the tissue or organism level. Here, authors investigate the comprehensive expression landscape of circRNAs in human and mouse at single-cell resolution, revealing highly specific and dynamic changes of circRNAs during multiple biological processes.
- Wanying Wu
- , Jinyang Zhang
- & Fangqing Zhao
-
Article
| Open Access A single-cell transcriptomic atlas characterizes the silk-producing organ in the silkworm
A single-cell transcriptomic atlas characterizes the silk-producing organ in the silkworm
The molecular underpinning of silk-producing organs is not well characterized. Here the authors use single-cell RNA sequencing to build an atlas of the silkworm silk gland and reveal the heterogeneity of silk gland cells.
- Yan Ma
- , Wenhui Zeng
- & Hanfu Xu
-
Article
| Open Access Salivary gland organoid culture maintains distinct glandular properties of murine and human major salivary glands
Salivary gland organoid culture maintains distinct glandular properties of murine and human major salivary glands
The long-term maintenance of diverse salivary gland cells remains challenging. Here the authors establish a protocol for long-term maintenance of murine and human salivary gland organoids exhibiting gland-specific gene expression, gland functions, and cellular diversity confirmed by scRNA-seq.
- Yeo-Jun Yoon
- , Donghyun Kim
- & Jae-Yol Lim
-
Article
| Open Access Mapping the cardiac vascular niche in heart failure
Mapping the cardiac vascular niche in heart failure
The cardiac vascular niche is of major importance in homeostasis and disease, but knowledge of its complexity in response to injury remains limited. Here we combine lineage tracing with single cell RNA sequencing to show alterations in fibroblasts, endothelial and mural cells in hypertrophic remodeling.
- Fabian Peisker
- , Maurice Halder
- & Rafael Kramann
-
Article
| Open Access Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project
Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project
Biobanks of genetic data have been primarily in European populations, which gives us an incomplete understanding of complex traits across populations. Here, the authors initiate the Westlake BioBank for Chinese (WBBC) pilot project with 4,535 whole genome sequences and 5,841 high-density genotypes from China, characterizing large-scale genomic variation in Chinese populations.
- Pei-Kuan Cong
- , Wei-Yang Bai
- & Hou-Feng Zheng
-
Article
| Open Access Peptide nano-blanket impedes fibroblasts activation and subsequent formation of pre-metastatic niche
Peptide nano-blanket impedes fibroblasts activation and subsequent formation of pre-metastatic niche
Primary tumors “spread the spark” by establishing a pre-metastatic niche. Here the authors develop an in-situ assembled peptide FR17 to serve as a “flame-retarding blanket” to extinguish the “fire” of the pre-metastatic microenvironment.
- Yi Zhou
- , Peng Ke
- & Jianqing Gao
-
Article
| Open Access Automated next-generation profiling of genomic alterations in human cancers
Automated next-generation profiling of genomic alterations in human cancers
The genomic profiling of tumours has not been widely adopted in the clinic due to technical and practical hurdles. Here, the authors develop PGDx elio tissue complete, a scalable, standardised and FDA-cleared test comprising a targeted gene panel and automated machine-learning analysis, which detects clinically relevant sequence biomarkers in cancer samples.
- Laurel A. Keefer
- , James R. White
- & Mark Sausen
-
Article
| Open Access Nuclear oligo hashing improves differential analysis of single-cell RNA-seq
Nuclear oligo hashing improves differential analysis of single-cell RNA-seq
Using spike-in controls with current single cell RNA-seq platforms remains a challenge. Here, the authors use a mixture of short, unmodified DNA oligos as a normalization standard for sci-RNAseq to improve the detection of global transcriptome changes.
- Hyeon-Jin Kim
- , Greg Booth
- & Cole Trapnell
-
Article
| Open Access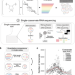 Characterization of RNA content in individual phase-separated coacervate microdroplets
Characterization of RNA content in individual phase-separated coacervate microdroplets
Here, the authors demonstrate that single cell RNA sequencing technology can be leveraged to characterize RNA content of individual membrane-free condensates formed by liquid-liquid phase separation processes such as coacervation.
- Damian Wollny
- , Benjamin Vernot
- & Barbara Treutlein
-
Article
| Open Access A rare variant analysis framework using public genotype summary counts to prioritize disease-predisposition genes
A rare variant analysis framework using public genotype summary counts to prioritize disease-predisposition genes
Sequencing studies in clinical and cancer genomics often utilize public data sets to identify genes enriched with pathogenic variants. Here, the authors propose a framework which controls for confounding factors that can bias the results in these studies.
- Wenan Chen
- , Shuoguo Wang
- & Gang Wu
-
Article
| Open Access Contribution of rare whole-genome sequencing variants to plasma protein levels and the missing heritability
Contribution of rare whole-genome sequencing variants to plasma protein levels and the missing heritability
Despite the success of genome-wide association studies, much of the genetic contribution to complex traits remains unexplained. Here, the authors identify effects by rare variants on plasma proteins, and estimate the contribution of rare variants to the heritability.
- Marcin Kierczak
- , Nima Rafati
- & Åsa Johansson
-
Article
| Open Access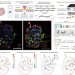 Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
The spatial context of epitranscriptomic features in the tumour microenvironment remains poorly understood. Here, a method for transcriptomic and epitranscriptomic analysis of immunofluorescence-stained tissue, Select-seq, is applied to stem cell-like microniches in triple negative breast cancer.
- Amos C. Lee
- , Yongju Lee
- & Sunghoon Kwon
-
Article
| Open Access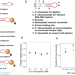 A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
This paper develops a multiplex RNA-seq method that reports tRNA abundance, modification, charging, and fragmentation. Results show stress-induced regulation in translational elongation and association of modification and tRNA fragment biogenesis.
- Christopher P. Watkins
- , Wen Zhang
- & Tao Pan
-
Article
| Open Access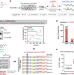 TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
RNA modifications are important regulators of RNA biology. Here we report N1-methyladenosine (m1A) enrichment on 22-nucleotide tRNA fragments and its effect on gene-silencing. Higher level of m1A in bladder cancer is accompanied by gene dysregulation in unfolded protein response.
- Zhangli Su
- , Ida Monshaugen
- & Anindya Dutta
-
Article
| Open Access Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium
Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium
The level of cellular diversity in the esophageal epithelium has yet to be classified at the single cell level. Here the authors analyze the transcriptome of 44,679 murine esophageal keratinocytes to identify an unexpected level of cellular heterogeneity.
- Mohammad Faujul Kabir
- , Adam L. Karami
- & Kelly A. Whelan
-
Article
| Open Access Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
T cell large granular lymphocyte leukemia (T-LGLL) and the cellular phenotype underlying response to therapy is not well understood. Here the authors use single cell sequencing to better understand changes in T cell clonal frequency and gene expression before and after therapy in T-LGLL.
- Shouguo Gao
- , Zhijie Wu
- & Neal S. Young
-
Article
| Open Access Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE)
Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE)
The design of highly multiplex PCR primers to amplify and enrich many different DNA sequences is increasing in biomedical importance as new mutations and pathogens are identified. The authors present and experimentally validate Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE), a stochastic algorithm for design of highly multiplex PCR primer sets that minimize primer dimer formation.
- Nina G. Xie
- , Michael X. Wang
- & David Yu Zhang
-
Article
| Open Access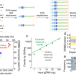 Ensemble of nucleic acid absolute quantitation modules for copy number variation detection and RNA profiling
Ensemble of nucleic acid absolute quantitation modules for copy number variation detection and RNA profiling
Highly multiplexed ddPCR for sensitive nucleic acid quantitation remains challenging due to limited fluorescence channels. Here the authors report quantitative amplicon sequencing (QASeq), a PCR-based molecular barcoding NGS approach which is compatible with high multiplexing.
- Lucia Ruojia Wu
- , Peng Dai
- & David Yu Zhang
-
Article
| Open Access Tissue extracellular matrix hydrogels as alternatives to Matrigel for culturing gastrointestinal organoids
Tissue extracellular matrix hydrogels as alternatives to Matrigel for culturing gastrointestinal organoids
The culture of gastrointestinal organoids relies on Matrigel that has several drawbacks for clinical application. Here, the authors report the feasibility of gastrointestinal tissue-mimetic matrices as effective alternatives to Matrigel for organoid culture and transplantation.
- Suran Kim
- , Sungjin Min
- & Seung-Woo Cho
-
Article
| Open Access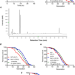 The flavonoid corylin exhibits lifespan extension properties in mouse
The flavonoid corylin exhibits lifespan extension properties in mouse
Various traditional Chinese herbs and complex formulas have been suggested to exhibit lifespan extension properties. Here, the authors isolated the flavonoid corylin from Psoralea corylifolia and demonstrated its longevity properties in yeast, human cells and mouse.
- Tong-Hong Wang
- , Wei-Che Tseng
- & Chin-Chuan Chen
-
Article
| Open Access Co-translational assembly orchestrates competing biogenesis pathways
Co-translational assembly orchestrates competing biogenesis pathways
The biogenesis of nuclear pores imposes a logistic challenge for cells. Here, the authors investigate structural motifs for co-translational interactions in nucleoporins and find that co-translational assembly events differ between paralogous assembly pathways thus contributing to faithful assembly.
- Maximilian Seidel
- , Anja Becker
- & Martin Beck
-
Article
| Open Access Antibody escape and global spread of SARS-CoV-2 lineage A.27
Antibody escape and global spread of SARS-CoV-2 lineage A.27
The A.27 SARS-CoV-2 lineage spread globally in 2021 but did not become dominant. Here, the authors show that A.27 shares some mutations in the spike gene that are present in variants of concern, but lacks the D614G mutation, indicating independent evolution of immune escape properties.
- Tamara Kaleta
- , Lisa Kern
- & Jonas Fuchs
-
Article
| Open Access High-throughput identification and quantification of single bacterial cells in the microbiota
High-throughput identification and quantification of single bacterial cells in the microbiota
Here, Jin et al., develop a method called Barcoding Bacteria for Identification and Quantification (BarBIQ), which allows to both characterize the global microbiome and to identify and quantify single-cell bacterial members in a high-throughput manner.
- Jianshi Jin
- , Reiko Yamamoto
- & Katsuyuki Shiroguchi
-
Article
| Open Access Whole exome sequencing in Alopecia Areata identifies rare variants in KRT82
Whole exome sequencing in Alopecia Areata identifies rare variants in KRT82
Common variants have been discovered to be associated with Alopecia Areata; however, rare variants have been less well studied. Here, the authors use whole-exome sequencing to identify associated rare variants in the hair keratin gene KRT82. Further, they find that individuals with Alopecia Areata have reduced expression of KRT82 in the skin and hair follicle.
- Stephanie O. Erjavec
- , Sahar Gelfman
- & Angela M. Christiano

 Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space
Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space