Featured
-
-
Article
| Open Access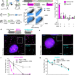 A rapid inducible RNA decay system reveals fast mRNA decay in P-bodies
A rapid inducible RNA decay system reveals fast mRNA decay in P-bodies
Studying RNA decay remains a challenging task. Here, the authors present a technology that enables inducible rapid degradation of targeted mRNAs. Visualizing mRNA decay dynamics unveils insights into P-body function in RNA metabolism.
- Lauren A. Blake
- , Leslie Watkins
- & Bin Wu
-
Article
| Open Access Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Here the authors uncovered CNOT3, a subunit of the CCR4-NOT complex, as an essential modulator of translation in leukemia. The work pointed to the potential of targeting the posttranscriptional circuitry via CNOT3 as a therapeutic vulnerability in acute myeloid leukemia.
- Maryam Ghashghaei
- , Yilin Liu
- & Ly P. Vu
-
Article
| Open Access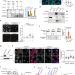 PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
R-loop accumulation at transcription sites poses a persistent threat to genome integrity. Here the authors demonstrate a role for PSIP1/LEDGF protein in reducing R-loop levels at the site of transcription and preventing transcription replication conflict to maintain genome integrity.
- Sundarraj Jayakumar
- , Manthan Patel
- & Madapura M. Pradeepa
-
Article
| Open Access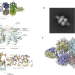 Acetylation regulates the oligomerization state and activity of RNase J, the Helicobacter pylori major ribonuclease
Acetylation regulates the oligomerization state and activity of RNase J, the Helicobacter pylori major ribonuclease
Here the authors find that RNase J, the major ribonuclease of the gastric pathogen Helicobacter pylori is post-translationally modified by acetylation. They show that acetylation can control RNase J activity.
- Alejandro Tejada-Arranz
- , Aleksei Lulla
- & Hilde De Reuse
-
Article
| Open Access ALKBH5 controls the meiosis-coupled mRNA clearance in oocytes by removing the N 6-methyladenosine methylation
ALKBH5 controls the meiosis-coupled mRNA clearance in oocytes by removing the N 6-methyladenosine methylation
N6-methyladenosine (m6A) maintains maternal RNA stability in oocytes. Here, the authors identify demethylase ALKBH5 as a key determinant of oocyte quality and unveil the facilitating role of ALKBH5-mediated m6A removal in maternal RNA decay.
- Long Bai
- , Yu Xiang
- & Yimin Zhu
-
Article
| Open Access Rio1 downregulates centromeric RNA levels to promote the timely assembly of structurally fit kinetochores
Rio1 downregulates centromeric RNA levels to promote the timely assembly of structurally fit kinetochores
Kinetochores assemble on centromeres via histone H3 variant CENP-A and low levels of centromere transcripts (cenRNAs). Here the authors show the Rio1 kinase limits cenRNA production by reducing RNAPII accessibility and promotes cenRNA degradation by the 5’− 3’exoribonuclease Rat1.
- Ksenia Smurova
- , Michela Damizia
- & Peter De Wulf
-
Article
| Open Access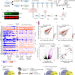 Buffering of transcription rate by mRNA half-life is a conserved feature of Rett syndrome models
Buffering of transcription rate by mRNA half-life is a conserved feature of Rett syndrome models
Rett syndrome (RTT) is a neurodevelopmental disorder caused by mutations in the transcriptional modulator MECP2. Here, the authors measured transcription rate and mRNA half-life changes in RTT patient-derived neurons to show transcription rate buffered by mRNA half-life changes.
- Deivid C. Rodrigues
- , Marat Mufteev
- & James Ellis
-
Article
| Open Access Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation
Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation
Transcriptional memory is key for cellular adaptation. Here the authors show that differences in mRNA stability and mRNA degradation machinery between naïve and primed cells facilitate faster gene expression response to repeated stimuli.
- Bingnan Li
- , Patrice Zeis
- & Vicent Pelechano
-
Article
| Open Access APOBEC3B drives PKR-mediated translation shutdown and protects stress granules in response to viral infection
APOBEC3B drives PKR-mediated translation shutdown and protects stress granules in response to viral infection
APOBEC’s are a family of cytidine deaminases that induce mutations in viruses to inhibit their replication and maintain cell integrity. Here, Manjunath et al show that APOBEC3B also inhibits viral replication by stimulating the innate immune sensor protein kinase R causing translational shutdown and stress granule formation independently of its cytidine deaminase activity.
- Lavanya Manjunath
- , Sunwoo Oh
- & Rémi Buisson
-
Article
| Open Access Mechanistic insights into RNA surveillance by the canonical poly(A) polymerase Pla1 of the MTREC complex
Mechanistic insights into RNA surveillance by the canonical poly(A) polymerase Pla1 of the MTREC complex
Here the authors show how the MTREC core protein Red1 binds to and sequesters Pla1 from the 3’-end processing machinery to hyperadenylate cryptic unstable transcripts and target them to the exosome for efficient degradation.
- Komal Soni
- , Anusree Sivadas
- & Tamás Fischer
-
Article
| Open Access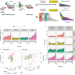 Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Dynamic RNA-protein interactions govern the co-transcriptional packaging of RNA polymerase II derived transcripts. Here the authors use temporal-iCLIP which combines transcriptional synchronisation with UV cross-linking of RNA-protein complexes to reveal dynamic RNA-protein interactions during the early phases of transcription and beyond.
- Ross A. Cordiner
- , Yuhui Dou
- & Torben Heick Jensen
-
Article
| Open Access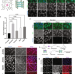 Stress promotes RNA G-quadruplex folding in human cells
Stress promotes RNA G-quadruplex folding in human cells
rG4s are G-quadruplex structures found in transcriptome. Many RNAs can fold into rG4s in vitro, yet their folding and functions in vivo are not well understood. Here the authors showed that rG4s folding is dynamically regulated under stress in human cells.
- Prakash Kharel
- , Marta Fay
- & Pavel Ivanov
-
Article
| Open Access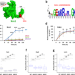 RNA-controlled nucleocytoplasmic shuttling of mRNA decay factors regulates mRNA synthesis and a novel mRNA decay pathway
RNA-controlled nucleocytoplasmic shuttling of mRNA decay factors regulates mRNA synthesis and a novel mRNA decay pathway
Nucleocytoplasmic shuttling of several yeast mRNA decay factors regulates transcription and initiates a novel mRNA decay pathway; shuttling is controlled by the decaying RNA and is critical for coping with environmental changes.
- Shiladitya Chattopadhyay
- , Jose Garcia-Martinez
- & Mordechai Choder
-
Article
| Open Access A nascent peptide code for translational control of mRNA stability in human cells
A nascent peptide code for translational control of mRNA stability in human cells
Here, the authors measure the mRNA levels of thousands of coding sequence motifs in human cells to find that nascent peptides with a combination of β-strand structures and bulky and positively charged sequences reduce mRNA levels by slowing translation.
- Phillip C. Burke
- , Heungwon Park
- & Arvind Rasi Subramaniam
-
Article
| Open Access Cryo-EM reveals the architecture of the PELP1-WDR18 molecular scaffold
Cryo-EM reveals the architecture of the PELP1-WDR18 molecular scaffold
PELP1 is a large scaffolding protein implicated in many cellular activities, including ribosome assembly as part of the Rix1 complex, comprising PELP1, WDR18, TEX10 and other components. Here, authors present the cryo-EM structure of PELP1 in complex with its binding partner WDR18, revealing the architecture of PELP1's numerous signaling motifs.
- Jacob Gordon
- , Fleur L. Chapus
- & Robin E. Stanley
-
Article
| Open Access 2′–5′ oligoadenylate synthetase‑like 1 (OASL1) protects against atherosclerosis by maintaining endothelial nitric oxide synthase mRNA stability
2′–5′ oligoadenylate synthetase‑like 1 (OASL1) protects against atherosclerosis by maintaining endothelial nitric oxide synthase mRNA stability
Maintaining optimal eNOS levels is important during cardiovascular events, although little is known regarding the mechanism of eNOS protection. Here, the authors show a regulatory role of endothelial OASL1 in maintaining eNOS mRNA stability and vascular biology under atheroprone conditions.
- Tae Kyeong Kim
- , Sejin Jeon
- & Goo Taeg Oh
-
Article
| Open Access The nucleolus is the site for inflammatory RNA decay during infection
The nucleolus is the site for inflammatory RNA decay during infection
The nucleolus is the traditional site for ribosomal RNA biogenesis. Here, the authors find that the nucleolus is a site of inflammatory pre-mRNA turnover and elucidated how immune homeostasis can be maintained by controlling inflammatory gene expression.
- Taeyun A. Lee
- , Heonjong Han
- & Boyoun Park
-
Article
| Open Access Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Mutations in tRNA processing factors can lead to myelin disorders. This study shows that differentiated oligodendrocytes, cells that make the myelin, are characterized by different tRNA modifications and mRNA decay compared to their precursor cells.
- Sophie Martin
- , Kevin C. Allan
- & Jeff Coller
-
Article
| Open Access Structural analysis of Red1 as a conserved scaffold of the RNA-targeting MTREC/PAXT complex
Structural analysis of Red1 as a conserved scaffold of the RNA-targeting MTREC/PAXT complex
Unwanted RNA transcripts are targeted for degradation by nuclear complexes such as MTREC/PAXT. Here, the authors structurally and functionally characterized three interfaces of the scaffold protein Red1, providing mechanistic insights into conserved features of MTREC/PAXT architecture.
- Anne-Emmanuelle Foucher
- , Leila Touat-Todeschini
- & Jan Kadlec
-
Article
| Open Access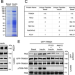 TRIM24 is an insulin-responsive regulator of P-bodies
TRIM24 is an insulin-responsive regulator of P-bodies
Insulin promotes hepatic lipogenesis, though underlying regulation remains unclear. Here the authors show that insulin translocates TRIM24 from the nucleus into cytosolic P-bodies to stabilise hepatic Pparγ mRNA, and that inactivation of TRIM24 promotes Pparγ degradation and alleviates hepatosteatosis.
- Wen Wei
- , Qiaoli Chen
- & Shuai Chen
-
Article
| Open Access Dynamic mRNA degradome analyses indicate a role of histone H3K4 trimethylation in association with meiosis-coupled mRNA decay in oocyte aging
Dynamic mRNA degradome analyses indicate a role of histone H3K4 trimethylation in association with meiosis-coupled mRNA decay in oocyte aging
Developmental potential of oocytes decreases in women of advanced age. Here the authors observe impaired meiosis-coupled mRNA decay and reduced CXXC1-maintained histone H3K4 trimethylation in old oocytes of mouse and humans.
- Yun-Wen Wu
- , Sen Li
- & Qian-Qian Sha
-
Article
| Open Access Gene-specific nonsense-mediated mRNA decay targeting for cystic fibrosis therapy
Gene-specific nonsense-mediated mRNA decay targeting for cystic fibrosis therapy
The W1282X nonsense mutation in the CFTR gene causes cystic fibrosis by reducing its mRNA and functional protein levels. Here the authors developed antisense-oligonucleotide cocktails that restore CFTR protein function by gene-specific stabilization of CFTR mRNA.
- Young Jin Kim
- , Tomoki Nomakuchi
- & Adrian R. Krainer
-
Article
| Open Access Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Processing bodies (PBs) are higher order RNA-protein complexes that determine mRNA fate. Here, the authors show Sbp1 is responsible for PB disassembly and pinpoint a repeats of arginine and glycine (RGG) motif in Sbp1 that play a key role.
- Raju Roy
- , Gitartha Das
- & Purusharth I. Rajyaguru
-
Article
| Open Access Pumilio protects Xbp1 mRNA from regulated Ire1-dependent decay
Pumilio protects Xbp1 mRNA from regulated Ire1-dependent decay
In Drosophila, ER-targeted mRNAs are degraded by Ire1-dependent pathway. Here the authors report that the fly mRNA binding protein Pumilio is phosphorylated by Ire1 and binds to Xbp1 mRNA, protecting it from the non-canonical endoribonuclease activity of Ire1.
- Fátima Cairrão
- , Cristiana C. Santos
- & Pedro M. Domingos
-
Article
| Open Access Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
The authors develop an RNA sequencing-based platform, PERSIST-seq, to simultaneously delineate in-cell mRNA stability, ribosome load, and in-solution stability of a diverse mRNA library to derive design principles for improved mRNA therapeutics.
- Kathrin Leppek
- , Gun Woo Byeon
- & Rhiju Das
-
Article
| Open Access LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3/ATG8 plays an essential role in autophagy. Here the authors show that LC3B exhibits RNA-binding ability and induces rapid degradation of target mRNAs via autophagic activation, highlighting the interplay between autophagy and RNA biology.
- Hyun Jung Hwang
- , Hongseok Ha
- & Yoon Ki Kim
-
Article
| Open Access RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
The CCR4-NOT complex is a key regulator of eukaryotic mRNA deadenylation. Here the authors identified the E3 ubiquitin ligase RNF219 as an acetylation-regulated cofactor of the complex, which inhibits mRNA decay through a direct interaction with NOT9.
- Fabian Poetz
- , Joshua Corbo
- & Georg Stoecklin
-
Article
| Open Access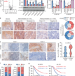 Nicotine-mediated OTUD3 downregulation inhibits VEGF-C mRNA decay to promote lymphatic metastasis of human esophageal cancer
Nicotine-mediated OTUD3 downregulation inhibits VEGF-C mRNA decay to promote lymphatic metastasis of human esophageal cancer
The mechanism of how VEGF-C is regulated to promote lymphatic metastasis of oesophageal cancer is unclear. Here the authors show that nicotine prevents the mRNA decay of VEGF-C mRNA and promotes lymphatic metastasis by downregulating deubiquitinase OTUD3 and RNA binding protein ZFP36.
- Meng Wang
- , Yue Li
- & Libing Song
-
Article
| Open Access Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
The authors show that post-transcriptional regulation of the cilia-driven leftward flow target dand5 is central to symmetry breakage in frog, fish and mouse and is mediated by a 139 nt Bicc1 responsive element in the dand5 3′UTR, and they present evidence that Pkd2 regulates this Bicc1/dand5 module.
- Markus Maerker
- , Maike Getwan
- & Axel Schweickert
-
Article
| Open Access Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
RNA polyadenosine tails are important for the export, translation and stability of mRNAs and play a role in non-coding RNA biogenesis. Here the authors measure yeast poly(A) tail lengths by direct RNA sequencing, revealing its dynamics in yeast exonuclease, deadenylase and poly(A) polymerase mutants.
- Agnieszka Tudek
- , Paweł S. Krawczyk
- & Andrzej Dziembowski
-
Article
| Open Access A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
Premature termination codons can cause early translation termination and lead to disease. Here the authors perform a screen to identify compounds with readthrough activity and show that these reduce eRF1 levels to suppress premature termination associated with cystic fibrosis.
- Jyoti Sharma
- , Ming Du
- & David M. Bedwell
-
Article
| Open Access Fluid flow-induced left-right asymmetric decay of Dand5 mRNA in the mouse embryo requires a Bicc1-Ccr4 RNA degradation complex
Fluid flow-induced left-right asymmetric decay of Dand5 mRNA in the mouse embryo requires a Bicc1-Ccr4 RNA degradation complex
Questioning what regulates left-right asymmetry breaking in the mouse node: the authors identify a 200 bp stretch of the Dand5 3’UTR where Bicc1 binds, and Cnot proteins downstream of calcium flow regulate the post-transcriptional regulation of Dand5 by Bicc1.
- Katsura Minegishi
- , Benjamin Rothé
- & Hiroshi Hamada
-
Article
| Open Access Terminal uridyltransferase 7 regulates TLR4-triggered inflammation by controlling Regnase-1 mRNA uridylation and degradation
Terminal uridyltransferase 7 regulates TLR4-triggered inflammation by controlling Regnase-1 mRNA uridylation and degradation
Terminal uridyltransferase 7 (TUT7) adds U-tails on diverse RNAs to promote degradation. Here the authors show that TUT7 is induced upon LPS treatment in macrophages and promotes decay of Regnase-1, thereby regulating the expression of a subset of cytokines, including IL-6.
- Chia-Ching Lin
- , Yi-Ru Shen
- & Li-Chung Hsu
-
Article
| Open Access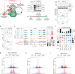 SMG5-SMG7 authorize nonsense-mediated mRNA decay by enabling SMG6 endonucleolytic activity
SMG5-SMG7 authorize nonsense-mediated mRNA decay by enabling SMG6 endonucleolytic activity
Degradation of nonsense mediated mRNA decay (NMD) substrates is carried out by two seemingly independent pathways, SMG6-mediated endonucleolytic cleavage and/or SMG5-SMG7-induced accelerated deadenylation. Here the authors show that SMG5-SMG7 maintain NMD activity by permitting SMG6 activation.
- Volker Boehm
- , Sabrina Kueckelmann
- & Niels H. Gehring
-
Article
| Open Access Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
22G-RNAs are single-stranded antisense small RNAs that are expressed in C. elegans germline. Here the authors show that CSR-1 dependent 22G-RNAs are produced in the cytosol on mRNAs actively engaged in translation and that codon usage of an mRNA regulates the biogenesis of CSR-1 dependent 22G-RNAs.
- Meetali Singh
- , Eric Cornes
- & Germano Cecere
-
Article
| Open Access Selectivity of mRNA degradation by autophagy in yeast
Selectivity of mRNA degradation by autophagy in yeast
Autophagic degradation of proteins and organelles is well studied, but the role of autophagy in RNA regulation is unclear. Here, the authors profile mRNAs targeted to the vacuole in yeast and observe autophagy-mediated mRNA degradation is not random but rather is a preferential process.
- Shiho Makino
- , Tomoko Kawamata
- & Yoshinori Ohsumi
-
Article
| Open Access The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis
The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis
TUTase mediated uridylation of mRNA promotes degradation. Here, Scheer, de Almeida et al. show that Arabidopsis TUTase URT1 interacts directly with the translation inhibitor and decay factor DECAPPING5 and suppresses siRNA biogenesis by preventing accumulation of deadenylated mRNAs
- Hélène Scheer
- , Caroline de Almeida
- & Dominique Gagliardi
-
Article
| Open Access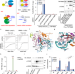 A scaffold lncRNA shapes the mitosis to meiosis switch
A scaffold lncRNA shapes the mitosis to meiosis switch
In fission yeast, the antagonistic RNA-binding proteins Mmi1 and Mei2 respectively promote and inhibit meiotic mRNA degradation during mitotic growth. Here the authors show that the lncRNA mamRNA scaffolds Mmi1 and Mei2 proteins to enable their mutual controls.
- Vedrana Andric
- , Alicia Nevers
- & Mathieu Rougemaille
-
Article
| Open Access Cryo-EM structures of the SARS-CoV-2 endoribonuclease Nsp15 reveal insight into nuclease specificity and dynamics
Cryo-EM structures of the SARS-CoV-2 endoribonuclease Nsp15 reveal insight into nuclease specificity and dynamics
Nsp15 is a uridine specific endoribonuclease present in all coronaviruses. Here, the authors determine the cryo-EM structures of SARS-CoV-2 Nsp15 in the apo and UTP-bound states, which together with biochemical experiments, mass spectrometry and molecular dynamics simulations provide insights into the catalytic mechanism of Nsp15 and its conformational dynamics.
- Monica C. Pillon
- , Meredith N. Frazier
- & Robin E. Stanley
-
Article
| Open Access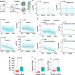 Dynamics and clinical relevance of maternal mRNA clearance during the oocyte-to-embryo transition in humans
Dynamics and clinical relevance of maternal mRNA clearance during the oocyte-to-embryo transition in humans
How maternal RNA clearance is regulated in human preimplantation embryos is unclear. Here, the authors show there is a potential correlation between maternal mRNA decay defects and early developmental arrest from in vitro fertilized human embryos, suggesting that M-decay and Z-decay pathways may regulate such early development.
- Qian-Qian Sha
- , Wei Zheng
- & Heng-Yu Fan
-
Article
| Open Access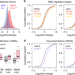 C9orf72 arginine-rich dipeptide repeats inhibit UPF1-mediated RNA decay via translational repression
C9orf72 arginine-rich dipeptide repeats inhibit UPF1-mediated RNA decay via translational repression
C9orf72 repeat expansion is the major genetic cause of familial amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). Here, the authors show that transcriptome aberrations commonly found in c9ALS/FTD are a result from defects in cellular RNA surveillance pathways that involve an RNA helicase UPF1.
- Yu Sun
- , Aziz Eshov
- & Junjie U. Guo
-
Article
| Open Access The RNA quality control pathway nonsense-mediated mRNA decay targets cellular and viral RNAs to restrict KSHV
The RNA quality control pathway nonsense-mediated mRNA decay targets cellular and viral RNAs to restrict KSHV
Cellular nonsense-mediated mRNA decay (NMD) has been shown to play a role in defense against RNA viruses. Here, Zhao et al. show that NMD restricts the DNA virus Kaposi sarcoma-associated herpesvirus (KSHV) via targeting both cellular and viral transcripts leading to inhibition of KSHV lytic reactivation.
- Yang Zhao
- , Xiang Ye
- & John Karijolich
-
Article
| Open Access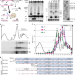 No-Go Decay mRNA cleavage in the ribosome exit tunnel produces 5′-OH ends phosphorylated by Trl1
No-Go Decay mRNA cleavage in the ribosome exit tunnel produces 5′-OH ends phosphorylated by Trl1
In the No-Go decay mRNA surveillance pathway, mRNAs containing stalled ribosomes are cleaved by an endoribonuclease. Here the authors show the endonucleolytic cleavage on the artificial No-Go decay target mRNAs, revealing downstream decay process by Trl1 kinase and the 5′ to 3′ exonuclease Dxo1.
- Albertas Navickas
- , Sébastien Chamois
- & Lionel Benard
-
Article
| Open Access Insights into the assembly and architecture of a Staufen-mediated mRNA decay (SMD)-competent mRNP
Insights into the assembly and architecture of a Staufen-mediated mRNA decay (SMD)-competent mRNP
The Staufen proteins recognize secondary structures in 3’-untranslated regions in mRNA transcripts and induce degradation of these mRNAs with the help of the RNA helicase UPF1. Here the authors report that the nonsense-mediated mRNA decay factor UPF2 mediates the interaction between Stau1 and UPF1 in Staufen-mediated mRNA decay.
- Manjeera Gowravaram
- , Juliane Schwarz
- & Sutapa Chakrabarti
-
Article
| Open Access Thermotolerance in the pathogen Cryptococcus neoformans is linked to antigen masking via mRNA decay-dependent reprogramming
Thermotolerance in the pathogen Cryptococcus neoformans is linked to antigen masking via mRNA decay-dependent reprogramming
The fungal pathogen Cryptococcus neoformans can adapt to mammalian core body temperature. Here, Bloom et al. show that Ccr4-mediated decay of ribosomal protein mRNAs is important for thermotolerance and immune evasion by promoting masking of cell wall glucans.
- Amanda L. M. Bloom
- , Richard M. Jin
- & John C. Panepinto
-
Article
| Open Access A human immune dysregulation syndrome characterized by severe hyperinflammation with a homozygous nonsense Roquin-1 mutation
A human immune dysregulation syndrome characterized by severe hyperinflammation with a homozygous nonsense Roquin-1 mutation
Roquin-1 is a posttranscriptional regulator that controls the expression of many immune-related genes such as ICOS and TNFA. Here, the authors report a homozygous R688* loss of function mutation in Roquin-1 in a patient with syndromic uncontrolled hyperinflammation associated with immune cell activation and hypercytokinemia.
- S. J. Tavernier
- , V. Athanasopoulos
- & F. Haerynck
-
Article
| Open Access RST1 and RIPR connect the cytosolic RNA exosome to the Ski complex in Arabidopsis
RST1 and RIPR connect the cytosolic RNA exosome to the Ski complex in Arabidopsis
Cytosolic RNA degradation by the RNA exosome requires the Ski complex. Here the authors show that the proteins RST1 and RIPR assist the RNA exosome and the Ski complex in RNA degradation, thereby preventing the production of secondary siRNAs from endogenous mRNAs.
- Heike Lange
- , Simon Y. A. Ndecky
- & Dominique Gagliardi
-
Article
| Open Access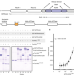 The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs
The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs
The human RNA exosome contains a nuclear co-factor MTR4, which unwinds structural RNAs and recruits adaptors for different RNA processing and decay pathways. Here the authors uncover new variations of the arch-interacting motif (AIM) in NVL and ZCCHC8 and characterise their interaction with MTR4.
- Mahesh Lingaraju
- , Dennis Johnsen
- & Elena Conti
-
Article
| Open Access Reconstitution of recombinant human CCR4-NOT reveals molecular insights into regulated deadenylation
Reconstitution of recombinant human CCR4-NOT reveals molecular insights into regulated deadenylation
The CCR4-NOT complex shortens poly(A) tails of messenger RNAs. By biochemical reconstitution of the entire human CCR4-NOT complex, the authors show the stimulatory roles of non-enzymatic subunits and the importance of the interaction between CAF40 and RNA binding proteins in targeted deadenylation.
- Tobias Raisch
- , Chung-Te Chang
- & Eugene Valkov

 Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq