Featured
-
-
Article
| Open Access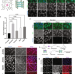 Stress promotes RNA G-quadruplex folding in human cells
Stress promotes RNA G-quadruplex folding in human cells
rG4s are G-quadruplex structures found in transcriptome. Many RNAs can fold into rG4s in vitro, yet their folding and functions in vivo are not well understood. Here the authors showed that rG4s folding is dynamically regulated under stress in human cells.
- Prakash Kharel
- , Marta Fay
- & Pavel Ivanov
-
Article
| Open Access Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
The authors develop an RNA sequencing-based platform, PERSIST-seq, to simultaneously delineate in-cell mRNA stability, ribosome load, and in-solution stability of a diverse mRNA library to derive design principles for improved mRNA therapeutics.
- Kathrin Leppek
- , Gun Woo Byeon
- & Rhiju Das
-
Article
| Open Access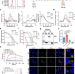 RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
Understanding the mechanisms of SARS-CoV-2 infection is important to control the pandemic. Here the authors show the biological and pathological role of RNA G-quadruplex structure in both SARS-CoV-2 genome and host factors, particularly TMPRSS2.
- Geng Liu
- , Wenya Du
- & Xianghui Fu
-
Article
| Open Access The RNA helicase Dbp7 promotes domain V/VI compaction and stabilization of inter-domain interactions during early 60S assembly
The RNA helicase Dbp7 promotes domain V/VI compaction and stabilization of inter-domain interactions during early 60S assembly
Early steps of large 60S ribosomal subunit biogenesis are not well understood. Here, the authors combine biochemical experiments with protein-RNA crosslinking and mass spectrometry to show that the RNA helicase Dbp7 is key player during early 60S ribosomal assembly. Dbp7 regulates a series of events driving compaction of domain V/VI in early pre60S ribosomal particles.
- Gerald Ryan R. Aquino
- , Philipp Hackert
- & Markus T. Bohnsack
-
Article
| Open Access Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
The molecular events underlying the assembly and maturation of the early pre-60S particles during eukaryotic ribosome synthesis are not well understood. Here, the authors combine yeast genetics and biochemical experiments to characterise the functions of two important players of eukaryotic ribosome biogenesis, the box C/D snoRNP snR190 and the helicase Dbp7, which both interact. They show that the snR190 snoRNA acts as a RNA chaperone that assists the structuring of the 25S rRNA during the maturation of early pre-60S particles and that Dbp7 is important for facilitating remodeling events in the peptidyl transferase center region of the 25S rRNAs during the maturation of early pre-60S particles.
- Mariam Jaafar
- , Julia Contreras
- & Anthony K. Henras
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Rapid and accurate determination of atomistic RNA dynamic ensemble models using NMR and structure prediction
Rapid and accurate determination of atomistic RNA dynamic ensemble models using NMR and structure prediction
Determining dynamic ensembles of biomolecules is still challenging. Here the authors present an approach for rapid RNA ensemble determination that combines RNA structure prediction tools and NMR residual dipolar coupling data and use it to determine atomistic ensemble models for a variety of RNAs.
- Honglue Shi
- , Atul Rangadurai
- & Hashim M. Al-Hashimi
-
Article
| Open Access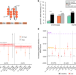 Integrative analysis reveals RNA G-quadruplexes in UTRs are selectively constrained and enriched for functional associations
Integrative analysis reveals RNA G-quadruplexes in UTRs are selectively constrained and enriched for functional associations
G-quadruplexes (G4s) are secondary structures that can form in both DNA and RNA from guanine-rich sequences which are enriched in untranslated regions (UTRs). Here, Lee et al. find that putative G4-forming sequences are evolutionarily constrained, enriched for RNA-binding protein interactions and enriched for disease genetic associations.
- David S. M. Lee
- , Louis R. Ghanem
- & Yoseph Barash
-
Article
| Open Access Structure mapping of dengue and Zika viruses reveals functional long-range interactions
Structure mapping of dengue and Zika viruses reveals functional long-range interactions
Here, the authors provide detailed analyses of viral RNA structure in virions and in infected cells for four dengue virus serotypes and four Zika virus strains, and identify conserved structures that are important for virus replication.
- Roland G. Huber
- , Xin Ni Lim
- & Yue Wan
-
Article
| Open Access RNA helicases mediate structural transitions and compositional changes in pre-ribosomal complexes
RNA helicases mediate structural transitions and compositional changes in pre-ribosomal complexes
Pre-ribosomes undergo numerous structural rearrangements during their assembly. Here the authors identify the binding sites of three essential RNA helicases on pre-ribosomal particles, enabling them to provide insights into the structural and compositional changes that occur during biogenesis of the large ribosomal subunit.
- Lukas Brüning
- , Philipp Hackert
- & Markus T. Bohnsack
-
Article
| Open Access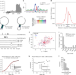 Roquin targets mRNAs in a 3′-UTR-specific manner by different modes of regulation
Roquin targets mRNAs in a 3′-UTR-specific manner by different modes of regulation
Roquin targets are known to contain two types of sequence-structure motifs, the constitutive and the alternative decay elements (CDE and ADE). Here, the authors describe a linear Roquin binding element (LBE) also involved in target recognition, and show that Roquin binding affects the translation of a subset of targeted mRNAs.
- Katharina Essig
- , Nina Kronbeck
- & Vigo Heissmeyer
-
Article
| Open Access LRPPRC-mediated folding of the mitochondrial transcriptome
LRPPRC-mediated folding of the mitochondrial transcriptome
The mitochondrial genome, being compressed to 16 kb, is an attractive model system to investigate how RNA-binding proteins chaperone mRNA lifecycles. Here the authors use RNase footprinting and PAR-CLIP to show that the LRPPRC–SLIRP complex stabilizes mRNA structures to expose sites required for translation and polyadenylation.
- Stefan J. Siira
- , Henrik Spåhr
- & Aleksandra Filipovska
-
Article
| Open Access High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
Ribosome biogenesis is a dynamic process that involves the ordered assembly of ribosomal proteins and numerous RNA structural rearrangements. Here the authors apply ChemModSeq, a high-throughput RNA structure probing method, to quantitatively measure changes in RNA flexibility during the nucleolar stages of 60S assembly in yeast.
- Elena Burlacu
- , Fredrik Lackmann
- & Sander Granneman
-
Article
| Open Access Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes
Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes
Ribosomes assemble through the hierarchical addition of proteins to a ribosomal RNA scaffold. Here the authors use three-color single-molecule FRET to show how the dynamics of the rRNA dictate the order in which multiple proteins assemble on the 5′ domain of the E. coli 16S rRNA.
- Sanjaya C. Abeysirigunawardena
- , Hajin Kim
- & Sarah A. Woodson
-
Article
| Open Access Visualizing the formation of an RNA folding intermediate through a fast highly modular secondary structure switch
Visualizing the formation of an RNA folding intermediate through a fast highly modular secondary structure switch
Short-lived RNA folding intermediates have important roles in the folding of RNA. Here, the authors combine 15N relaxation dispersion NMR with chemical probing to visualise one of these intermediates, and are able to show it is a secondary structural switch, that might help with folding.
- Yi Xue
- , Brant Gracia
- & Hashim M. Al-Hashimi
-
Article
| Open Access Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Eukaryotic ribosomal proteins contain nuclear localization signals (NLSs) that their bacterial counterparts lack. Here the authors compare homologous proteins from bacterial and eukaryotic ribosomes to show how NLSs could emerge in the course of evolution, and use this knowledge to identify novel NLSs.
- Sergey Melnikov
- , Adam Ben-Shem
- & Marat Yusupov

 Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect