Article
|
Open Access
Featured
-
-
Article
| Open Access FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons
FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons
The RNA binding protein FUS functions in several RNA biosynthetic processes and has been linked to the pathogenesis of amyotrophic lateral sclerosis (ALS). Here the authors show that FUS controls back-splicing reactions leading to circular RNA (circRNA) production in stem cell-derived motor neurons and that ALS-associated FUS mutations affect the biogenesis of circRNAs.
- Lorenzo Errichelli
- , Stefano Dini Modigliani
- & Irene Bozzoni
-
Article
| Open Access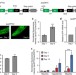 RNA surveillance via nonsense-mediated mRNA decay is crucial for longevity in daf-2/insulin/IGF-1 mutant C. elegans
RNA surveillance via nonsense-mediated mRNA decay is crucial for longevity in daf-2/insulin/IGF-1 mutant C. elegans
The decline of DNA and protein quality control contributes to organismal ageing. Here, Sonet al. report that nonsense-mediated mRNA decay, a RNA quality control mechanism, is enhanced in long-lived daf-2 mutant worms and contributes to their longevity by regulating expression of the yars-2/tyrosyl tRNA synthetase.
- Heehwa G. Son
- , Mihwa Seo
- & Seung-Jae V. Lee
-
Article
| Open Access CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
CLK-dependent exon recognition and conjoined gene formation revealed with a novel small molecule inhibitor
The phosphorylation of serine/arginine-rich proteins by CDC-like kinase is a central regulatory mechanism for RNA splicing reactions. Here, the authors synthesize a novel small molecule CLK inhibitor and map CLK-responsive alternative splicing events and discover an effect on conjoined gene transcription.
- Tyler Funnell
- , Shinya Tasaki
- & Samuel Aparicio
-
Article
| Open Access A CAF40-binding motif facilitates recruitment of the CCR4-NOT complex to mRNAs targeted by Drosophila Roquin
A CAF40-binding motif facilitates recruitment of the CCR4-NOT complex to mRNAs targeted by Drosophila Roquin
Roquin proteins downregulate target mRNA expression by recruiting effectors such as the CCR4-NOT deadenylase complex. Here the authors provide molecular details of how Roquin proteins recruit the CCR4-NOT complex to repress the expression of its targets.
- Annamaria Sgromo
- , Tobias Raisch
- & Elisa Izaurralde
-
Article
| Open Access ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
Nonsense-mediated mRNA decay (NMD) is a quality control pathway that recognizes and degrades transcripts harbouring nonsense mutations. Here the authors show that the ATPase activity of UPF1 mediates functional interactions between the NMD machinery and ribosomes required for efficient ribosome release at premature termination codons.
- Lucas D. Serdar
- , DaJuan L. Whiteside
- & Kristian E. Baker
-
Article
| Open Access Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
mRNA surveillance is essential to maintain homeostasis in eukaryotes and is activated by mRNAs lacking a stop codon. Here the authors describe a high resolution cryo-EM structure of a nonstop complex that shows how arrested ribosome recognition is achieved during Dom34-mediated mRNA surveillance.
- Tarek Hilal
- , Hiroshi Yamamoto
- & Christian M.T. Spahn
-
Article
| Open Access Interrogating the degradation pathways of unstable mRNAs with XRN1-resistant sequences
Interrogating the degradation pathways of unstable mRNAs with XRN1-resistant sequences
Degradation of messenger RNA is a key regulatory step in controlling eukaryotic gene expression. Here the authors present xrFrag, a molecular tool to interrogate the extent and directionality of mRNA turnover by the detection of stabilized decay intermediates produced by several common decay pathways.
- Volker Boehm
- , Jennifer V. Gerbracht
- & Niels H. Gehring
-
Article
| Open Access Structure of the RBM7–ZCCHC8 core of the NEXT complex reveals connections to splicing factors
Structure of the RBM7–ZCCHC8 core of the NEXT complex reveals connections to splicing factors
RBM7 and ZCCHC8 are two core subunits of the Nuclear Exosome Targeting complex, which regulates the degradation of selected non-coding RNAs in human cells. Here, the authors use structural and biochemical methods to show how ZCCHC8 recruits RBM7 in the complex, leaving the RNA binding site accessible and revealing possible implications for splicing.
- Sebastian Falk
- , Ksenia Finogenova
- & Elena Conti
-
Article
| Open Access AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
The chromatin-associated protein AKAP95 is known for its chromatin-related functions including enhancing transcription. Here the authors show that AKAP95 interacts with the splicing regulatory factors as well as RNAs to regulate the inclusion of exons and pre-mRNA splicing.
- Jing Hu
- , Alireza Khodadadi-Jamayran
- & Hao Jiang
-
Article
| Open Access Nop9 is a PUF-like protein that prevents premature cleavage to correctly process pre-18S rRNA
Nop9 is a PUF-like protein that prevents premature cleavage to correctly process pre-18S rRNA
Nop9 is a conserved small ribosomal subunit biogenesis factor. Here, Zhang et al. show that Nop9, in complex with RNA, adopts a C-shaped fold formed from 11 Pumillo repeats and propose that Nop9 inhibits premature cleavage of 20S pre-rRNA by inhibiting the Nob1 nuclease.
- Jun Zhang
- , Kathleen L. McCann
- & Traci M. Tanaka Hall
-
Article
| Open Access Classical non-homologous end-joining pathway utilizes nascent RNA for error-free double-strand break repair of transcribed genes
Classical non-homologous end-joining pathway utilizes nascent RNA for error-free double-strand break repair of transcribed genes
Most adult mammalian cells prefer to repair double-strand DNA breaks though the classical nonhomologous end-joining pathway. Here the authors present evidence that a nascent RNA transcript can serve as a template to facilitate error-free repair.
- Anirban Chakraborty
- , Nisha Tapryal
- & Tapas K. Hazra
-
Article
| Open Access YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex
YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex
The YTHDF family of proteins are able to bind and regulate the stability of methylated N6 RNA. Here the authors show that this decreased m6A RNA stability is mediated by direct recruitment of the CCR4–NOT deadenylase complex through YTHDF proteins.
- Hao Du
- , Ya Zhao
- & Ligang Wu
-
Article
| Open Access Hyperphosphorylation amplifies UPF1 activity to resolve stalls in nonsense-mediated mRNA decay
Hyperphosphorylation amplifies UPF1 activity to resolve stalls in nonsense-mediated mRNA decay
Gene expression is regulated by a range of mechanisms, including post-translational modifications such as phosphorylation. Here the authors present evidence for a feedback mechanism whereby hyperphosphorylation of UPF1 in response to delays in nonsense-mediated decay enhances recruitment of mRNA decay machinery.
- Sébastien Durand
- , Tobias M. Franks
- & Jens Lykke-Andersen
-
Article
| Open Access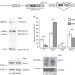 Sec16 alternative splicing dynamically controls COPII transport efficiency
Sec16 alternative splicing dynamically controls COPII transport efficiency
The transport of secretory proteins from the endoplasmic reticulum to the Golgi depends on COPII-coated vesicles. Here, the authors show that activation-induced alternative splicing of Sec16 controls adaptation of COPII transport to increased secretory cargo upon T cell activation.
- Ilka Wilhelmi
- , Regina Kanski
- & Florian Heyd
-
Article
| Open Access The ATPase hCINAP regulates 18S rRNA processing and is essential for embryogenesis and tumour growth
The ATPase hCINAP regulates 18S rRNA processing and is essential for embryogenesis and tumour growth
Perturbations in ribosome biogenesis affect development and increase cancer susceptibility. Here, the authors show that hCINAP is required for 18S rRNA processing, is highly expressed in cancers, and promotes cancer cell growth by upregulating the translation of cancer-associated genes.
- Dongmei Bai
- , Jinfang Zhang
- & Xiaofeng Zheng
-
Article
| Open Access RNA editing generates cellular subsets with diverse sequence within populations
RNA editing generates cellular subsets with diverse sequence within populations
RNA editing rate detected from bulk RNA-seq data can vary widely. Here, by constructing a hierarchical Bayesian model, the authors report substantial variance in editing signatures detected by RNA-seq data from both single cells and a cognate bulk sample.
- Dewi Harjanto
- , Theodore Papamarkou
- & Anastasia Papavasiliou
-
Article
| Open Access Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses
Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses
Abnormal functions of RNA-binding proteins TAF15, FUS and TDP43 are associated with amyotrophic lateral sclerosis. Here, Kapeli et al. characterize the RNA targets of TAF15 and identify points of convergence and divergence between the targets of TAF15, FUS and TDP43 in several neuronal systems.
- Katannya Kapeli
- , Gabriel A. Pratt
- & Gene W. Yeo
-
Article
| Open Access A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
The assembly of the splicesome involves several distinct stages that require the sequential action of DExD/H-box RNA helicases. Here, the authors uncover a new intermediate, the pre-B complex, that accumulates in the presence of an inactive form of the DEAD-box protein Prp28.
- Carsten Boesler
- , Norbert Rigo
- & Reinhard Lührmann
-
Article
| Open Access Comprehensive identification of internal structure and alternative splicing events in circular RNAs
Comprehensive identification of internal structure and alternative splicing events in circular RNAs
Circular RNAs are increasingly understood to have important biological roles and have several subclasses. Here, the authors develop CIRI-AS to analyse sequencing data, identifying the prevalence of alternative splicing and circular RNA isoforms.
- Yuan Gao
- , Jinfeng Wang
- & Fangqing Zhao
-
Article
| Open Access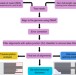 A survey of the sorghum transcriptome using single-molecule long reads
A survey of the sorghum transcriptome using single-molecule long reads
Alternative splicing and alternative polyadenylation (APA) contribute to mRNA diversity but are difficult to assess using short read RNA-seq data. Here, the authors use single molecule long-read isoform sequencing and develop a computational pipeline to identify full-length splice isoforms and APA sites in sorghum.
- Salah E. Abdel-Ghany
- , Michael Hamilton
- & Anireddy S. N. Reddy
-
Article
| Open Access Visualizing the formation of an RNA folding intermediate through a fast highly modular secondary structure switch
Visualizing the formation of an RNA folding intermediate through a fast highly modular secondary structure switch
Short-lived RNA folding intermediates have important roles in the folding of RNA. Here, the authors combine 15N relaxation dispersion NMR with chemical probing to visualise one of these intermediates, and are able to show it is a secondary structural switch, that might help with folding.
- Yi Xue
- , Brant Gracia
- & Hashim M. Al-Hashimi
-
Article
| Open Access A splicing isoform of TEAD4 attenuates the Hippo–YAP signalling to inhibit tumour proliferation
A splicing isoform of TEAD4 attenuates the Hippo–YAP signalling to inhibit tumour proliferation
The Hippo/Yap signalling pathway is found deregulated in several cancers. Here, the authors uncover an additional mechanism of YAP regulation that occurs via alternately spliced isoform of TEAD4, which acts as a dominant negative regulator of YAP-TEAD signalling.
- Yangfan Qi
- , Jing Yu
- & Zefeng Wang
-
Article
| Open Access Splicing factors control C. elegans behavioural learning in a single neuron by producing DAF-2c receptor
Splicing factors control C. elegans behavioural learning in a single neuron by producing DAF-2c receptor
Little is known about the molecular mechanisms regulating neuron-specific alternative splicing. Here, the authors identify a combination of RNA-binding proteins regulating neuron-specific expression of the C. elegansinsulin receptor isoform DAF-2c and find disrupting these factors leads to learning deficits.
- Masahiro Tomioka
- , Yasuki Naito
- & Yuichi Iino
-
Article
| Open Access The complete local genotype–phenotype landscape for the alternative splicing of a human exon
The complete local genotype–phenotype landscape for the alternative splicing of a human exon
Genotype–phenotype landscapes are an important characteristic for understanding the evolution of traits. Here the authors construct the local landscape for the alternative splicing of FAS/CD95 exon 6, revealing the regulation of splicing and the evolution of regulatory information between species.
- Philippe Julien
- , Belén Miñana
- & Ben Lehner
-
Article
| Open Access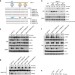 Alternative splicing of MALT1 controls signalling and activation of CD4+ T cells
Alternative splicing of MALT1 controls signalling and activation of CD4+ T cells
MALT1 regulates NFκB signalling both as a scaffolding protein and as a protease. Here the authors show that during T cell activation the expression of MALT1 gene switches to an alternatively spliced variant, which increases TCR signal transduction due to enhanced TRAF6 binding.
- Isabel Meininger
- , Richard A. Griesbach
- & Daniel Krappmann
-
Article
| Open Access Splicing misregulation of SCN5A contributes to cardiac-conduction delay and heart arrhythmia in myotonic dystrophy
Splicing misregulation of SCN5A contributes to cardiac-conduction delay and heart arrhythmia in myotonic dystrophy
Patients with myotonic dystrophy (MD) suffer from severe cardiac issues of unknown aetiology. Freyermuth et al. show that fatal changes in cardiac electrophysiological properties in humans and mice with MD may arise from misregulation of the alternative splicing of the cardiac Na+ channel SCN5Atranscript, resulting in expression of its fetal form.
- Fernande Freyermuth
- , Frédérique Rau
- & Nicolas Charlet-Berguerand
-
Article
| Open Access Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Post-transcriptional control of RNA is important in health and disease. Here, the authors show that the RNA-binding protein Quaking guides pre-mRNA splicing and transcript abundance during monocyte to macrophage differentiation, and that Quaking depletion impairs pro-atherogenic foam cell formation.
- Ruben G. de Bruin
- , Lily Shiue
- & Eric P. van der Veer
-
Article
| Open Access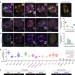 Cajal bodies are linked to genome conformation
Cajal bodies are linked to genome conformation
Nuclear bodies can nucleate at sites of active transcription and are beneficial for efficient gene expression. Here, the authors show that Cajal bodies, a prominent type of nuclear body, contribute to genome organization with global effects on gene expression and RNA splicing fidelity.
- Qiuyan Wang
- , Iain A. Sawyer
- & Miroslav Dundr
-
Article
| Open Access Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage
Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage
Mutations in the splicing factor SF3B1 are found in uveal melanoma. Here, Alsafadi et al. use RNA-sequencing data from these cancers and experimental models, and show that mutant SF3B1 promotes alternative branchpoints in a specific gene subset and that the mutant protein gains a new function.
- Samar Alsafadi
- , Alexandre Houy
- & Marc-Henri Stern
-
Article
| Open Access The RNA helicase DHX34 functions as a scaffold for SMG1-mediated UPF1 phosphorylation
The RNA helicase DHX34 functions as a scaffold for SMG1-mediated UPF1 phosphorylation
UPF1 is a central Nonsense-mediated mRNA decay—(NMD), a mechanism to degrade mRNAs containing premature translation termination codons-factor—whose phosphorylation is key to triggering NMD. Here the authors show that the DHX34 helicase acts as a scaffold in promoting UPF1 phosphorylation by SMG1 to promotes NMD.
- Roberto Melero
- , Nele Hug
- & Oscar Llorca
-
Article
| Open Access ADAR-mediated RNA editing suppresses sleep by acting as a brake on glutamatergic synaptic plasticity
ADAR-mediated RNA editing suppresses sleep by acting as a brake on glutamatergic synaptic plasticity
Sleep is postulated to offset buildup in net synaptic strength that occurs during waking experience. Here, the authors identify a role for the RNA editing gene Adar in regulating glutamatergic synaptic plasticity and show that disruption in Adarexpression impairs normal waking in flies.
- J. E. Robinson
- , J. Paluch
- & W. J. Joiner
-
Article
| Open Access Structural basis of RNA recognition and dimerization by the STAR proteins T-STAR and Sam68
Structural basis of RNA recognition and dimerization by the STAR proteins T-STAR and Sam68
Sam68 and T-STAR are members of the STAR family of proteins, which regulate various aspects of RNA metabolism. Here, the authors reveal structural features required for alternative splicing regulation by these proteins.
- Mikael Feracci
- , Jaelle N. Foot
- & Cyril Dominguez
-
Article
| Open Access Activation-induced deoxycytidine deaminase (AID) co-transcriptional scanning at single-molecule resolution
Activation-induced deoxycytidine deaminase (AID) co-transcriptional scanning at single-molecule resolution
Activation-induced deoxycytidine deaminase (AID) induces somatic hypermutation and class-switch recombination during transcription of immunoglobulin genes. Here the authors use single-molecule FRET to show that AID translocates together with RNA polymerase and scans within stalled transcription bubbles.
- Gayan Senavirathne
- , Jeffrey G. Bertram
- & David Rueda
-
Article
| Open Access Alternative splicing of Drosophila Nmnat functions as a switch to enhance neuroprotection under stress
Alternative splicing of Drosophila Nmnat functions as a switch to enhance neuroprotection under stress
Nicotinamide mononucleotide adenylyltransferase (NMNAT) acts in the NAD biosynthesis pathway and has neuroprotective activity. Ruan et al. show that the neuroprotective activity of NMNAT is restricted to a splice variant of the enzyme, and that this variant is preferentially spliced in response to stress.
- Kai Ruan
- , Yi Zhu
- & R. Grace Zhai
-
Article
| Open Access ESRP2 controls an adult splicing programme in hepatocytes to support postnatal liver maturation
ESRP2 controls an adult splicing programme in hepatocytes to support postnatal liver maturation
Alternative RNA splicing is important during organismal development. Here, the authors perform RNA-Seq on mouse and human liver samples to provide a comprehensive view of splicing events during liver development and growth, and identify Espr2 as a main regulator of these splicing processes.
- Amruta Bhate
- , Darren J. Parker
- & Auinash Kalsotra
-
Article
| Open Access Compound heterozygous mutations in the noncoding RNU4ATAC cause Roifman Syndrome by disrupting minor intron splicing
Compound heterozygous mutations in the noncoding RNU4ATAC cause Roifman Syndrome by disrupting minor intron splicing
Roifman Syndrome is a rare disorder whose disease manifestations include growth retardation, spondyloepiphyseal dysplasia and immunodeficiency. Here, the authors use whole-genome sequencing to discover that rare compound heterozygous variants disrupting the small nuclear RNA gene RNU4ATACcause Roifman Syndrome.
- Daniele Merico
- , Maian Roifman
- & Stephen W. Scherer
-
Article
| Open Access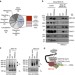 A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C
A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C
mRNA deadenylation, the first step in regulated degradation, is mediated by the action of the CCR4-NOT and PAN2-PAN3 complexes. Here the authors show that the RNA-binding E3 ubiquitin-ligase MEX-3C associates with the CCR4-NOT complex and ubiquitinates the catalytic subunit CNOT7 to regulate its deadenylation activity.
- Florencia Cano
- , Radu Rapiteanu
- & Paul J. Lehner
-
Article
| Open Access The alternative splicing factor Nova2 regulates vascular development and lumen formation
The alternative splicing factor Nova2 regulates vascular development and lumen formation
The alternative splicing factor Nova2 is best known for its pivotal function in the brain. Giampietro et al. reveal an important role for Nova2 in the regulation of alternative splicing of transcripts in the vascular endothelium that are crucial for the maintenance of endothelial cell polarity and vessel lumen formation in zebrafish.
- Costanza Giampietro
- , Gianluca Deflorian
- & Claudia Ghigna
-
Article
| Open Access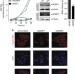 mRNA export through an additional cap-binding complex consisting of NCBP1 and NCBP3
mRNA export through an additional cap-binding complex consisting of NCBP1 and NCBP3
The processing of RNAs transcribed by RNA polymerase II requires a cap-binding complex (CBC), consisting of NCBP1 and NCBP2. Here, the authors report an alternative CBC formed by NCBP1 and a previously uncharacterized protein, NCBP3 that is critical for RNA processing under cellular stress conditions.
- Anna Gebhardt
- , Matthias Habjan
- & Andreas Pichlmair
-
Article
| Open Access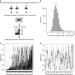 Genetic mapping uncovers cis-regulatory landscape of RNA editing
Genetic mapping uncovers cis-regulatory landscape of RNA editing
Adenosine-to-inosine (A-to-I) RNA editing plays an important role in neurological functions. Here, by a quantitative trait loci (QTL) mapping approach in 131 Drosophila melanogasterstrains, the authors identify 545 QTLs associated with differences in RNA editing.
- Gokul Ramaswami
- , Patricia Deng
- & Jin Billy Li
-
Article |
 Efficient AID targeting of switch regions is not sufficient for optimal class switch recombination
Efficient AID targeting of switch regions is not sufficient for optimal class switch recombination
Activation-induced cytidine deaminase can induce both somatic hypermutation (SHM) and class switch recombination in immunoglobulin loci. Here the authors show that spliced and repetitive switch regions are exquisite SHM targets, but are not sufficient by themselves for class switch recombination.
- Amélie Bonaud
- , Fabien Lechouane
- & Christophe Sirac
-
Article
| Open Access Human Upf1 is a highly processive RNA helicase and translocase with RNP remodelling activities
Human Upf1 is a highly processive RNA helicase and translocase with RNP remodelling activities
Upf1 is a multifunctional helicase involved in various DNA- and RNA-related processes, including nonsense-mediated mRNA decay (NMD). Here the authors demonstrate that Upf1 is a highly processive ribonucleoprotein complex remodeler—a capability likely important for Upf1’s NMD function.
- Francesca Fiorini
- , Debjani Bagchi
- & Vincent Croquette
-
Article
| Open Access Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Eukaryotic ribosomal proteins contain nuclear localization signals (NLSs) that their bacterial counterparts lack. Here the authors compare homologous proteins from bacterial and eukaryotic ribosomes to show how NLSs could emerge in the course of evolution, and use this knowledge to identify novel NLSs.
- Sergey Melnikov
- , Adam Ben-Shem
- & Marat Yusupov
-
Article
| Open Access Widespread disruption of host transcription termination in HSV-1 infection
Widespread disruption of host transcription termination in HSV-1 infection
Herpes simplex virus 1 (HSV-1) efficiently shuts down host gene expression in infected cells. Here Rutkowski et al. analyse the genome-wide changes in transcription and translation in infected cells, and show that HSV-1 triggers an extensive disruption of transcription termination of cellular genes.
- Andrzej J. Rutkowski
- , Florian Erhard
- & Lars Dölken
-
Article |
 Inhibition of vemurafenib-resistant melanoma by interference with pre-mRNA splicing
Inhibition of vemurafenib-resistant melanoma by interference with pre-mRNA splicing
BRAF inhibitors have shown encouraging clinical effects in melanoma patients; however, patients rapidly develop resistance via different mechanisms including alternative splicing. Here the authors find a specific mutation affecting BRAF splicing and highlight the therapeutic potential of splicing interference.
- Maayan Salton
- , Wojciech K. Kasprzak
- & Tom Misteli
-
Article
| Open Access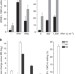 APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages
APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages
Aberrant RNA editing is linked to a range of neuropsychiatric and chronic diseases. Here Sharma et al. show that APOBEC3A can function as an RNA editing protein in response to physiological stimuli, significantly expanding our understanding of RNA editing and the role this may play in diseases.
- Shraddha Sharma
- , Santosh K. Patnaik
- & Bora E. Baysal
-
Article |
 Alternative splicing modulates Kv channel clustering through a molecular ball and chain mechanism
Alternative splicing modulates Kv channel clustering through a molecular ball and chain mechanism
Ion channel clustering at specific membrane sites plays a fundamental role in action potential transmission. Here, the authors show that alternative splice variants of the intrinsically disordered C-terminal segment of the Shaker Kv channel support distinct patterns of scaffold protein-mediated channel clustering.
- Nitzan Zandany
- , Shir Marciano
- & Ofer Yifrach
-
Article |
 Attenuation of nonsense-mediated mRNA decay facilitates the response to chemotherapeutics
Attenuation of nonsense-mediated mRNA decay facilitates the response to chemotherapeutics
Nonsense-mediated mRNA decay (NMD) is a pathway that controls endogenous transcript levels and limits the production of aberrant mRNAs. Here the authors show that NMD is attenuated in cells treated with chemotherapeutic compounds through caspase-mediated proteolytic cleavage of UPF1, a key NMD effector.
- Maximilian W. Popp
- & Lynne E. Maquat
-
Article |
 Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways
Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways
ADAR1 is an adenosine deaminase that converts adenosine to inosine (A-to-I) mostly on Alu repeats in human RNA. Here by analysing transcriptome-wide ADAR1–RNA interactions, the authors show that ADAR1 also binds non-Alusequences to regulate alternative 3′ UTR usage and miRNA biogenesis in the nucleus.
- Jae Hoon Bahn
- , Jaegyoon Ahn
- & Xinshu Xiao

 Programming mRNA decay to modulate synthetic circuit resource allocation
Programming mRNA decay to modulate synthetic circuit resource allocation