Featured
-
-
Article
| Open Access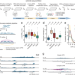 scCircle-seq unveils the diversity and complexity of extrachromosomal circular DNAs in single cells
scCircle-seq unveils the diversity and complexity of extrachromosomal circular DNAs in single cells
Extrachromosomal circular DNAs (eccDNAs) affect gene expression and tumour progression. Here, the authors report a method, scCircle-seq, for eccDNA profiling in single cells, demonstrating the stochasticity, cell type specificity, and dynamics of eccDNAs in cell lines and primary tumour samples.
- Jinxin Phaedo Chen
- , Constantin Diekmann
- & Nicola Crosetto
-
Article
| Open Access scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
Single-cell chromatin accessibility sequencing (scCAS) data suffers from high sparsity and dimensionality. Here, authors propose an accurate and interpretable computational framework for enhancing scCAS data that considers cell-to-cell similarity.
- Songming Tang
- , Xuejian Cui
- & Shengquan Chen
-
Article
| Open Access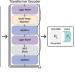 ECOLE: Learning to call copy number variants on whole exome sequencing data
ECOLE: Learning to call copy number variants on whole exome sequencing data
Copy number variants (CNV) are shown to contribute to the etiology of various genetic disorders. Here, authors present ECOLE, a deep learning-based somatic and germline CNV caller for WES data. Utilising a variant of the transformer architecture, the model is trained to call CNVs per exon.
- Berk Mandiracioglu
- , Furkan Ozden
- & A. Ercument Cicek
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access Long-read whole-genome analysis of human single cells
Long-read whole-genome analysis of human single cells
Here the authors introduce a new method to study DNA in single cells by long-read sequencing. Their method gives a more complete view of the genomic structure of individual cells and allows to study genetic differences at the single-cell level.
- Joanna Hård
- , Jeff E. Mold
- & Adam Ameur
-
Article
| Open Access Diploid and tetraploid genomes of Acorus and the evolution of monocots
Diploid and tetraploid genomes of Acorus and the evolution of monocots
Acorales is sister to all other monocots and contains only one family with just one genus, Acorus. Here, the authors assemble the genome of the diploid Ac. gramineus and the tetraploid Ac. calamus, reconstruct an ancestral monocot karyotype and gene toolkit, and discuss the origin and evolution of the two species and other monocots.
- Liang Ma
- , Ke-Wei Liu
- & Zhong-Jian Liu
-
Article
| Open Access The genome of Acorus deciphers insights into early monocot evolution
The genome of Acorus deciphers insights into early monocot evolution
Monocots are one of the most diverse and dominant clades of flowering plants. Here, the authors assemble the genome of Acorus gramineus, confirm its phylogenetic position as sister to the rest of monocots and reveal the absence of tau (τ) whole-genome duplication observed in the majority of monocot clades.
- Xing Guo
- , Fang Wang
- & Huan Liu
-
Article
| Open Access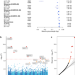 Rare genetic variants impact muscle strength
Rare genetic variants impact muscle strength
Here, the authors provide an exome study of hand grip strength, a proxy of generalized muscle strength. They identify six exome-wide significant genes, with links to disease, and additivity of rare and common genetic variant effects on muscle strength.
- Yunfeng Huang
- , Dora Bodnar
- & Heiko Runz
-
Article
| Open Access Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Prostate cancer is known to have a variable response to androgen receptor signalling inhibitors. Here, the authors use machine learning to predict response to therapy from genomic, transcriptomic and clinical data.
- Anouk C. de Jong
- , Alexandra Danyi
- & Martijn P. Lolkema
-
Article
| Open Access Germline TP53 mutations undergo copy number gain years prior to tumor diagnosis
Germline TP53 mutations undergo copy number gain years prior to tumor diagnosis
Li-Fraumeni syndrome (LFS) is associated with pathogenic germline TP53 variants and predisposes patients to cancer; understanding the evolution and drivers of LFS-related tumours remains crucial. Here, the authors analyse 22 LFS tumours using whole-genome sequencing and reconstruct the evolution and timing of somatic driver alterations.
- Nicholas Light
- , Mehdi Layeghifard
- & Adam Shlien
-
Article
| Open Access Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
The interplay between histone modifications and DNA methylation plays a crucial role in establishing and maintaining the epigenomic landscape. Here, the authors develop a nanopore sequencing based method for mapping histone modifications and DNA methylation from native, long, single DNA molecules.
- Xue Yue
- , Zhiyuan Xie
- & Yimeng Yin
-
Article
| Open Access Whole genome sequencing identifies structural variants contributing to hematologic traits in the NHLBI TOPMed program
Whole genome sequencing identifies structural variants contributing to hematologic traits in the NHLBI TOPMed program
Most genetic association studies have been done on single nucleotide polymorphisms and small indels, while other types of variants have been less studied. Here, the authors use whole genome sequencing in a diverse population to identify and provide experimental evidence for associations between structural variants and blood-cell traits.
- Marsha M. Wheeler
- , Adrienne M. Stilp
- & Alex P. Reiner
-
Article
| Open Access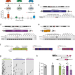 Retrotransposon instability dominates the acquired mutation landscape of mouse induced pluripotent stem cells
Retrotransposon instability dominates the acquired mutation landscape of mouse induced pluripotent stem cells
Retrotransposons are mobile genetic elements normally repressed by DNA methylation in differentiated cells. Here, the authors show that DNA hypomethylation in mouse induced pluripotent stem cells allows retrotransposons to jump, but this can be blocked with a reverse transcriptase inhibitor.
- Patricia Gerdes
- , Sue Mei Lim
- & Geoffrey J. Faulkner
-
Article
| Open Access VeChat: correcting errors in long reads using variation graphs
VeChat: correcting errors in long reads using variation graphs
Consensus sequence-based methods for self-correction of long-read sequencing data are affected by biases that can mask true variants characterizing little-covered or low-frequency haplotypes. Here, to address this issue, the authors develop a variation graph-based method for performing haplotype-aware self-correction of long reads.
- Xiao Luo
- , Xiongbin Kang
- & Alexander Schönhuth
-
Article
| Open Access Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Adding library adaptors to DNA samples is an essential step in preparing samples for next-generation sequencing. Here, Gunter et al. describe the development of Control Library Adaptors (CAPTORs), that correct sequencing errors and normalise quantitative biases in Nanopore libraries.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Characterization of somatic structural variations in 528 Chinese individuals with Esophageal squamous cell carcinoma
Characterization of somatic structural variations in 528 Chinese individuals with Esophageal squamous cell carcinoma
The biological and clinical implications of high genome instability in esophageal squamous cell carcinoma (ESCC) remain to be explored. Here, the authors analyse 528 whole genomes and investigate the landscape and molecular mechanisms underlying structural variations in ESCC patients.
- Heyang Cui
- , Yong Zhou
- & Qimin Zhan
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Association of mitochondrial DNA content, heteroplasmies and inter-generational transmission with autism
Association of mitochondrial DNA content, heteroplasmies and inter-generational transmission with autism
Most genetic studies of autism spectrum disorder (ASD) have focused on the nuclear genome. Here, the authors show that variations in mitochondrial DNA, detectable at birth, are also associated with risk of ASD.
- Yiqin Wang
- , Xiaoxian Guo
- & Zhenglong Gu
-
Article
| Open Access Phasing analysis of lung cancer genomes using a long read sequencer
Phasing analysis of lung cancer genomes using a long read sequencer
Long-read sequencing technologies are useful for the multifaceted task of characterising somatic mutations, including structural variants, in cancers. Here, the authors combine short and long read sequencing for the phasing analysis, which enables them to resolve the chromosomal backgrounds of somatic mutations in Japanese non-small cell lung cancers.
- Yoshitaka Sakamoto
- , Shuhei Miyake
- & Ayako Suzuki
-
Article
| Open Access Whole exome sequencing in Alopecia Areata identifies rare variants in KRT82
Whole exome sequencing in Alopecia Areata identifies rare variants in KRT82
Common variants have been discovered to be associated with Alopecia Areata; however, rare variants have been less well studied. Here, the authors use whole-exome sequencing to identify associated rare variants in the hair keratin gene KRT82. Further, they find that individuals with Alopecia Areata have reduced expression of KRT82 in the skin and hair follicle.
- Stephanie O. Erjavec
- , Sahar Gelfman
- & Angela M. Christiano
-
Article
| Open Access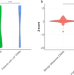 A de novo paradigm for male infertility
A de novo paradigm for male infertility
Germline de novo mutations can impact individual fitness, but their role in human male infertility is understudied. Trio-based exome sequencing identifies many new candidate genes affecting male fertility, including an essential regulator of male germ cell pre-mRNA splicing.
- M. S. Oud
- , R. M. Smits
- & J. A. Veltman
-
Article
| Open Access Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Obtaining accurate variant calls from multiple displacement amplified single cell DNA sequencing data needs dedicated models that account for amplification bias and copy errors. Here, the authors describe ProSolo, a model for calling single nucleotide variants with control over the false discovery rate.
- David Lähnemann
- , Johannes Köster
- & Alexander Schönhuth
-
Article
| Open Access Mapping epigenetic divergence in the massive radiation of Lake Malawi cichlid fishes
Mapping epigenetic divergence in the massive radiation of Lake Malawi cichlid fishes
The Lake Malawi cichlid fishes are an example of extreme vertebrate radiation; however, there is very little sequence divergence among the species. Here the authors present a comparative genome-wide methylome study to suggest DNA methylation played a major role in the extensive phenotypic diversity amongst these fishes.
- Grégoire Vernaz
- , Milan Malinsky
- & Eric A. Miska
-
Article
| Open Access Analysis of 427 genomes reveals moso bamboo population structure and genetic basis of property traits
Analysis of 427 genomes reveals moso bamboo population structure and genetic basis of property traits
Moso bamboo is an economically and ecologically important nontimber forestry species. Here, the authors analyze 427 genomes collected from 15 representative geographic areas, and identify genes under balancing selection, putative patterns of historic demography, and candidate genes associated with important traits.
- Hansheng Zhao
- , Shuai Sun
- & Zhimin Gao
-
Article
| Open Access Haploinsufficiency of SF3B2 causes craniofacial microsomia
Haploinsufficiency of SF3B2 causes craniofacial microsomia
Despite being a common congenital facial anomaly, the genetic etiology of craniofacial microsomia (CFM) is not well understood. Here, the authors use exome and genome sequencing of 146 individuals with CFM to identify haploinsufficient variants in SF3B2 as a prevalent underlying cause.
- Andrew T. Timberlake
- , Casey Griffin
- & Daniela V. Luquetti
-
Article
| Open Access COVseq is a cost-effective workflow for mass-scale SARS-CoV-2 genomic surveillance
COVseq is a cost-effective workflow for mass-scale SARS-CoV-2 genomic surveillance
Genomic surveillance of SARS-CoV-2 is crucial to monitor the spread of variants of concern. A new sequencing method enables cost-effective SARS-CoV-2 genomic surveillance at scale and is easily adaptable to other viruses.
- Michele Simonetti
- , Ning Zhang
- & Nicola Crosetto
-
Article
| Open Access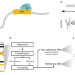 Cas9 targeted enrichment of mobile elements using nanopore sequencing
Cas9 targeted enrichment of mobile elements using nanopore sequencing
Mobile element insertions (MEIs) are a source of repetitive genetic variation and can lead to genetic disorders. Here the authors use Cas9-targeted nanopore sequencing to efficiently saturate enrichment for known and non-reference MEIs.
- Torrin L. McDonald
- , Weichen Zhou
- & Alan P. Boyle
-
Article
| Open Access Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing
Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing
Several existing algorithms predict the methylation of DNA using Nanopore sequencing signals, but it is unclear how they compare in performance. Here, the authors benchmark the performance of several such tools, and propose METEORE, a consensus tool that improves prediction accuracy.
- Zaka Wing-Sze Yuen
- , Akanksha Srivastava
- & Eduardo Eyras
-
Article
| Open Access Joint profiling of DNA and proteins in single cells to dissect genotype-phenotype associations in leukemia
Joint profiling of DNA and proteins in single cells to dissect genotype-phenotype associations in leukemia
It is currently difficult to map DNA variants and surface phenotypes in the same cells, preventing direct linkage of phenotype and genotype. Here the authors report DAb-seq for simultaneous capture of DNA genotype and cell surface phenotype from single cells at high throughput.
- Benjamin Demaree
- , Cyrille L. Delley
- & Adam R. Abate
-
Article
| Open Access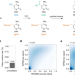 Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Specific and quantitative sequencing of cytosine modifications is challenging at base-resolution. Here the authors present TAPSβ and CAPS for subtraction-free whole genome sequencing of 5mC and 5hmC.
- Yibin Liu
- , Zhiyuan Hu
- & Chun-Xiao Song
-
Article
| Open Access Complete sequences of Schizosaccharomyces pombe subtelomeres reveal multiple patterns of genome variation
Complete sequences of Schizosaccharomyces pombe subtelomeres reveal multiple patterns of genome variation
Sequencing and mapping of long repetitive regions can be challenging due to technical difficulties in sequencing and assembly of the sequence data. Here authors report the complete sequences of subtelomeric homologous (SH) regions of the fission yeast Schizosaccharomyces pombe to reveal highly polymorphic and hot spots for genome variation features.
- Yusuke Oizumi
- , Takuto Kaji
- & Junko Kanoh
-
Article
| Open Access Pan-cancer circulating tumor DNA detection in over 10,000 Chinese patients
Pan-cancer circulating tumor DNA detection in over 10,000 Chinese patients
The detection of aberrations in circulating tumour DNA represents a non-invasive method to survey the oncogenes and tumour suppressors that are modified within a patient’s cancer. Here, the authors analysed more than 10,000 patients using a targeted sequencing panel and report on the frequencies of the mutations that they found.
- Yongliang Zhang
- , Yu Yao
- & Qiang Zeng
-
Article
| Open Access Efficient assembly of nanopore reads via highly accurate and intact error correction
Efficient assembly of nanopore reads via highly accurate and intact error correction
Nanopore reads have been advantageous for de novo genome assembly; however these reads have high error rates. Here, the authors develop an error correction and de novo assembly tool, NECAT, which produces efficient, high quality assemblies of nanopore reads.
- Ying Chen
- , Fan Nie
- & Chuan-Le Xiao
-
Article
| Open Access Photon-directed multiplexed enzymatic DNA synthesis for molecular digital data storage
Photon-directed multiplexed enzymatic DNA synthesis for molecular digital data storage
Writing data in DNA is still a bottleneck due to the reliance on chemical synthesis methods. Here the authors report multiplexed enzymatic DNA synthesis using maskless photolithography.
- Howon Lee
- , Daniel J. Wiegand
- & George M. Church
-
Article
| Open Access Whole genome sequence analysis of pulmonary function and COPD in 19,996 multi-ethnic participants
Whole genome sequence analysis of pulmonary function and COPD in 19,996 multi-ethnic participants
Chronic obstructive pulmonary disease is a leading cause of morbidity and mortality. Here, the authors analyse whole genome sequence data and find new loci associated with lung function and COPD.
- Xutong Zhao
- , Dandi Qiao
- & Ani Manichaikul
-
Article
| Open Access Framework for quality assessment of whole genome cancer sequences
Framework for quality assessment of whole genome cancer sequences
Working with cancer genomes from multiple projects can increase investigative power, but quality of sequences can vary. Here, the authors present a framework for comparing whole genome sequencing quality to help researchers guide downstream analyses and exclude poor quality samples.
- Justin P. Whalley
- , Ivo Buchhalter
- & Ivo G. Gut
-
Article
| Open Access Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment
Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment
Off-target mutations that occur at a frequency below 0.5% can be difficult to detect. Here the authors use predicted off-target amplification to increase detection sensitivity.
- Seung-Hun Kang
- , Wi-jae Lee
- & Seung Hwan Lee
-
Article
| Open Access Disruption of the tumour-associated EMP3 enhances erythroid proliferation and causes the MAM-negative phenotype
Disruption of the tumour-associated EMP3 enhances erythroid proliferation and causes the MAM-negative phenotype
The molecular basis of the clinically important MAM blood group antigen present in most humans is unknown. We identify EMP3 as its encoding gene, establishing MAM as a new blood group system, and demonstrate the role of EMP3 in erythropoiesis through its interaction with the signalling molecule CD44.
- Nicole Thornton
- , Vanja Karamatic Crew
- & David J. Anstee
-
Article
| Open Access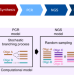 Quantifying molecular bias in DNA data storage
Quantifying molecular bias in DNA data storage
DNA is an attractive digital data storing medium due to high information density and longevity. Here the authors use millions of sequences to investigate inherent biases in DNA synthesis and PCR amplification.
- Yuan-Jyue Chen
- , Christopher N. Takahashi
- & Karin Strauss
-
Article
| Open Access The mutREAD method detects mutational signatures from low quantities of cancer DNA
The mutREAD method detects mutational signatures from low quantities of cancer DNA
Sequencing tumour genomes can reveal information about the processes that drive the formation of cancer. Here, the authors describe a method that can detect these mutational signatures from small amounts of DNA and degraded samples.
- Juliane Perner
- , Sujath Abbas
- & Rebecca C. Fitzgerald
-
Article
| Open Access Functional annotation of rare structural variation in the human brain
Functional annotation of rare structural variation in the human brain
Structural variants (SVs) contribute to the genetic architecture of many brain-related disorders. Here, the authors integrate SV calls from genome sequencing (n = 755) with RNA-seq data (n = 629) from post-mortem dorsal lateral prefrontal cortex to annotate the gene regulatory effects of SVs in the human brain and their potential to contribute to disease.
- Lide Han
- , Xuefang Zhao
- & Douglas M. Ruderfer
-
Article
| Open Access Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
The complexity of structural variation (SV) and short tandem repeats (STRs) makes it necessary to apply different calling and filtering strategies to sequencing datasets. Here, Jakubosky et al. report a comprehensive SV and STR callset from whole-genome sequencing of 477 individuals from iPSCORE and HipSci using five algorithms.
- David Jakubosky
- , Erin N. Smith
- & Kelly A. Frazer
-
Article
| Open Access Ancient genomes from northern China suggest links between subsistence changes and human migration
Ancient genomes from northern China suggest links between subsistence changes and human migration
Northern China contains some of the world’s earliest farming societies. Here, authors use 55 ancient genomes to trace the genetic history of human migrations across northern China for the last 7500 years, and document genetic changes mirroring shifts in subsistence strategy.
- Chao Ning
- , Tianjiao Li
- & Yinqiu Cui
-
Article
| Open Access ORF Capture-Seq as a versatile method for targeted identification of full-length isoforms
ORF Capture-Seq as a versatile method for targeted identification of full-length isoforms
Most human protein-coding genes are expressed as multiple isoforms. Here the authors present ORF Capture-seq that uses cloned ORFs as probes to capture and sequence full length transcript sequences. This enables highly sensitive characterization of eukaryotic transcriptomes.
- Gloria M. Sheynkman
- , Katharine S. Tuttle
- & Marc Vidal
-
Article
| Open Access Increased burden of ultra-rare structural variants localizing to boundaries of topologically associated domains in schizophrenia
Increased burden of ultra-rare structural variants localizing to boundaries of topologically associated domains in schizophrenia
Common variants identified by large-scale genomewide association studies cannot account fully account for the heritability of schizophrenia (SCZ). Here, the authors report high-coverage whole-genome sequencing of 1162 SCZ cases and 936 controls and explore the contribution of different types of variants to SCZ.
- Matthew Halvorsen
- , Ruth Huh
- & Jin P. Szatkiewicz
-
Article
| Open Access Integrated proteogenomic deep sequencing and analytics accurately identify non-canonical peptides in tumor immunopeptidomes
Integrated proteogenomic deep sequencing and analytics accurately identify non-canonical peptides in tumor immunopeptidomes
Non-canonical HLA-bound peptides from presumed non-coding regions are potential targets for cancer immunotherapy, but their discovery remains challenging. Here, the authors integrate exome sequencing, transcriptomics, ribosome profiling, and immunopeptidomics to identify tumor-specific non-canonical HLA-bound peptides.
- Chloe Chong
- , Markus Müller
- & Michal Bassani-Sternberg
-
Article
| Open Access Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha is an extremely fast-growing invasive plant species that can cause serious damage to natural ecosystems. Here, the authors assemble its chromosome-scale reference genome and explore possible mechanisms that contribute to its rapid growth.
- Bo Liu
- , Jian Yan
- & Fanghao Wan
-
Article
| Open Access Identification of recurrent FHL2-GLI2 oncogenic fusion in sclerosing stromal tumors of the ovary
Identification of recurrent FHL2-GLI2 oncogenic fusion in sclerosing stromal tumors of the ovary
Little is known about the genetics of sclerosing stromal tumor of the ovary, a rare type of sex cord-stromal tumor. Here, the authors use sequencing strategies to show that in a cohort of 26 tumor samples 65% carry a FHL2-GLI2 fusion gene and demonstrate in vitro that the fusion gene has oncogenic properties.
- Sarah H. Kim
- , Arnaud Da Cruz Paula
- & Britta Weigelt
-
Article
| Open Access A 5700 year-old human genome and oral microbiome from chewed birch pitch
A 5700 year-old human genome and oral microbiome from chewed birch pitch
Birch pitch is thought to have been used in prehistoric times as hafting material or antiseptic and tooth imprints suggest that it was chewed. Here, the authors report a 5,700 year-old piece of chewed birch pitch from Denmark from which they successfully recovered a complete ancient human genome and oral microbiome DNA.
- Theis Z. T. Jensen
- , Jonas Niemann
- & Hannes Schroeder

 High clonal diversity and spatial genetic admixture in early prostate cancer and surrounding normal tissue
High clonal diversity and spatial genetic admixture in early prostate cancer and surrounding normal tissue