Featured
-
-
Article
| Open Access Apparent bias toward long gene misregulation in MeCP2 syndromes disappears after controlling for baseline variations
Apparent bias toward long gene misregulation in MeCP2 syndromes disappears after controlling for baseline variations
Recent studies have suggested that long genes (>100 kb) are more likely to be misregulated in some neurological diseases, such as autism and Rett syndrome. Here the authors find that the apparent length-dependent trends previously observed in MeCP2 microarray and RNA-sequencing datasets disappeared after controlling for baseline variations.
- Ayush T. Raman
- , Amy E. Pohodich
- & Zhandong Liu
-
Article
| Open Access Global pairwise RNA interaction landscapes reveal core features of protein recognition
Global pairwise RNA interaction landscapes reveal core features of protein recognition
RNA–protein interactions often depend on the recognition of extended RNA elements but the identification of these motifs is challenging. Here, the authors present a global integrated approach to analyze RNA–protein binding landscapes, mapping extended RNA interaction motifs for four RNA-binding proteins.
- Qin Zhou
- , Nikesh Kunder
- & Zachary T. Campbell
-
Article
| Open Access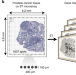 Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity
Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity
Heterogeneity within tumors presents a challenge to cancer treatment. Here, the authors investigate transcriptional heterogeneity in prostate cancer, examining expression profiles of different tissue components and highlighting expression gradients in the tumor microenvironment.
- Emelie Berglund
- , Jonas Maaskola
- & Joakim Lundeberg
-
Article
| Open Access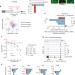 Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
The function and consequences of alternative polyadenylation (APA) in stressed cells are largely unclear. Here, the authors show that stress-induced mRNA degradation depends on 3′UTR length and that APA-mediated 3′UTR shortening is an adaptive stress response mechanism for selective transcript stabilization.
- Dinghai Zheng
- , Ruijia Wang
- & Bin Tian
-
Article
| Open Access Distinct epigenomic patterns are associated with haploinsufficiency and predict risk genes of developmental disorders
Distinct epigenomic patterns are associated with haploinsufficiency and predict risk genes of developmental disorders
Predicting haploinsufficient genes helps to understand the genetic risk underlying developmental disorders. Here, the authors develop a Random Forest-based method that uses epigenomic data to predict haploinsufficiency, Episcore, which is complementary to methods based on mutation intolerance scores.
- Xinwei Han
- , Siying Chen
- & Yufeng Shen
-
Article
| Open Access Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity
Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity
Epithelial subpopulations are present in the human breast but how these differentiate or form is unclear. Here, the authors use single-cell RNA sequencing of primary human breast epithelial cells to define previously undescribed luminal, basal epithelial subpopulations and ZEB1-positive basal cells.
- Quy H. Nguyen
- , Nicholas Pervolarakis
- & Kai Kessenbrock
-
Article
| Open Access High-throughput screening of prostate cancer risk loci by single nucleotide polymorphisms sequencing
High-throughput screening of prostate cancer risk loci by single nucleotide polymorphisms sequencing
Functional characterization of disease-causing variants at risk loci in cancer is challenging. Here, in prostate cancer the authors report a pipeline for high-throughput single-nucleotide polymorphisms sequencing (SNPs-seq) for large scale screening of functional SNPs at disease risk loci.
- Peng Zhang
- , Ji-Han Xia
- & Liang Wang
-
Article
| Open Access The human Vδ2+ T-cell compartment comprises distinct innate-like Vγ9+ and adaptive Vγ9- subsets
The human Vδ2+ T-cell compartment comprises distinct innate-like Vγ9+ and adaptive Vγ9- subsets
Human Vδ2+ γδ T cells are thought to be an innate-like T-cell population. Here the authors show the Vδ2+ compartment contains both innate-like Vγ9+ and an adaptive Vγ9- subset that undergoes clonal expansion during viral infection and can infiltrate liver tissue.
- Martin S. Davey
- , Carrie R. Willcox
- & Benjamin E. Willcox
-
Article
| Open Access Deep whole-genome sequencing reveals recent selection signatures linked to evolution and disease risk of Japanese
Deep whole-genome sequencing reveals recent selection signatures linked to evolution and disease risk of Japanese
Recent natural selection left signals in human genomes. Here, Okada et al. generate high-depth whole-genome sequence (WGS) data (25.9×) from 2,234 Japanese people of the BioBank Japan Project (BBJ), and identify signals of recent natural selection which overlap variants associated with human traits.
- Yukinori Okada
- , Yukihide Momozawa
- & Yoichiro Kamatani
-
Article
| Open Access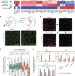 Histone H3.3 sub-variant H3mm7 is required for normal skeletal muscle regeneration
Histone H3.3 sub-variant H3mm7 is required for normal skeletal muscle regeneration
Incorporation of histone H3 variant H3.3 into chromatin regulates transcription. Here the authors find that H3.3 sub-variant H3mm7 is required for skeletal muscle regeneration and that H3mm7 nucleosomes are unstable and exhibit higher mobility, with H3mm7 promoting open chromatin around promoters.
- Akihito Harada
- , Kazumitsu Maehara
- & Yasuyuki Ohkawa
-
Article
| Open Access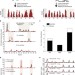 Genome-wide identification of natural RNA aptamers in prokaryotes and eukaryotes
Genome-wide identification of natural RNA aptamers in prokaryotes and eukaryotes
Riboswitches recognize and respond to specific metabolites by altering gene expression. Here, the authors developed a high-throughput method (PARCEL) to experimentally identify RNA aptamers across transcriptomes.
- Sidika Tapsin
- , Miao Sun
- & Yue Wan
-
Article
| Open Access A complex epistatic network limits the mutational reversibility in the influenza hemagglutinin receptor-binding site
A complex epistatic network limits the mutational reversibility in the influenza hemagglutinin receptor-binding site
The receptor-binding site (RBS) of influenza A viruses evolves to evade immune pressure, while maintaining efficient attachment to the host receptor. Wu et al. here identify the complex epistatic network in RBS of H3N2 viruses that limits reversibility of naturally occurring mutations to retain infectivity.
- Nicholas C. Wu
- , Andrew J. Thompson
- & Ian A. Wilson
-
Article
| Open Access BEARscc determines robustness of single-cell clusters using simulated technical replicates
BEARscc determines robustness of single-cell clusters using simulated technical replicates
Single cell messenger RNAseq allows the study of heterogeneity in tissue samples. Here the authors present BEARscc, a tool that uses RNA spike-in controls to simulate experiment-specific technical replicates to estimate the robustness of computational predictions of subpopulations of cells.
- D. T. Severson
- , R. P. Owen
- & B. Schuster-Böckler
-
Article
| Open Access Nanoplanktonic diatoms are globally overlooked but play a role in spring blooms and carbon export
Nanoplanktonic diatoms are globally overlooked but play a role in spring blooms and carbon export
Diatoms are major oceanic primary producers, but some species belonging to the nano- and even picoplankton size are poorly characterized. Here the authors describe a massive spring bloom of the smallest known diatom in the Mediterranean Sea and reveal their general oversight at the global scale.
- Karine Leblanc
- , Bernard Quéguiner
- & Pascal Conan
-
Article
| Open Access Somatic mutagenesis in satellite cells associates with human skeletal muscle aging
Somatic mutagenesis in satellite cells associates with human skeletal muscle aging
Aging skeletal muscle shows declining numbers and activity of satellite cells. Here, Franco et al. show that in satellite cells of the human leg muscle vastus lateralis, somatic mutations accumulate with age and that these mutations become enriched in exons and promoters of genes involved in muscle function.
- Irene Franco
- , Anna Johansson
- & Maria Eriksson
-
Article
| Open Access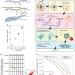 Single-cell full-length total RNA sequencing uncovers dynamics of recursive splicing and enhancer RNAs
Single-cell full-length total RNA sequencing uncovers dynamics of recursive splicing and enhancer RNAs
Total RNA sequencing has been used to profile poly(A) and non-poly(A) RNA expression, processing and the activity of enhancers. Here the authors develop RamDA-seq, a method for full-length total RNA sequencing in single cells.
- Tetsutaro Hayashi
- , Haruka Ozaki
- & Itoshi Nikaido
-
Article
| Open Access High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell
High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell
Long-read sequencing technologies facilitate efficient and high quality genome assembly. Here Michael et al. achieve a fast reference assembly for Arabidopsis thaliana KBS-Mac-74 accession using the handheld Oxford Nanopore MinION sequencer and consumer computing hardware, and demonstrate its usefulness in resolving complex structural variation.
- Todd P. Michael
- , Florian Jupe
- & Joseph R. Ecker
-
Article
| Open Access Cellular stressors contribute to the expansion of hematopoietic clones of varying leukemic potential
Cellular stressors contribute to the expansion of hematopoietic clones of varying leukemic potential
Cellular stressors can impact clonal hematopoiesis. Here, the authors explore the impact of cytotoxic therapy and hematopoietic transplantation on clonal expansion, suggesting different stressors can promote expansion of distinct long-lived clones.
- Terrence N. Wong
- , Christopher A. Miller
- & Daniel C. Link
-
Article
| Open Access Nucleotide resolution mapping of influenza A virus nucleoprotein-RNA interactions reveals RNA features required for replication
Nucleotide resolution mapping of influenza A virus nucleoprotein-RNA interactions reveals RNA features required for replication
Influenza A virus packaging depends on interactions between nucleoprotein (NP) and viral RNA (vRNA), but the pattern of NP binding is unclear. Using PAR-CLIP, Williams et al. here show that NP binds vRNA non-uniformly and that RNA structures in low-NP binding regions are important for packaging.
- Graham D. Williams
- , Dana Townsend
- & Adrianus C. M. Boon
-
Article
| Open Access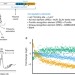 PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs
PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs
A large number of RNA binding proteins (RBPs) and miRNAs bind to the 3′ untranslated regions of mRNA, but methods to dissect their function and interactions are lacking. Here the authors introduce post-transcriptional regulatory element sequencing (PTRE-seq) to dissect sequence preferences, interactions and consequences of RBP and miRNA binding.
- Kyle A. Cottrell
- , Hemangi G. Chaudhari
- & Sergej Djuranovic
-
Article
| Open Access Extreme haplotype variation in the desiccation-tolerant clubmoss Selaginella lepidophylla
Extreme haplotype variation in the desiccation-tolerant clubmoss Selaginella lepidophylla
Selaginella lepidophylla is a clubmoss with extreme desiccation tolerance. Here, the authors assemble its highly heterozygotic haplotypes and examine gene expression changes during desiccation, which shed light on the mechanisms for maintaining a small genome size and adaptation to extreme drying.
- Robert VanBuren
- , Ching Man Wai
- & Todd P. Michael
-
Article
| Open Access Reading and editing the Pleurodeles waltl genome reveals novel features of tetrapod regeneration
Reading and editing the Pleurodeles waltl genome reveals novel features of tetrapod regeneration
The Iberian ribbed newt Pleurodeles waltl has a wide spectrum of regeneration abilities. Here, Elewa et al. sequence its ~20 Gb genome and transcriptome to investigate the molecular features underlying its regenerative capacities.
- Ahmed Elewa
- , Heng Wang
- & András Simon
-
Article
| Open Access Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Regeneration capable flatworms have emerged as powerful models for studying stem cell biology and patterning, however their study has been hindered by the lack of transgenesis methods. Here, the authors describe a transgenesis method for Macrostomum lignano, as well as a new annotated genome sequence.
- Jakub Wudarski
- , Daniil Simanov
- & Eugene Berezikov
-
Article
| Open Access The Kalanchoë genome provides insights into convergent evolution and building blocks of crassulacean acid metabolism
The Kalanchoë genome provides insights into convergent evolution and building blocks of crassulacean acid metabolism
Crassulacean acid metabolism (CAM) is a metabolic adaptation of photosynthesis that enhances water use efficiency. Here, via genomic analysis of Kalanchoë, the authors provide evidence for convergent evolution of protein sequence and temporal gene expression underpinning the multiple independent emergences of CAM.
- Xiaohan Yang
- , Rongbin Hu
- & Gerald A. Tuskan
-
Article
| Open Access Scallop genome reveals molecular adaptations to semi-sessile life and neurotoxins
Scallop genome reveals molecular adaptations to semi-sessile life and neurotoxins
Bivalve molluscs have evolved various characteristics to adapt to benthic filter-feeding. Here, Li et al investigate the genome, transcriptomes and proteomes of scallop Chlamys farreri, revealing evidences of molecular adaptations to semi-sessile life and neurotoxins.
- Yuli Li
- , Xiaoqing Sun
- & Zhenmin Bao
-
Article
| Open Access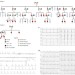 Loss-of-activity-mutation in the cardiac chloride-bicarbonate exchanger AE3 causes short QT syndrome
Loss-of-activity-mutation in the cardiac chloride-bicarbonate exchanger AE3 causes short QT syndrome
Mutations in potassium and calcium channel genes have been associated with cardiac arrhythmias. Here, Jensen et al. show that an anion transporter chloride-bicarbonate exchanger AE3 is also responsible for the genetically-induced mechanism of cardiac arrhythmia, suggesting new therapeutic targets for this disease
- Kasper Thorsen
- , Vibeke S. Dam
- & Henrik K. Jensen
-
Article
| Open Access Prevalence and detection of low-allele-fraction variants in clinical cancer samples
Prevalence and detection of low-allele-fraction variants in clinical cancer samples
High-throughput sequencing is used to identify somatic variants in cancer patients. Here, the authors perform panel-based profiling of 5095 clinical samples and demonstrate that many clinically-actionable variants have low variant allele fractions, requiring assays with high detection sensitivity.
- Hyun-Tae Shin
- , Yoon-La Choi
- & Woong-Yang Park
-
Article
| Open Access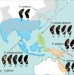 Tracing the origin and evolution of supergene mimicry in butterflies
Tracing the origin and evolution of supergene mimicry in butterflies
Wing pattern mimicry in the butterfly Papilio polytes is controlled by a single Mendelian locus, the mimicry supergene doublesex. Here, Zhang and colleagues reconstruct the complex evolutionary history of the doublesex supergene and mimicry in the Papilio polytes species group.
- Wei Zhang
- , Erica Westerman
- & Marcus R. Kronforst
-
Article
| Open Access Mapping and phasing of structural variation in patient genomes using nanopore sequencing
Mapping and phasing of structural variation in patient genomes using nanopore sequencing
The detection of structural variants can be difficult with short-read sequencing technology, especially when variants are highly complex. Here, the authors use a MinION nanopore sequencer to analyse two patient genomes and develop NanoSV to map known and novel structural variants in long read data.
- Mircea Cretu Stancu
- , Markus J. van Roosmalen
- & Wigard P. Kloosterman
-
Article
| Open Access Dense and accurate whole-chromosome haplotyping of individual genomes
Dense and accurate whole-chromosome haplotyping of individual genomes
Haplotype information is important in investigating many biological phenomena. Here, Porubsky et al. combine Strand-seq with long-read or linked-read sequencing to obtain complete and genome-wide haplotypes of a single individual genome at manageable costs.
- David Porubsky
- , Shilpa Garg
- & Tobias Marschall
-
Article
| Open Access Coding and noncoding landscape of extracellular RNA released by human glioma stem cells
Coding and noncoding landscape of extracellular RNA released by human glioma stem cells
While circulating DNA has been extensively explored as a potential cancer biomarker, RNA potential has been overlooked so far. Here the authors present a comprehensive analysis of extracellular RNA secreted by glioblastoma cells that could prove a valuable resource for biomarker discovery and a means of intercellular communication.
- Zhiyun Wei
- , Arsen O. Batagov
- & Anna M. Krichevsky
-
Article
| Open Access Single nucleus sequencing reveals spermatid chromosome fragmentation as a possible cause of maize haploid induction
Single nucleus sequencing reveals spermatid chromosome fragmentation as a possible cause of maize haploid induction
Plant breeders can produce haploid maize lines using haploid inducer lines as pollen donors. Here, by sequencing the genomes of single pollen nuclei, Li et al. show that haploid inducer spermatids are frequently aneuploid and suggest chromosome fragmentation as a possible cause of haploid induction.
- Xiang Li
- , Dexuan Meng
- & Jianbing Yan
-
Article
| Open Access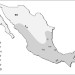 Demographic history and biologically relevant genetic variation of Native Mexicans inferred from whole-genome sequencing
Demographic history and biologically relevant genetic variation of Native Mexicans inferred from whole-genome sequencing
People of Mexico have diverse historical and genetic background. Here, Romero-Hidalgo and colleagues sequence whole genomes of Native Americans of Mexico, and show demographic history and genetic variation shared among subgroups of Native Americans.
- Sandra Romero-Hidalgo
- , Adrián Ochoa-Leyva
- & Xavier Soberón
-
Article
| Open Access Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis
Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis
Amyotrophic lateral sclerosis (ALS) is a rapidly progressing neurodegenerative disease. Here, Wray and colleagues identify association of the GPX3-TNIP1 locus with ALS using cross-ethnic meta-analyses.
- Beben Benyamin
- , Ji He
- & Dongsheng Fan
-
Article
| Open Access Identifying DNase I hypersensitive sites as driver distal regulatory elements in breast cancer
Identifying DNase I hypersensitive sites as driver distal regulatory elements in breast cancer
Cancer driver mutations can occur within noncoding genomic sequences. Here, the authors develop a statistical approach to identify candidate noncoding driver mutations in DNase I hypersensitive sites in breast cancer and experimentally demonstrate they are regulatory elements of known cancer genes.
- Matteo D′Antonio
- , Donate Weghorn
- & Kelly A Frazer
-
Article
| Open Access Annotating pathogenic non-coding variants in genic regions
Annotating pathogenic non-coding variants in genic regions
While non-coding synonymous and intronic variants are often not under strong selective constraint, they can be pathogenic through affecting splicing or transcription. Here, the authors develop a score that uses sequence context alterations to predict pathogenicity of synonymous and non-coding genetic variants, and provide a web server of pre-computed scores.
- Sahar Gelfman
- , Quanli Wang
- & David B. Goldstein
-
Article
| Open Access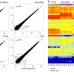 Nanogrid single-nucleus RNA sequencing reveals phenotypic diversity in breast cancer
Nanogrid single-nucleus RNA sequencing reveals phenotypic diversity in breast cancer
Single cell RNA sequencing is a powerful tool for understanding cellular diversity but is limited by cost, throughput and sample preparation. Here the authors use nanogrid technology with integrated imaging to sequence thousands of cancer nuclei in parallel from fresh or frozen tissue.
- Ruli Gao
- , Charissa Kim
- & Nicholas Navin
-
Article
| Open Access PLATE-Seq for genome-wide regulatory network analysis of high-throughput screens
PLATE-Seq for genome-wide regulatory network analysis of high-throughput screens
Despite the importance of pharmacological and functional genomic screens the readouts are of low complexity. Here the authors introduce PLATE-Seq, a low-cost genome-wide mRNA profiling method to complement high-throughput screening.
- Erin C. Bush
- , Forest Ray
- & Peter A. Sims
-
Article
| Open Access Nanopore long-read RNAseq reveals widespread transcriptional variation among the surface receptors of individual B cells
Nanopore long-read RNAseq reveals widespread transcriptional variation among the surface receptors of individual B cells
Short-read RNA-seq is limited in its ability to resolve complex transcript isoforms since it cannot sequence full-length cDNA. Here the authors use Oxford Nanopore MinION and their Mandalorion analysis pipeline to measure complex isoforms in B1a cells.
- Ashley Byrne
- , Anna E. Beaudin
- & Christopher Vollmers
-
Article
| Open Access Generation and comparison of CRISPR-Cas9 and Cre-mediated genetically engineered mouse models of sarcoma
Generation and comparison of CRISPR-Cas9 and Cre-mediated genetically engineered mouse models of sarcoma
Site-specific recombination and CRISPR-Cas9 have been used to generate genetically engineered mouse models of cancer. Here the authors compare sarcomas generated using both systems and see similar genetic and cellular phenotypes, suggesting CRISPR-Cas9 can be used to rapidly generate sarcoma models.
- Jianguo Huang
- , Mark Chen
- & David G. Kirsch
-
Article
| Open Access Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis
Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis
RNA-seq is widely used for transcriptome analysis. Here, the authors analyse a wide spectrum of RNA-seq workflows and present a comprehensive analysis protocol named RNACocktail as well as a computational pipeline leveraging the widely used tools for accurate RNA-seq analysis.
- Sayed Mohammad Ebrahim Sahraeian
- , Marghoob Mohiyuddin
- & Hugo Y. K. Lam
-
Article
| Open Access Flipping between Polycomb repressed and active transcriptional states introduces noise in gene expression
Flipping between Polycomb repressed and active transcriptional states introduces noise in gene expression
Polycomb repressive complexes modify histones but it is unclear how changes in chromatin states alter kinetics of transcription. Here, the authors use single-cell RNAseq and ChIPseq to find that actively transcribed genes with Polycomb marks have greater cell-to-cell variation in expression.
- Gozde Kar
- , Jong Kyoung Kim
- & Sarah A. Teichmann
-
Article
| Open Access Single-virus genomics reveals hidden cosmopolitan and abundant viruses
Single-virus genomics reveals hidden cosmopolitan and abundant viruses
Viruses play an important role in microbial communities but, due to limitations of available techniques, our understanding of viral diversity is limited. Here, the authors use SVGs and identify highly abundant viruses in marine communities that have been previously overlooked.
- Francisco Martinez-Hernandez
- , Oscar Fornas
- & Manuel Martinez-Garcia
-
Article
| Open Access Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
In single-cell RNA sequencing data of heterogeneous cell populations, cell cycle stage of individual cells would often be informative. Here, the authors introduce a computational model to reconstruct a pseudo-time series from single cell transcriptome data, identify the cell cycle stages, identify candidate cell cycle-regulated genes and recover the methylome changes during the cell cycle.
- Zehua Liu
- , Huazhe Lou
- & Ting Chen
-
Article
| Open Access Genetic diagnosis of Mendelian disorders via RNA sequencing
Genetic diagnosis of Mendelian disorders via RNA sequencing
Genome sequencing alone fails to provide a genetic diagnosis for many Mendelian disorder patients. Here, the authors utilize RNA sequencing to complement genotyping of patients with a rare mitochondrial disease by detecting aberrant RNA expression, splicing and allele-specific expression.
- Laura S. Kremer
- , Daniel M. Bader
- & Holger Prokisch
-
Article
| Open Access Cis-perturbation of cancer drivers by the HTLV-1/BLV proviruses is an early determinant of leukemogenesis
Cis-perturbation of cancer drivers by the HTLV-1/BLV proviruses is an early determinant of leukemogenesis
Human T-cell leukaemia virus type-1 and bovine leukaemia virus infect T and B lymphocytes and lead to aggressive leukaemia. Here, the authors show these proviruses integrate near cancer drivers perturbing transcription termination or antisense RNA-dependent interaction, suggesting post-transcriptional mechanisms in some cases.
- Nicolas Rosewick
- , Keith Durkin
- & Anne Van den Broeke
-
Article
| Open Access Ultrasensitive and high-efficiency screen of de novo low-frequency mutations by o2n-seq
Ultrasensitive and high-efficiency screen of de novo low-frequency mutations by o2n-seq
Detection ofde novo, low frequency mutations is important for characterising heterogeneous cell populations, such as those found in cancer cell populations. Here the authors present o2n-seq, an ultrasensitive method with highly efficient data usage for detection of rare mutations.
- Kaile Wang
- , Shujuan Lai
- & Jue Ruan
-
Article
| Open Access Whole genome analysis of a schistosomiasis-transmitting freshwater snail
Whole genome analysis of a schistosomiasis-transmitting freshwater snail
Biomphalaria glabrata is a fresh water snail that acts as a host for trematode Schistosoma mansoni that causes intestinal infection in human. This work describes the genome and transcriptome analyses from 12 different tissues of B glabrata, and identify genes for snail behavior and evolution.
- Coen M. Adema
- , LaDeana W. Hillier
- & Richard K. Wilson
-
Article
| Open Access BLISS is a versatile and quantitative method for genome-wide profiling of DNA double-strand breaks
BLISS is a versatile and quantitative method for genome-wide profiling of DNA double-strand breaks
Double-strand breaks are a major DNA lesion that can occur by endogenous and exogenous processes. Here the authors present BLISS—Breaks LabellingIn Situand Sequencing—to map breaks across the genome.
- Winston X. Yan
- , Reza Mirzazadeh
- & Nicola Crosetto

 Single-cell analysis reveals that stochasticity and paracrine signaling control interferon-alpha production by plasmacytoid dendritic cells
Single-cell analysis reveals that stochasticity and paracrine signaling control interferon-alpha production by plasmacytoid dendritic cells