Featured
-
-
Article
| Open Access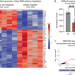 Modulation of insulin secretion by RBFOX2-mediated alternative splicing
Modulation of insulin secretion by RBFOX2-mediated alternative splicing
Insulin secretion from the pancreatic beta cell is a tightly regulated process that is vital for maintaining blood glucose homeostasis. Here, the authors show that the RNA binding protein RBFOX2 is a regulator of insulin secretion through the alternative splicing of genes required for insulin granule docking and exocytosis.
- Nicole D. Moss
- , Kristen L. Wells
- & Lori Sussel
-
Article
| Open Access Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Dawicki-McKenna and Felix et al comprehensively map binding and alternative splicing by PTBP2 in human brain and neurons, thus identifying splice switching therapeutic strategies for the neurodevelopmental disorder associated gene SYNGAP1.
- Jennine M. Dawicki-McKenna
- , Alex J. Felix
- & Benjamin L. Prosser
-
Comment
| Open Access The problem of selection bias in studies of pre-mRNA splicing
The problem of selection bias in studies of pre-mRNA splicing
In this comment, the authors discuss the potentially widespread problem of selection bias in drawing biological conclusions from RNA sequencing data.
- Zachary W. Dwyer
- & Jeffrey A. Pleiss
-
Article
| Open Access The m6A reader YTHDC1 and the RNA helicase DDX5 control the production of rhabdomyosarcoma-enriched circRNAs
The m6A reader YTHDC1 and the RNA helicase DDX5 control the production of rhabdomyosarcoma-enriched circRNAs
Rhabdomyosarcoma (RMS) is the most diffused soft tissue sarcoma in children and adolescents. Herein, the authors identify the m6A machinery and the RNA helicase DDX5 as factors responsible for the increase of a subset of circRNAs in RMS, providing protein and RNA candidates for the study of its tumorigenicity.
- Dario Dattilo
- , Gaia Di Timoteo
- & Irene Bozzoni
-
Article
| Open Access Endothelial deletion of PTBP1 disrupts ventricular chamber development
Endothelial deletion of PTBP1 disrupts ventricular chamber development
Alternative splicing crucially affects various biological processes, however, its function in heart development is largely unknown. Here, the authors show an essential role of alternative splicing factor PTBP1 in ventricular chamber development.
- Hongyu Liu
- , Ran Duan
- & Yi-Han Chen
-
Article
| Open Access Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Analysing the regulatory consequences of mutations and splice variants at large scale in cancer requires efficient computational tools. Here, the authors develop RegTools, a software package that can identify splice-associated variants from large-scale genomics and transcriptomics data with efficiency and flexibility.
- Kelsy C. Cotto
- , Yang-Yang Feng
- & Malachi Griffith
-
Article
| Open Access Splicing factor SRSF1 deficiency in the liver triggers NASH-like pathology and cell death
Splicing factor SRSF1 deficiency in the liver triggers NASH-like pathology and cell death
Nonalcoholic steatohepatitis (NASH) is an advanced form of fatty liver disease with complex pathogenic mechanisms. Here, the authors report that SRSF1 deficiency in mice livers provokes deleterious R-loop formation and genotoxicity, which impedes hepatocellular gene expression, metabolism, and lipid trafficking, resulting in NASH-like pathology.
- Waqar Arif
- , Bhoomika Mathur
- & Auinash Kalsotra
-
Article
| Open Access Cryo-EM structure of hnRNPDL-2 fibrils, a functional amyloid associated with limb-girdle muscular dystrophy D3
Cryo-EM structure of hnRNPDL-2 fibrils, a functional amyloid associated with limb-girdle muscular dystrophy D3
The authors report the Cryo-EM of hnRNPDL-2 fibrils. The structure highlights features of a functional amyloid associated with limb-girdle muscular dystrophy-3 and explains how alternative splicing controls the assembly of this ribonucleoprotein.
- Javier Garcia-Pardo
- , Andrea Bartolomé-Nafría
- & Salvador Ventura
-
Article
| Open Access High-throughput mutagenesis identifies mutations and RNA-binding proteins controlling CD19 splicing and CART-19 therapy resistance
High-throughput mutagenesis identifies mutations and RNA-binding proteins controlling CD19 splicing and CART-19 therapy resistance
Multiple alternative splicing events in CD19 mRNA have been associated with resistance/relapse to CD19 CAR-T therapy in patients with B cell malignancies. Here, by combining patient data and a high-throughput mutagenesis screen, the authors identify single point mutations and RNA-binding proteins that can control CD19 splicing and be associated with CD19 CAR-T therapy resistance.
- Mariela Cortés-López
- , Laura Schulz
- & Julian König
-
Article
| Open Access Heterogeneous nuclear ribonucleoprotein U (HNRNPU) safeguards the developing mouse cortex
Heterogeneous nuclear ribonucleoprotein U (HNRNPU) safeguards the developing mouse cortex
HNRNPU is an RNA splicing protein associated with brain disorders such as early onset seizures. Here they show that HNRNPU functions to maintain neural progenitors and their progeny by regulating splicing of key neuronal genes.
- Tamar Sapir
- , Aditya Kshirsagar
- & Orly Reiner
-
Article
| Open Access hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells
hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells
Coordinated regulation of alternative splicing is essential for germ cell development. Here, the authors report that hnRNPH1 interacts with alternative splicing factors PTBP2 and SRSF3 in the germline to regulate pre-mRNA alternative splicing.
- Shenglei Feng
- , Jinmei Li
- & Shuiqiao Yuan
-
Perspective
| Open Access Autophagy regulation by RNA alternative splicing and implications in human diseases
Autophagy regulation by RNA alternative splicing and implications in human diseases
Both alternative splicing and autophagy are core cell biological processes, but where they intersect has received little attention. Here, the authors reflect on recent connections identified between these pathways and consider their impact on human disease.
- Patricia González-Rodríguez
- , Daniel J. Klionsky
- & Bertrand Joseph
-
Article
| Open Access A multi-factor trafficking site on the spliceosome remodeling enzyme BRR2 recruits C9ORF78 to regulate alternative splicing
A multi-factor trafficking site on the spliceosome remodeling enzyme BRR2 recruits C9ORF78 to regulate alternative splicing
Bergfort et al. use biochemistry, cryoEM, structure-guided mutagenesis, transcriptomics and proteomics to reveal how the intrinsically unstructured C9ORF78 protein binds BRR2 and PRPF8, regulating cassette exons and alternative 3’-splice sites.
- Alexandra Bergfort
- , Marco Preußner
- & Markus C. Wahl
-
Article
| Open Access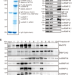 Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
MeCP2 mutations can cause Rett syndrome, a severe childhood neurological disorder. Here the authors show that MeCP2 mediates the higher-order assembly of a large splicing complex Rbfox/LASR, which is disrupted in the mouse models of Rett syndrome.
- Yan Jiang
- , Xing Fu
- & Jingyi Hui
-
Article
| Open Access A ligand-insensitive UNC5B splicing isoform regulates angiogenesis by promoting apoptosis
A ligand-insensitive UNC5B splicing isoform regulates angiogenesis by promoting apoptosis
UNC5B is a Netrin-1 receptor expressed in endothelial cells that in the absence of ligand induces apoptosis. Here the authors identify an UNC5B splicing isoform that is insensitive to the pro-survival ligand Netrin-1 and is required for apoptosis-dependent blood vessel development.
- Davide Pradella
- , Gianluca Deflorian
- & Claudia Ghigna
-
Article
| Open Access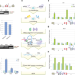 The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
We know that most splicing reactions take place co-transcriptionally, but how the transcription machinery facilitate splicing of introns is unknown. Here the authors show that the 5′ splice site remains associated with the transcription machinery during intron synthesis through U1 snRNP, providing a basis for the rapid splicing reaction of introns.
- Yodfat Leader
- , Galit Lev Maor
- & Gil Ast
-
Article
| Open Access Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
The Argonaute protein CSR-1 is essential for fertility and viability in C. elegans. Here the authors show that CSR-1A isoform associates preferentially with small RNAs mapping to spermatogenesis-specific genes while CSR-1B isoform binds small RNAs mapping to oogenesis-specific genes. Arginine methylation of CSR-1A promotes small RNA-binding specificity.
- Dieu An H. Nguyen
- & Carolyn M. Phillips
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Arginine methyltransferases (PRMTs) are involved in the regulation of various physiological and pathological conditions. Using proteomics, the authors here profile the methylation substrates of PRMTs 4, 5 and 7 and characterize the roles of these enzymes in cancer-associated splicing regulation.
- Wen-juan Li
- , Yao-hui He
- & Wen Liu
-
Article
| Open Access A conserved role for the ALS-linked splicing factor SFPQ in repression of pathogenic cryptic last exons
A conserved role for the ALS-linked splicing factor SFPQ in repression of pathogenic cryptic last exons
SFPQ is a splicing factor and its mutations are associated to amyotrophic lateral sclerosis (ALS) patients. Here, the authors show that SFPQ represses the use of pathogenic cryptic last exons in zebrafish, mouse and human cells.
- Patricia M. Gordon
- , Fursham Hamid
- & Corinne Houart
-
Article
| Open Access Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Chromatin is known to regulate splicing by modulating recruitment of splicing factors. Using machine learning approaches, the authors have underlined a chromatin code for alternative splicing regulation that is conserved amongst cell lines.
- E. Agirre
- , A. J. Oldfield
- & R. F. Luco
-
Article
| Open Access isoCirc catalogs full-length circular RNA isoforms in human transcriptomes
isoCirc catalogs full-length circular RNA isoforms in human transcriptomes
Circular RNAs have been identified using short-read RNA sequencing. Here, the authors report isoCirc, a long-read sequencing method to characterize full-length circRNA isoforms and generate a catalogue of full-length circRNA isoforms in 12 human tissues and one human cell line.
- Ruijiao Xin
- , Yan Gao
- & Yi Xing
-
Article
| Open Access CRISPR artificial splicing factors
CRISPR artificial splicing factors
Control over splicing could be used for both therapeutic and engineering applications. Here the authors create artificial splicing factors using RNA-targeting CRISPR systems under small molecule control.
- Menghan Du
- , Nathaniel Jillette
- & Albert Wu Cheng
-
Article
| Open Access Ribosome profiling at isoform level reveals evolutionary conserved impacts of differential splicing on the proteome
Ribosome profiling at isoform level reveals evolutionary conserved impacts of differential splicing on the proteome
Genes express multiple mRNA isoforms through alternative processing. Here the authors analyze ribosome profiling data with ORQAS (ORF quantification pipeline for alternative splicing) and report that 40–50% of the expressed mRNA isoforms are translated.
- Marina Reixachs-Solé
- , Jorge Ruiz-Orera
- & Eduardo Eyras
-
Article
| Open Access Cis- and trans-regulations of pre-mRNA splicing by RNA editing enzymes influence cancer development
Cis- and trans-regulations of pre-mRNA splicing by RNA editing enzymes influence cancer development
RNA editing and RNA splicing are involved in tumorigenesis. Here the authors report crosstalk between RNA editing and splicing by identifying ADAR1/2-dependent splicing events in esophageal squamous carcinoma cells.
- Sze Jing Tang
- , Haoqing Shen
- & Leilei Chen
-
Article
| Open Access Structural basis for adhesion G protein-coupled receptor Gpr126 function
Structural basis for adhesion G protein-coupled receptor Gpr126 function
The extracellular regions (ECRs) of adhesion GPCRs have diverse biological functions, but their structures and mechanisms of action remain unclear. Here, the authors solve the ECR structure of the Gpr126 receptor and show that ECR conformation and signaling functions are regulated by alternative splicing.
- Katherine Leon
- , Rebecca L. Cunningham
- & Demet Araç
-
Article
| Open Access Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Alternative splicing is regulated by multiple mechanisms. Here the authors employed designed splice site libraries and massively parallel reporter assays to dissect the regulatory complexity and cell-to-cell variability of splicing decisions and to build accurate predictive models.
- Martin Mikl
- , Amit Hamburg
- & Eran Segal
-
Article
| Open Access Specific inhibition of splicing factor activity by decoy RNA oligonucleotides
Specific inhibition of splicing factor activity by decoy RNA oligonucleotides
Alternative splicing, critical for gene expression, is deregulated in many diseases. Here the authors develop decoy oligonucleotides to specifically downregulate splicing factors activity.
- Polina Denichenko
- , Maxim Mogilevsky
- & Rotem Karni
-
Article
| Open Access An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
CTCF plays key roles in gene regulation, chromatin insulation and organizing the higher-order chromatin architecture of mammalian genomes. Here the authors investigate the function an alternatively spliced shorter CTCF isoform, finding that this isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis.
- Jiao Li
- , Kaimeng Huang
- & Hongjie Yao
-
Article
| Open Access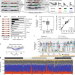 Decoding a cancer-relevant splicing decision in the RON proto-oncogene using high-throughput mutagenesis
Decoding a cancer-relevant splicing decision in the RON proto-oncogene using high-throughput mutagenesis
Alternative splicing is a critical step in eukaryotic gene expression but its molecular rules are not fully understood. Here, the authors develop a high-throughput mutagenesis approach to comprehensively characterise determinants of alternative splicing for the RON proto-oncogene.
- Simon Braun
- , Mihaela Enculescu
- & Kathi Zarnack
-
Article
| Open Access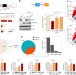 Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Cytoplasmic PTEN is a tumor suppressor that antagonises PI3K signalling. Here, the authors show that nuclear PTEN can interact with the spliceosomal proteins and drive pre-mRNA splicing in a phosphatase-independent manner, in particular, PTEN depletion promotes Golgi extension and secretion through GOLGA2 exon skipping.
- Shao-Ming Shen
- , Yan Ji
- & Guo-Qiang Chen
-
Article
| Open Access Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Alternative splicing of influenza A virus (IAV) M transcript is regulated by hnRNP K and NS1-BP, but mechanistic details are unknown. Here, Thompson et al. show how hnRNP K and NS1-BP bind M mRNA and that these proteins regulate splicing of host transcripts in both the absence and presence of IAV infection.
- Matthew G. Thompson
- , Raquel Muñoz-Moreno
- & Kristen W. Lynch
-
Article
| Open Access Precise temporal regulation of alternative splicing during neural development
Precise temporal regulation of alternative splicing during neural development
The precise timing of neurodevelopmental splicing switches and the underlying regulatory mechanisms remain poorly understood. This study identifies two major waves of developmental switches under the control of distinct combinations of RNA-binding proteins in central and peripheral nervous systems.
- Sebastien M. Weyn-Vanhentenryck
- , Huijuan Feng
- & Chaolin Zhang
-
Article
| Open Access rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
Myotonic dystrophy (DM) type 2 is a neuromuscular pathology caused by large expansions of CCTG repeats. Here the authors find that rbFOX1 RNA binding protein binds to CCUG RNA repeats and competes with MBNL1 for the binding to CCUG repeats, releasing MBNL1 from sequestration in DM2 muscle cells.
- Chantal Sellier
- , Estefanía Cerro-Herreros
- & Nicolas Charlet-Berguerand
-
Article
| Open Access Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention (IR) can increase protein diversity and function, and yet unregulated IR may be detrimental to cellular health. This study shows that aberrant IR occurs in ALS and finds nuclear loss of an RNA-binding protein called SFPQ as a new molecular hallmark in this devastating condition.
- Raphaelle Luisier
- , Giulia E. Tyzack
- & Rickie Patani
-
Article
| Open Access ZFR coordinates crosstalk between RNA decay and transcription in innate immunity
ZFR coordinates crosstalk between RNA decay and transcription in innate immunity
Type I interferon signaling is critical for the control of infection. Here the authors show that zinc finger RNA-binding protein (ZFR) can control type I interferon responses, and that this control is itself regulated by distinct ZFR truncation patterns that differ between monocytes and macrophages.
- Nazmul Haque
- , Ryota Ouda
- & J. Robert Hogg
-
Article
| Open Access Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers
Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers
Small molecules correcting the splicing deficit of the survival of motor neuron 2 (SMN2) gene have been identified as having therapeutic potential. Here, the authors provide evidence that SMN2 mRNA forms a ribonucleoprotein complex that can be specifically targeted by these small molecules.
- Manaswini Sivaramakrishnan
- , Kathleen D. McCarthy
- & Friedrich Metzger
-
Article
| Open Access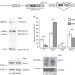 Sec16 alternative splicing dynamically controls COPII transport efficiency
Sec16 alternative splicing dynamically controls COPII transport efficiency
The transport of secretory proteins from the endoplasmic reticulum to the Golgi depends on COPII-coated vesicles. Here, the authors show that activation-induced alternative splicing of Sec16 controls adaptation of COPII transport to increased secretory cargo upon T cell activation.
- Ilka Wilhelmi
- , Regina Kanski
- & Florian Heyd
-
Article
| Open Access A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
The assembly of the splicesome involves several distinct stages that require the sequential action of DExD/H-box RNA helicases. Here, the authors uncover a new intermediate, the pre-B complex, that accumulates in the presence of an inactive form of the DEAD-box protein Prp28.
- Carsten Boesler
- , Norbert Rigo
- & Reinhard Lührmann
-
Article
| Open Access Comprehensive identification of internal structure and alternative splicing events in circular RNAs
Comprehensive identification of internal structure and alternative splicing events in circular RNAs
Circular RNAs are increasingly understood to have important biological roles and have several subclasses. Here, the authors develop CIRI-AS to analyse sequencing data, identifying the prevalence of alternative splicing and circular RNA isoforms.
- Yuan Gao
- , Jinfeng Wang
- & Fangqing Zhao
-
Article
| Open Access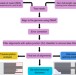 A survey of the sorghum transcriptome using single-molecule long reads
A survey of the sorghum transcriptome using single-molecule long reads
Alternative splicing and alternative polyadenylation (APA) contribute to mRNA diversity but are difficult to assess using short read RNA-seq data. Here, the authors use single molecule long-read isoform sequencing and develop a computational pipeline to identify full-length splice isoforms and APA sites in sorghum.
- Salah E. Abdel-Ghany
- , Michael Hamilton
- & Anireddy S. N. Reddy
-
Article
| Open Access A splicing isoform of TEAD4 attenuates the Hippo–YAP signalling to inhibit tumour proliferation
A splicing isoform of TEAD4 attenuates the Hippo–YAP signalling to inhibit tumour proliferation
The Hippo/Yap signalling pathway is found deregulated in several cancers. Here, the authors uncover an additional mechanism of YAP regulation that occurs via alternately spliced isoform of TEAD4, which acts as a dominant negative regulator of YAP-TEAD signalling.
- Yangfan Qi
- , Jing Yu
- & Zefeng Wang
-
Article
| Open Access Splicing factors control C. elegans behavioural learning in a single neuron by producing DAF-2c receptor
Splicing factors control C. elegans behavioural learning in a single neuron by producing DAF-2c receptor
Little is known about the molecular mechanisms regulating neuron-specific alternative splicing. Here, the authors identify a combination of RNA-binding proteins regulating neuron-specific expression of the C. elegansinsulin receptor isoform DAF-2c and find disrupting these factors leads to learning deficits.
- Masahiro Tomioka
- , Yasuki Naito
- & Yuichi Iino
-
Article
| Open Access The complete local genotype–phenotype landscape for the alternative splicing of a human exon
The complete local genotype–phenotype landscape for the alternative splicing of a human exon
Genotype–phenotype landscapes are an important characteristic for understanding the evolution of traits. Here the authors construct the local landscape for the alternative splicing of FAS/CD95 exon 6, revealing the regulation of splicing and the evolution of regulatory information between species.
- Philippe Julien
- , Belén Miñana
- & Ben Lehner
-
Article
| Open Access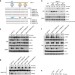 Alternative splicing of MALT1 controls signalling and activation of CD4+ T cells
Alternative splicing of MALT1 controls signalling and activation of CD4+ T cells
MALT1 regulates NFκB signalling both as a scaffolding protein and as a protease. Here the authors show that during T cell activation the expression of MALT1 gene switches to an alternatively spliced variant, which increases TCR signal transduction due to enhanced TRAF6 binding.
- Isabel Meininger
- , Richard A. Griesbach
- & Daniel Krappmann
-
Article
| Open Access Splicing misregulation of SCN5A contributes to cardiac-conduction delay and heart arrhythmia in myotonic dystrophy
Splicing misregulation of SCN5A contributes to cardiac-conduction delay and heart arrhythmia in myotonic dystrophy
Patients with myotonic dystrophy (MD) suffer from severe cardiac issues of unknown aetiology. Freyermuth et al. show that fatal changes in cardiac electrophysiological properties in humans and mice with MD may arise from misregulation of the alternative splicing of the cardiac Na+ channel SCN5Atranscript, resulting in expression of its fetal form.
- Fernande Freyermuth
- , Frédérique Rau
- & Nicolas Charlet-Berguerand
-
Article
| Open Access Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage
Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage
Mutations in the splicing factor SF3B1 are found in uveal melanoma. Here, Alsafadi et al. use RNA-sequencing data from these cancers and experimental models, and show that mutant SF3B1 promotes alternative branchpoints in a specific gene subset and that the mutant protein gains a new function.
- Samar Alsafadi
- , Alexandre Houy
- & Marc-Henri Stern
-
Article
| Open Access Alternative splicing of Drosophila Nmnat functions as a switch to enhance neuroprotection under stress
Alternative splicing of Drosophila Nmnat functions as a switch to enhance neuroprotection under stress
Nicotinamide mononucleotide adenylyltransferase (NMNAT) acts in the NAD biosynthesis pathway and has neuroprotective activity. Ruan et al. show that the neuroprotective activity of NMNAT is restricted to a splice variant of the enzyme, and that this variant is preferentially spliced in response to stress.
- Kai Ruan
- , Yi Zhu
- & R. Grace Zhai
-
Article
| Open Access The alternative splicing factor Nova2 regulates vascular development and lumen formation
The alternative splicing factor Nova2 regulates vascular development and lumen formation
The alternative splicing factor Nova2 is best known for its pivotal function in the brain. Giampietro et al. reveal an important role for Nova2 in the regulation of alternative splicing of transcripts in the vascular endothelium that are crucial for the maintenance of endothelial cell polarity and vessel lumen formation in zebrafish.
- Costanza Giampietro
- , Gianluca Deflorian
- & Claudia Ghigna

 Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing
Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing