Featured
-
-
Article
| Open Access A universal molecular control for DNA, mRNA and protein expression
A universal molecular control for DNA, mRNA and protein expression
Multi-omics analyses powerfully combine gene expression and translation, however no available controls can be used across these techniques. Here the authors develop pREF, a universal control construct designed for use in DNA, RNA and protein analyses.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
The impact of other pathogens on disease outcome was studied in European infants with RSV infection. Additional viruses were commonly co-detected during infection but were weakly linked to severity. However, presence of Haemophilus bacteria strongly associated with severe cases.
- Gu-Lung Lin
- , Simon B. Drysdale
- & Andrew J. Pollard
-
Article
| Open Access ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
Identifying small proteins is challenging. ProTInSeq uses modified transposons to express markers inserted in-frame to protein-coding genes. This method identifies 153 unannotated small proteins in M. pneumoniae and additional proteomic information.
- Samuel Miravet-Verde
- , Rocco Mazzolini
- & Luis Serrano
-
Article
| Open Access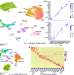 Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
A profibrotic, proinflammatory kidney cell population has been identified as a driver of chronic kidney disease. Here, authors generate a human kidney single cell multiomic dataset and apply a regularised regression approach to identify transcription factors underpinning this cell population.
- Nicolas Ledru
- , Parker C. Wilson
- & Benjamin D. Humphreys
-
Article
| Open Access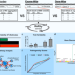 A method to estimate the contribution of rare coding variants to complex trait heritability
A method to estimate the contribution of rare coding variants to complex trait heritability
The contribution of rare variants to complex traits has not been well studied. Here, the authors present RARity, a method to assess rare variant heritability without assuming a particular genetic architecture and enabling both gene-level and exome-wide heritability estimation of continuous traits.
- Nazia Pathan
- , Wei Q. Deng
- & Guillaume Paré
-
Article
| Open Access Human whole-exome genotype data for Alzheimer’s disease
Human whole-exome genotype data for Alzheimer’s disease
The heterogeneity of whole-exome sequencing (WES) data generation methods presents a challenge to joint analysis. Here, the authors present a bioinformatics strategy to generate high-quality data from processing diversely generated WES samples, as applied in the Alzheimer’s Disease Sequencing Project.
- Yuk Yee Leung
- , Adam C. Naj
- & Li-San Wang
-
Article
| Open Access Terminal modifications independent cell-free RNA sequencing enables sensitive early cancer detection and classification
Terminal modifications independent cell-free RNA sequencing enables sensitive early cancer detection and classification
Cell free RNA is a potentially valuable resource to detect cancer, however, its low concentration in plasma can limit usefulness. Here, the authors devise a library preparation method from 100ul of plasma, and apply to multiple cancer types to detect and classify cancer patients
- Jun Wang
- , Jinyong Huang
- & Deming Gou
-
Article
| Open Access ACIDES: on-line monitoring of forward genetic screens for protein engineering
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Screening mutated proteins is a versatile strategy in protein research, producing massive datasets when combined with NGS. Here, authors present ACIDES to estimate mutated protein fitness and aid protein engineering pipelines in a range of applications, including gene therapy.
- Takahiro Nemoto
- , Tommaso Ocari
- & Ulisse Ferrari
-
Article
| Open Access Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Current sequencing technologies can shed light on the stepwise progression of lung adenocarcinoma. Here, the authors characterize tumor progression in lung adenocarcinomas from an early stage using short and long read whole-genome sequencing, bulk and spatial transcriptomics, and epigenomics.
- Yasuhiko Haga
- , Yoshitaka Sakamoto
- & Ayako Suzuki
-
Article
| Open Access Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
The role of clonal evolution on the actionable proteome and response to therapy in childhood acute lymphoblastic leukemia (ALL) remains unknown. Here, targeted sequencing and proteomic analysis of paired ALL diagnosis and relapsed samples revealed PARP1 as a potential therapeutic target.
- Amanda C. Lorentzian
- , Jenna Rever
- & Philipp F. Lange
-
Article
| Open Access MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Methylation is the dominant modification in mRNA and occurs at a variety of sites. Here, Hartstock et al. show that a clickable analogue of the key cosubstrate S-adenosyl-L-methionine (SAM) can be produced in cells, allowing for identification and mapping of different methylated nucleosides in mRNA.
- Katja Hartstock
- , Nadine A. Kueck
- & Andrea Rentmeister
-
Article
| Open Access Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Chameleolyser enables the accurate identification of genetic variants hidden within complex regions of the genome. Its application uncovers the disease-explanatory variant in 25 previously undiagnosed patients.
- Wouter Steyaert
- , Lonneke Haer-Wigman
- & Christian Gilissen
-
Article
| Open Access Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Unnatural base pairing xenonucleic acids (XNAs) can be used to expand life’s alphabet beyond ATGC. Here, authors show strategies for enzymatic synthesis and next-generation nanopore sequencing of XNA base pairs for reading and writing 12-letter DNA (ATGCBSPZXKJV).
- Hinako Kawabe
- , Christopher A. Thomas
- & Jorge A. Marchand
-
Article
| Open Access A landscape of complex tandem repeats within individual human genomes
A landscape of complex tandem repeats within individual human genomes
Haplotype-resolved long, complex tandem repeats remain largely hidden despite their potential relevance to disease. Here, the authors reveal and analyze the genome-wide landscape of these repeats using a high-precision algorithm.
- Kazuki Ichikawa
- , Riki Kawahara
- & Shinichi Morishita
-
Article
| Open Access Metagenomic sequencing of post-mortem tissue samples for the identification of pathogens associated with neonatal deaths
Metagenomic sequencing of post-mortem tissue samples for the identification of pathogens associated with neonatal deaths
Rapid identification of pathogens in neonatal infection, and corresponding antimicrobial susceptibility profiles, would improve patient outcomes and assist in antibiotic stewardship. In this work, the authors utilize metagenomic next-generation sequencing of post-mortem tissue samples to identify pathogens associated with neonatal deaths.
- Vicky L. Baillie
- , Shabir A. Madhi
- & Courtney P. Olwagen
-
Article
| Open Access Spatial transcriptomics reveal markers of histopathological changes in Duchenne muscular dystrophy mouse models
Spatial transcriptomics reveal markers of histopathological changes in Duchenne muscular dystrophy mouse models
Duchenne muscular dystrophy is a fatal neuromuscular disorder affecting one in 5000 male births. To enrich our understanding of the underlying pathology, the authors apply spatial transcriptomics on dystrophic skeletal muscle to unravel markers related to histopathological changes in Duchenne mouse models.
- L.G.M. Heezen
- , T. Abdelaal
- & P. Spitali
-
Article
| Open Access Three-dimensional molecular architecture of mouse organogenesis
Three-dimensional molecular architecture of mouse organogenesis
Qu et al. present a detailed three-dimensional spatial transcriptome atlas of all major organs in the mouse embryo at E13.5, providing a better understanding of organ development and cellular interactions during mammalian development.
- Fangfang Qu
- , Wenjia Li
- & Guangdun Peng
-
Article
| Open Access Increased interregional virus exchange and nucleotide diversity outline the expansion of chikungunya virus in Brazil
Increased interregional virus exchange and nucleotide diversity outline the expansion of chikungunya virus in Brazil
Chikungunya virus is endemic in Brazil and cases have been rapidly increasing in recent years. Here, the authors describe the expansion of a genomic surveillance program across the country allowing them to characterise the emergence and dispersal of two distinct subclades mainly seeded from the north eastern region.
- Joilson Xavier
- , Luiz Carlos Junior Alcantara
- & Marta Giovanetti
-
Article
| Open Access Multiplexed ddPCR-amplicon sequencing reveals isolated Plasmodium falciparum populations amenable to local elimination in Zanzibar, Tanzania
Multiplexed ddPCR-amplicon sequencing reveals isolated Plasmodium falciparum populations amenable to local elimination in Zanzibar, Tanzania
Sequencing malaria parasites from low density infections in small amounts of dried blood is important for large-scale genomic surveillance. Here, the authors develop and validate a highly multiplexed droplet digital PCR-based amplicon deep sequencing assay and apply it to data from Zanzibar, Tanzania.
- Aurel Holzschuh
- , Anita Lerch
- & Cristian Koepfli
-
Comment
| Open Access Unravelling the genetic architecture of human complex traits through whole genome sequencing
Unravelling the genetic architecture of human complex traits through whole genome sequencing
Whole genome sequencing has enabled new insights into the genetic architecture of complex traits, especially through access to low-frequency and rare variation. This Comment highlights the key contributions from this technology and discusses considerations for its use and future perspectives.
- Ozvan Bocher
- , Cristen J. Willer
- & Eleftheria Zeggini
-
Article
| Open Access A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
Authors carry out a longitudinal genomic analysis of Salmonella enterica serovar Concord isolates from various geographical locations, to reconstruct population diversity, evolution and antimicrobial resistance distribution.
- Wim L. Cuypers
- , Pieter Meysman
- & Sandra Van Puyvelde
-
Article
| Open Access High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
Long-read single-cell RNA isoform sequencing can elucidate the intricate landscape of alternative RNA splicing in individual cells, but it suffers from a low read throughput. Here, the authors develop circular consensus sequencing methods to allow high-throughput and high-accuracy single-cell RNA isoform sequencing.
- Zhuo-Xing Shi
- , Zhi-Chao Chen
- & Yi-Zhi Liu
-
Article
| Open Access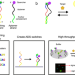 A massively parallel screening platform for converting aptamers into molecular switches
A massively parallel screening platform for converting aptamers into molecular switches
Efforts to convert aptamers into molecular switches using rational design are often unsuccessful. Here the authors describe a massively parallel screening-based strategy whereby millions of potential aptamer switches are synthesised, sequenced and screened directly on a flow-cell.
- Alex M. Yoshikawa
- , Alexandra E. Rangel
- & H. Tom Soh
-
Article
| Open Access Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Implementing high-throughput single-cell DNA sequencing for the study of solid tumours has been challenging. Here, the authors present an optimised approach for snap-frozen tissue single nuclei extraction and DNA sequencing, which can be applied to study pancreatic ductal adenocarcinoma evolution and heterogeneity.
- Haochen Zhang
- , Elias-Ramzey Karnoub
- & Christine A. Iacobuzio-Donahue
-
Article
| Open Access The rapid and highly parallel identification of antibodies with defined biological activities by SLISY
The rapid and highly parallel identification of antibodies with defined biological activities by SLISY
The covid pandemic has highlighted the need for rapid antibody development. Here, authors develop an approach called SLISY, which uses NGS with phage display to simultaneously assess millions of clones to rapidly isolate specific antibodies against SARS-CoV-2 and its evolving variants.
- Steve Lu
- , Austin K. Mattox
- & Kenneth W. Kinzler
-
Article
| Open Access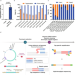 TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
Methods to predict genome-wide off-target activities of prime editors (PEs) are currently lacking. Here the authors report a cell-based assay, TAgmentation of Prime Editor sequencing (TAPE-seq), that provides genome-wide off-target candidates for PEs.
- Jeonghun Kwon
- , Minyoung Kim
- & Jungjoon K. Lee
-
Article
| Open Access Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Adding library adaptors to DNA samples is an essential step in preparing samples for next-generation sequencing. Here, Gunter et al. describe the development of Control Library Adaptors (CAPTORs), that correct sequencing errors and normalise quantitative biases in Nanopore libraries.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota
HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota
Here, the authors construct 102 complete metagenome-assembled genomes (cMAGs) from Pacific Biosciences (PacBio) high-accuracy long-read (HiFi) metagenomic sequencing, showing as high nucleotide accuracy as reference genomes and revealing that regions hard to assemble by short-read sequencing comprise mostly of genomic islands and rRNAs.
- Chan Yeong Kim
- , Junyeong Ma
- & Insuk Lee
-
Article
| Open Access GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
Plant genotype alone appears to be insufficient to explain trait variations. This study integrates GWAS, MWAS and mGWAS in 827 foxtail millet cultivars, revealing that root-associated microbiota affect plant phenotypes in a host genotype-dependent manner.
- Yayu Wang
- , Xiaolin Wang
- & Huan Liu
-
Article
| Open Access Heterogeneity and transcriptome changes of human CD8+ T cells across nine decades of life
Heterogeneity and transcriptome changes of human CD8+ T cells across nine decades of life
The characterisation of T cells during aging is important to predict functional outcomes in vaccination or infection. Here the authors use flow cytometry and scRNA sequencing to transcriptionally age CD8 T cells and then use a machine learning model to interpret cell age from transcriptional profiles.
- Jian Lu
- , Raheel Ahmad
- & Nan-ping Weng
-
Article
| Open Access Integrated multi-omics reveals cellular and molecular interactions governing the invasive niche of basal cell carcinoma
Integrated multi-omics reveals cellular and molecular interactions governing the invasive niche of basal cell carcinoma
The role of reciprocal tumour-stroma interactions in tumour invasion remains poorly characterised. Here, single-cell and spatial transcriptomics identifies the cell populations and their transcriptional reprogramming contributing to the spatial organization of the basal cell carcinoma invasive niche.
- Laura Yerly
- , Christine Pich-Bavastro
- & François Kuonen
-
Article
| Open Access Cutaneous and acral melanoma cross-OMICs reveals prognostic cancer drivers associated with pathobiology and ultraviolet exposure
Cutaneous and acral melanoma cross-OMICs reveals prognostic cancer drivers associated with pathobiology and ultraviolet exposure
While cutaneous melanoma is linked to UV radiation, acral melanoma is not. Epigenetic mechanisms function as sensors to exposures and determinants of cell identity. Here, the authors use DNA methylation data to identify dysregulated pathways associated with UV radiation and pathobiology in cutaneous and acral melanomas.
- Anna Luiza Silva Almeida Vicente
- , Alexei Novoloaca
- & Akram Ghantous
-
Article
| Open Access Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project
Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project
Biobanks of genetic data have been primarily in European populations, which gives us an incomplete understanding of complex traits across populations. Here, the authors initiate the Westlake BioBank for Chinese (WBBC) pilot project with 4,535 whole genome sequences and 5,841 high-density genotypes from China, characterizing large-scale genomic variation in Chinese populations.
- Pei-Kuan Cong
- , Wei-Yang Bai
- & Hou-Feng Zheng
-
Article
| Open Access A rare variant analysis framework using public genotype summary counts to prioritize disease-predisposition genes
A rare variant analysis framework using public genotype summary counts to prioritize disease-predisposition genes
Sequencing studies in clinical and cancer genomics often utilize public data sets to identify genes enriched with pathogenic variants. Here, the authors propose a framework which controls for confounding factors that can bias the results in these studies.
- Wenan Chen
- , Shuoguo Wang
- & Gang Wu
-
Article
| Open Access Contribution of rare whole-genome sequencing variants to plasma protein levels and the missing heritability
Contribution of rare whole-genome sequencing variants to plasma protein levels and the missing heritability
Despite the success of genome-wide association studies, much of the genetic contribution to complex traits remains unexplained. Here, the authors identify effects by rare variants on plasma proteins, and estimate the contribution of rare variants to the heritability.
- Marcin Kierczak
- , Nima Rafati
- & Åsa Johansson
-
Article
| Open Access Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
T cell large granular lymphocyte leukemia (T-LGLL) and the cellular phenotype underlying response to therapy is not well understood. Here the authors use single cell sequencing to better understand changes in T cell clonal frequency and gene expression before and after therapy in T-LGLL.
- Shouguo Gao
- , Zhijie Wu
- & Neal S. Young
-
Article
| Open Access Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE)
Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE)
The design of highly multiplex PCR primers to amplify and enrich many different DNA sequences is increasing in biomedical importance as new mutations and pathogens are identified. The authors present and experimentally validate Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE), a stochastic algorithm for design of highly multiplex PCR primer sets that minimize primer dimer formation.
- Nina G. Xie
- , Michael X. Wang
- & David Yu Zhang
-
Article
| Open Access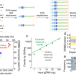 Ensemble of nucleic acid absolute quantitation modules for copy number variation detection and RNA profiling
Ensemble of nucleic acid absolute quantitation modules for copy number variation detection and RNA profiling
Highly multiplexed ddPCR for sensitive nucleic acid quantitation remains challenging due to limited fluorescence channels. Here the authors report quantitative amplicon sequencing (QASeq), a PCR-based molecular barcoding NGS approach which is compatible with high multiplexing.
- Lucia Ruojia Wu
- , Peng Dai
- & David Yu Zhang
-
Article
| Open Access Antibody escape and global spread of SARS-CoV-2 lineage A.27
Antibody escape and global spread of SARS-CoV-2 lineage A.27
The A.27 SARS-CoV-2 lineage spread globally in 2021 but did not become dominant. Here, the authors show that A.27 shares some mutations in the spike gene that are present in variants of concern, but lacks the D614G mutation, indicating independent evolution of immune escape properties.
- Tamara Kaleta
- , Lisa Kern
- & Jonas Fuchs
-
Article
| Open Access High-throughput identification and quantification of single bacterial cells in the microbiota
High-throughput identification and quantification of single bacterial cells in the microbiota
Here, Jin et al., develop a method called Barcoding Bacteria for Identification and Quantification (BarBIQ), which allows to both characterize the global microbiome and to identify and quantify single-cell bacterial members in a high-throughput manner.
- Jianshi Jin
- , Reiko Yamamoto
- & Katsuyuki Shiroguchi
-
Article
| Open Access Cell-fate transition and determination analysis of mouse male germ cells throughout development
Cell-fate transition and determination analysis of mouse male germ cells throughout development
The full-term developmental profile of male germ cells remains undefined. Here, the authors use single-cell sequencing to investigate the transcriptome landscapes of mouse male germ cells throughout development and find several critical regulators for prenatal cell-fate determination.
- Jiexiang Zhao
- , Ping Lu
- & Xiao-Yang Zhao
-
Article
| Open Access Calibration-free NGS quantitation of mutations below 0.01% VAF
Calibration-free NGS quantitation of mutations below 0.01% VAF
Quantification of rare somatic mutations is essential for basic research and translational applications. Here the authors present Quantitative Blocker Displacement Amplification allowing for accurate detection of mutations below 0.01% VAF.
- Peng Dai
- , Lucia Ruojia Wu
- & David Yu Zhang
-
Article
| Open Access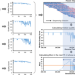 Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Current methods to identify G-quadruplex structures in DNA require specialized protocols and multiple rounds of sequencing. Here, the authors develop a method to detect G-quadruplex structures in DNA based on fluctuations in sequencing quality in a standard sequencing experiment.
- Jing Tu
- , Mengqin Duan
- & Zuhong Lu
-
Article
| Open Access Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Identifying associations of rare variants with disease is challenging due to small effect sizes, technical artefacts and population structure heterogeneity. Here, the authors present RV-EXCALIBER, a method that uses large summary-level exome data to robustly calibrate rare variant burden.
- Ricky Lali
- , Michael Chong
- & Guillaume Paré
-
Article
| Open Access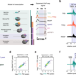 Promoter-proximal elongation regulates transcription in archaea
Promoter-proximal elongation regulates transcription in archaea
Transcription in archaea is known to be regulated through the recruitment of RNA polymerase to promoters. Here, the authors show that the archaeon Saccharolobus solfataricus regulates transcription globally through a rate-limiting promoter-proximal elongation step.
- Fabian Blombach
- , Thomas Fouqueau
- & Finn Werner
-
Article
| Open Access Somatic genetic rescue of a germline ribosome assembly defect
Somatic genetic rescue of a germline ribosome assembly defect
Shwachman-Diamond syndrome (SDS) is a leukemia predisposition disorder that is caused by defective release of eIF6 during ribosome assembly. Here the authors show that acquired somatic EIF6 mutations are frequent in the hematopoietic cells from individuals with SDS and provide a selective advantage over non-modified cells.
- Shengjiang Tan
- , Laëtitia Kermasson
- & Patrick Revy
-
Article
| Open Access The genomic landscape of 85 advanced neuroendocrine neoplasms reveals subtype-heterogeneity and potential therapeutic targets
The genomic landscape of 85 advanced neuroendocrine neoplasms reveals subtype-heterogeneity and potential therapeutic targets
Metastatic and locally-advanced neuroendocrine neoplasms (aNEN) display heterogeneous clinical and genetic characteristics. Here, the authors investigate the mutational landscape of 85 aNEN by whole genome sequencing and identify distinct subpopulations, tumour mutational burden patterns, drivers and actionable somatic alterations.
- Job van Riet
- , Harmen J. G. van de Werken
- & Bianca Mostert
-
Article
| Open Access A deep learning model for predicting next-generation sequencing depth from DNA sequence
A deep learning model for predicting next-generation sequencing depth from DNA sequence
DNA probes used in next generation sequencing (NGS) have variable hybridisation kinetics, resulting in non-uniform coverage. Here, the authors develop a deep learning model to predict NGS depth using DNA probe sequences and apply to human and non-human sequencing panels.
- Jinny X. Zhang
- , Boyan Yordanov
- & David Yu Zhang
-
Article
| Open Access Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Pooling participant-level genetic data into a single analysis can result in variance stratification, reducing statistical performance. Here, the authors develop variant-specific inflation factors to assess variance stratification and apply this to pooled individual-level data from whole genome sequencing.
- Tamar Sofer
- , Xiuwen Zheng
- & Kenneth M. Rice
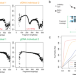
 FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA