Featured
-
-
Article
| Open Access Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Changes in glucose metabolism can lead to tumor development, but the involvement of splicing factors is unclear. Here, the authors screened for SR proteins and identified SRSF5 stability is enhanced in response to glucose elevation to promote alternative splicing of CCAR1 which facilitates tumor growth.
- Yuhan Chen
- , Qingyang Huang
- & Lingqiang Zhang
-
Article
| Open Access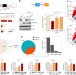 Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Cytoplasmic PTEN is a tumor suppressor that antagonises PI3K signalling. Here, the authors show that nuclear PTEN can interact with the spliceosomal proteins and drive pre-mRNA splicing in a phosphatase-independent manner, in particular, PTEN depletion promotes Golgi extension and secretion through GOLGA2 exon skipping.
- Shao-Ming Shen
- , Yan Ji
- & Guo-Qiang Chen
-
Article
| Open Access Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Alternative splicing of influenza A virus (IAV) M transcript is regulated by hnRNP K and NS1-BP, but mechanistic details are unknown. Here, Thompson et al. show how hnRNP K and NS1-BP bind M mRNA and that these proteins regulate splicing of host transcripts in both the absence and presence of IAV infection.
- Matthew G. Thompson
- , Raquel Muñoz-Moreno
- & Kristen W. Lynch
-
Article
| Open Access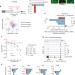 Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
The function and consequences of alternative polyadenylation (APA) in stressed cells are largely unclear. Here, the authors show that stress-induced mRNA degradation depends on 3′UTR length and that APA-mediated 3′UTR shortening is an adaptive stress response mechanism for selective transcript stabilization.
- Dinghai Zheng
- , Ruijia Wang
- & Bin Tian
-
Article
| Open Access HuR regulates telomerase activity through TERC methylation
HuR regulates telomerase activity through TERC methylation
Mutations in the RNA component TERC can cause telomerase dysfunction but the underlying mechanisms are largely unknown. Here, the authors show that RNA-binding protein HuR regulates telomerase function by enhancing the methylation of TERC, which is impaired by several disease-relevant TERC mutations.
- Hao Tang
- , Hu Wang
- & Wengong Wang
-
Article
| Open Access Precise temporal regulation of alternative splicing during neural development
Precise temporal regulation of alternative splicing during neural development
The precise timing of neurodevelopmental splicing switches and the underlying regulatory mechanisms remain poorly understood. This study identifies two major waves of developmental switches under the control of distinct combinations of RNA-binding proteins in central and peripheral nervous systems.
- Sebastien M. Weyn-Vanhentenryck
- , Huijuan Feng
- & Chaolin Zhang
-
Article
| Open Access rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
Myotonic dystrophy (DM) type 2 is a neuromuscular pathology caused by large expansions of CCTG repeats. Here the authors find that rbFOX1 RNA binding protein binds to CCUG RNA repeats and competes with MBNL1 for the binding to CCUG repeats, releasing MBNL1 from sequestration in DM2 muscle cells.
- Chantal Sellier
- , Estefanía Cerro-Herreros
- & Nicolas Charlet-Berguerand
-
Article
| Open Access Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention (IR) can increase protein diversity and function, and yet unregulated IR may be detrimental to cellular health. This study shows that aberrant IR occurs in ALS and finds nuclear loss of an RNA-binding protein called SFPQ as a new molecular hallmark in this devastating condition.
- Raphaelle Luisier
- , Giulia E. Tyzack
- & Rickie Patani
-
Article
| Open Access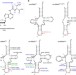 CO2-sensitive tRNA modification associated with human mitochondrial disease
CO2-sensitive tRNA modification associated with human mitochondrial disease
Transfer RNA modifications play critical roles in protein synthesis. Here the authors reveal the t6A37 tRNA modification is dynamically regulated by sensing intracellular CO2 concentration in mitochondria, implying metabolic regulation of protein synthesis.
- Huan Lin
- , Kenjyo Miyauchi
- & Tsutomu Suzuki
-
Article
| Open Access Nuclear fate of yeast snoRNA is determined by co-transcriptional Rnt1 cleavage
Nuclear fate of yeast snoRNA is determined by co-transcriptional Rnt1 cleavage
Small nucleolar ribonucleoprotein complexes (snoRNP) are fundamental for ribosome biogenesis. Here the authors provide insight into the 5ʹend processing of S. cerevisiae snoRNA and its important role in downstream nuclear events.
- Pawel Grzechnik
- , Sylwia A. Szczepaniak
- & Nicholas J. Proudfoot
-
Article
| Open Access Structural analysis of human ARS2 as a platform for co-transcriptional RNA sorting
Structural analysis of human ARS2 as a platform for co-transcriptional RNA sorting
Arsenic resistance protein 2 (ARS2) plays an important role in nuclear RNA metabolism and interacts with the nuclear cap-binding complex (CBC). Here the authors present the human ARS2 structure and identify regions important for its interactions with binding partners supporting that mutually exclusive higher order CBC-ARS2 complexes are formed.
- Wiebke Manuela Schulze
- , Frank Stein
- & Stephen Cusack
-
Article
| Open Access A SUMO-dependent feedback loop senses and controls the biogenesis of nuclear pore subunits
A SUMO-dependent feedback loop senses and controls the biogenesis of nuclear pore subunits
The nuclear pore complex is crucial for mediating nucleocytoplasmic exchanges. Here the authors use budding yeast to reveal a mechanism responsible of maintaining nucleoporin homeostasis by sensing changes in the complex integrity and further altering the metabolism of the corresponding mRNAs.
- Jérôme O. Rouvière
- , Manuel Bulfoni
- & Benoit Palancade
-
Article
| Open Access ZFR coordinates crosstalk between RNA decay and transcription in innate immunity
ZFR coordinates crosstalk between RNA decay and transcription in innate immunity
Type I interferon signaling is critical for the control of infection. Here the authors show that zinc finger RNA-binding protein (ZFR) can control type I interferon responses, and that this control is itself regulated by distinct ZFR truncation patterns that differ between monocytes and macrophages.
- Nazmul Haque
- , Ryota Ouda
- & J. Robert Hogg
-
Article
| Open Access uvCLAP is a fast and non-radioactive method to identify in vivo targets of RNA-binding proteins
uvCLAP is a fast and non-radioactive method to identify in vivo targets of RNA-binding proteins
RNA-binding proteins have important roles in gene expression and regulation. Here the authors develop uvCLAP to purify proteins and determine their binding sites without the use of radioactivity.
- Daniel Maticzka
- , Ibrahim Avsar Ilik
- & Asifa Akhtar
-
Article
| Open Access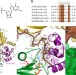 Structure of the activated Edc1-Dcp1-Dcp2-Edc3 mRNA decapping complex with substrate analog poised for catalysis
Structure of the activated Edc1-Dcp1-Dcp2-Edc3 mRNA decapping complex with substrate analog poised for catalysis
The decapping enzyme Dcp2 removes the 5′ eukaryotic cap from mRNA transcripts and acts in concert with its essential activator Dcp1 and various coactivators. Here the authors present the structure of the fully-activated mRNA decapping complex, which reveals how Dcp2 recognizes the cap substrate and coactivators Edc1 and Edc3 activate catalysis.
- Jeffrey S. Mugridge
- , Ryan W. Tibble
- & John D. Gross
-
Article
| Open Access p38-MK2 signaling axis regulates RNA metabolism after UV-light-induced DNA damage
p38-MK2 signaling axis regulates RNA metabolism after UV-light-induced DNA damage
UV-light-induced DNA damage affects RNA metabolism but the underlying signalling pathways are largely unexplored. Here, the authors show that UV light triggers p38-MK2-mediated phosphorylation of the NELF complex, promoting its release from chromatin and concurrent transcriptional elongation.
- Marina E. Borisova
- , Andrea Voigt
- & Petra Beli
-
Article
| Open Access Decapping protein EDC4 regulates DNA repair and phenocopies BRCA1
Decapping protein EDC4 regulates DNA repair and phenocopies BRCA1
Mutations in BRCA1 are associated with an increased risk of breast and ovarian cancer. Here the authors show that EDC4, a component of P-bodies, is a member of the BRCA1 complex with roles in stimulating end resection at breaks.
- Gonzalo Hernández
- , María José Ramírez
- & Jordi Surrallés
-
Article
| Open Access Detecting RNA base methylations in single cells by in situ hybridization
Detecting RNA base methylations in single cells by in situ hybridization
Methylated RNA bases influence many life processes, but current detection methods lack the ability to detect individual methylations in single cells. Here, the authors use fluorescence hybridization probes sensitive to methylation to detect specific epitranscriptomic modifications at the single-cell level.
- Rohan T. Ranasinghe
- , Martin R. Challand
- & David Klenerman
-
Article
| Open Access HTLV-1 Tax plugs and freezes UPF1 helicase leading to nonsense-mediated mRNA decay inhibition
HTLV-1 Tax plugs and freezes UPF1 helicase leading to nonsense-mediated mRNA decay inhibition
UPF1 is a central protein in nonsense-mediated mRNA decay (NMD), but contribution of its RNA processivity to NMD is unclear. Here, the authors show how the retroviral Tax protein interacts with and inhibits UPF1, and demonstrate that UPF1’s translocase activity contributes to NMD.
- Francesca Fiorini
- , Jean-Philippe Robin
- & Vincent Mocquet
-
Article
| Open Access Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
RNA-binding protein hnRNP A2/B1 is suggested to promote miRNA processing as a m6A 'reader'. Here, the authors determine crystal structures of RRM domains of hnRNP A2/B1 in complex with various RNA substrates and determine that hnRNP A2/B1 may function as an auxiliary factor in 'm6A switch' instead.
- Baixing Wu
- , Shichen Su
- & Jinbiao Ma
-
Article
| Open Access Kruppel-like factor 4-dependent Staufen1-mediated mRNA decay regulates cortical neurogenesis
Kruppel-like factor 4-dependent Staufen1-mediated mRNA decay regulates cortical neurogenesis
While being known as a transcription factor, Kruppel-like factor 4 (Klf4) may have other molecular functions. This study shows that Klf4 in neural progenitor cells regulate neurogenesis and self-renewal by interacting with RNA-binding protein Staufen1 and RNA helicase Ddx5/17 to control mRNA decay.
- Byoung-San Moon
- , Jinlun Bai
- & Wange Lu
-
Article
| Open Access Binding of NUFIP2 to Roquin promotes recognition and regulation of ICOS mRNA
Binding of NUFIP2 to Roquin promotes recognition and regulation of ICOS mRNA
The RNA-binding proteins Roquin-1 and Roquin-2 are essential for immune cell function and postnatal survival in mice. Here, the authors identify NUFIP2 as a cofactor of Roquin; Roquin binds and stabilizes NUFIP2 in cells while NUFIP2 regulates Roquin mRNA target recognition.
- Nina Rehage
- , Elena Davydova
- & Vigo Heissmeyer
-
Article
| Open Access Structural analysis of mtEXO mitochondrial RNA degradosome reveals tight coupling of nuclease and helicase components
Structural analysis of mtEXO mitochondrial RNA degradosome reveals tight coupling of nuclease and helicase components
The mitochondrial RNA degradosome (mtEXO) plays an essential role in the regulation of mitochondrial gene expression and is composed of the 3′-to-5′ exoribonuclease Dss1 and the helicase Suv3. Here the authors present the RNA bound mtEXO crystal structure and give insights into its mechanism.
- Michal Razew
- , Zbigniew Warkocki
- & Marcin Nowotny
-
Article
| Open Access RNA degradation by the plant RNA exosome involves both phosphorolytic and hydrolytic activities
RNA degradation by the plant RNA exosome involves both phosphorolytic and hydrolytic activities
The yeast and human RNA exosome is structurally related to prokaryotic phosphorylases but degrades RNA only via associated hydrolytic activities. Here the authors show that the RNA exosome of plants, and likely those of a few basal eukaryotes, combines phosphorolytic and hydrolytic activities to degrade RNA.
- Natalia Sikorska
- , Hélène Zuber
- & Dominique Gagliardi
-
Article
| Open Access Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Several families of natural compounds target core components of the pre-mRNA splicing machinery and display anti-tumor activity. Here the authors show that particular sequence features can be linked to drug response, and that drugs with very similar chemical structures display substantially different effects on splicing regulation.
- Luisa Vigevani
- , André Gohr
- & Juan Valcárcel
-
Article
| Open Access Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Most mammalian genes contain alternative polyadenylation sites. Here, the authors provide evidence that mRNA can be cleaved post-transcriptionally to generate mRNAs with shorter 3-´UTRs and stable autonomous uncapped 3´-UTR sequences.
- Yuval Malka
- , Avital Steiman-Shimony
- & Michael Berger
-
Article
| Open Access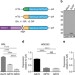 RAN translation at C9orf72-associated repeat expansions is selectively enhanced by the integrated stress response
RAN translation at C9orf72-associated repeat expansions is selectively enhanced by the integrated stress response
A nucleotide repeat expansion in C9orf72 is a common genetic cause of neurodegenerative disorders. Here, the authors provide insight into the molecular mechanism by which this repeat undergoes Repeat-Associated Non-AUG (RAN) translation, implicating the integrated stress response and eIF2α phosphorylation.
- Katelyn M. Green
- , M. Rebecca Glineburg
- & Peter K. Todd
-
Article
| Open Access BEX1 is an RNA-dependent mediator of cardiomyopathy
BEX1 is an RNA-dependent mediator of cardiomyopathy
Little is known about the changes in mRNA splicing, processing and stability that can alter gene expression during heart failure. Here, the authors show that BEX1 is induced during heart failure and is part of a ribonucleoprotein complex enhancing the expression and stability of proinflammatory genes.
- Federica Accornero
- , Tobias G. Schips
- & Jeffery D. Molkentin
-
Article
| Open Access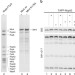 Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Excision of internal transcribed spacer 2 (ITS2) within eukaryotic pre-ribosomal RNA is essential for ribosome function. Here, the authors reconstitute the entire cycle of ITS2 processing in vitro using purified components, providing insights into the cleavage process and demonstrating that 26S pre-rRNA processing necessarily precedes 7S pre-rRNA processing.
- Lisa Fromm
- , Sebastian Falk
- & Ed Hurt
-
Article
| Open Access Regulation of gene expression and RNA editing in Drosophila adapting to divergent microclimates
Regulation of gene expression and RNA editing in Drosophila adapting to divergent microclimates
Environmental adaptation is generally studied at the genomic level, but it may also be driven by transcriptional processes. Here, the authors investigate variation in gene expression and RNA editing across diverging populations of Drosophila melanogaster from two microclimates.
- Arielle L. Yablonovitch
- , Jeremy Fu
- & Jin Billy Li
-
Article
| Open Access LRPPRC-mediated folding of the mitochondrial transcriptome
LRPPRC-mediated folding of the mitochondrial transcriptome
The mitochondrial genome, being compressed to 16 kb, is an attractive model system to investigate how RNA-binding proteins chaperone mRNA lifecycles. Here the authors use RNase footprinting and PAR-CLIP to show that the LRPPRC–SLIRP complex stabilizes mRNA structures to expose sites required for translation and polyadenylation.
- Stefan J. Siira
- , Henrik Spåhr
- & Aleksandra Filipovska
-
Article
| Open Access Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers
Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers
Small molecules correcting the splicing deficit of the survival of motor neuron 2 (SMN2) gene have been identified as having therapeutic potential. Here, the authors provide evidence that SMN2 mRNA forms a ribonucleoprotein complex that can be specifically targeted by these small molecules.
- Manaswini Sivaramakrishnan
- , Kathleen D. McCarthy
- & Friedrich Metzger
-
Article
| Open Access A post-transcriptional program coordinated by CSDE1 prevents intrinsic neural differentiation of human embryonic stem cells
A post-transcriptional program coordinated by CSDE1 prevents intrinsic neural differentiation of human embryonic stem cells
Unlike transcriptional regulation of hESC identity, little is known post-transcriptionally. Here, the authors show that the RNA binding protein CSDE1 regulates core components of hESC identity, neurectoderm commitment and neurogenesis to maintain pluripotency and prevent neural differentiation.
- Hyun Ju Lee
- , Deniz Bartsch
- & David Vilchez
-
Article
| Open Access Structural basis for mutually exclusive co-transcriptional nuclear cap-binding complexes with either NELF-E or ARS2
Structural basis for mutually exclusive co-transcriptional nuclear cap-binding complexes with either NELF-E or ARS2
The nuclear cap-binding complex (CBC) binds to the 5′-cap structure of Pol II transcripts. Here, the authors give structural insights into CBC-mediated transcript processing and show that CBC forms mutual exclusive complexes with NELF and ARS2, which might act in earlier and later phases of transcription, respectively.
- Wiebke Manuela Schulze
- & Stephen Cusack
-
Article
| Open Access Post-transcriptional gene silencing mediated by microRNAs is controlled by nucleoplasmic Sfpq
Post-transcriptional gene silencing mediated by microRNAs is controlled by nucleoplasmic Sfpq
MicroRNAs have been best characterized for their functions in the cytoplasm; however, there is growing evidence of a nuclear localized role. Here, the authors identify Sfpq as an Ago2-interacting protein that modulates miRNA activity in both the nucleus and cytoplasm.
- Silvia Bottini
- , Nedra Hamouda-Tekaya
- & Michele Trabucchi
-
Article
| Open Access CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
U1 snRNP is critical for 5′ splicing site recognition in pre-mRNA splicing. Here the authors describe the cryo-EM structure of the yeast U1 snRNP and suggest that PrpF39 is an alternative splicing factor essential for the successful recruitment of U1 snRNP by other alternative splicing factors.
- Xueni Li
- , Shiheng Liu
- & Rui Zhao
-
Article
| Open Access Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease
Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease
“mRNA abundance is determined by the rates of transcription and decay. Here, the authors propose a method for estimating the rate of differential mRNA decay from RNA-seq data and model mRNA stability in the brain, suggesting a link between mRNA stability and Alzheimer’s disease.”
- Rached Alkallas
- , Lisa Fish
- & Hamed S. Najafabadi
-
Article
| Open Access Extracellular matrix stiffness and cell contractility control RNA localization to promote cell migration
Extracellular matrix stiffness and cell contractility control RNA localization to promote cell migration
Adenomatous polyposis coli (APC) regulates the localization of some mRNAs at cellular protrusions but the underlying mechanisms and functional roles are not known. Here the authors show that APC-dependent RNAs are enriched in contractile protrusions, via detyrosinated microtubules, and enhance cell migration.
- Tianhong Wang
- , Susan Hamilla
- & Stavroula Mili
-
Article
| Open Access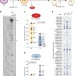 High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
Ribosome biogenesis is a dynamic process that involves the ordered assembly of ribosomal proteins and numerous RNA structural rearrangements. Here the authors apply ChemModSeq, a high-throughput RNA structure probing method, to quantitatively measure changes in RNA flexibility during the nucleolar stages of 60S assembly in yeast.
- Elena Burlacu
- , Fredrik Lackmann
- & Sander Granneman
-
Article
| Open Access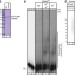 The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
FAM46C is one of the most frequently mutated genes in multiple myeloma (MM), but its molecular function remains unknown. Here the authors show that FAM46C is a poly(A) polymerase and that loss of function of FAM46C drives multiple myeloma through the destabilisation of ER response transcripts.
- Seweryn Mroczek
- , Justyna Chlebowska
- & Andrzej Dziembowski
-
Article
| Open Access RNA localization is a key determinant of neurite-enriched proteome
RNA localization is a key determinant of neurite-enriched proteome
Subcellular localization of RNAs and proteins is important for polarized cells such as neurons. Here the authors differentiate mouse embryonic stem cells into neurons, and analyze the local transcriptome, proteome, and translated transcriptome in their cell bodies and neurites, providing a unique resource for future studies on neuronal polarity.
- Alessandra Zappulo
- , David van den Bruck
- & Marina Chekulaeva
-
Article
| Open Access P-body proteins regulate transcriptional rewiring to promote DNA replication stress resistance
P-body proteins regulate transcriptional rewiring to promote DNA replication stress resistance
P-bodies form in response to stress and act as sites of mRNA storage and degradation. Here the authors identify the mRNA targets of P-bodies during DNA replication stress, and show that P-body proteins act to prevent toxic accumulation of these target transcripts.
- Raphael Loll-Krippleber
- & Grant W. Brown
-
Article
| Open Access Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes
Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes
Ribosomes assemble through the hierarchical addition of proteins to a ribosomal RNA scaffold. Here the authors use three-color single-molecule FRET to show how the dynamics of the rRNA dictate the order in which multiple proteins assemble on the 5′ domain of the E. coli 16S rRNA.
- Sanjaya C. Abeysirigunawardena
- , Hajin Kim
- & Sarah A. Woodson
-
Article
| Open Access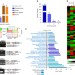 Differential alternative splicing coupled to nonsense-mediated decay of mRNA ensures dietary restriction-induced longevity
Differential alternative splicing coupled to nonsense-mediated decay of mRNA ensures dietary restriction-induced longevity
Alternative splicing coupled to nonsense-mediated decay (AS-NMD) is a conserved mechanism for post-transcriptional gene regulation. Here, the authors provide evidence that AS-NMD is enhanced during dietary restriction (DR) and is required for DR-mediated longevity assurance in C. elegans.
- Syed Shamsh Tabrez
- , Ravi Datta Sharma
- & Arnab Mukhopadhyay
-
Article
| Open Access The m6A pathway facilitates sex determination in Drosophila
The m6A pathway facilitates sex determination in Drosophila
N6-methyladenosine (m6A) is a conserved RNA modification that has recently emerged as an important regulator of messenger RNA processing and activity. Here, the authors provide evidence that m6A pathway facilitates female-specific splicing of Sxl, regulating sex determination in Drosophila.
- Lijuan Kan
- , Anya V. Grozhik
- & Eric C. Lai
-
Article
| Open Access Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation
Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation
TDP-43 aggregation is observed in amyotrophic lateral sclerosis. Here the authors combine X-ray crystallography, nuclear magnetic resonance and electron microscopy studies and show that physiological oligomerization of TDP-43 is mediated through its N-terminal domain, which forms functional and dynamic oligomers antagonizing pathologic aggregation.
- Tariq Afroz
- , Eva-Maria Hock
- & Magdalini Polymenidou
-
Article
| Open Access Crystal structures of U6 snRNA-specific terminal uridylyltransferase
Crystal structures of U6 snRNA-specific terminal uridylyltransferase
After transcription the 3′-end of U6 snRNA is oligo-uridylylated by the terminal uridylyltransferase TUT1. Here the authors present the crystal structure of human TUT1 and give insights into the mechanism of 3′-end uridylylation by the enzyme.
- Seisuke Yamashita
- , Yuko Takagi
- & Kozo Tomita
-
Article
| Open Access R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
The R2TP/Prefoldin-like cochaperone complex is involved in the assembly of a number of protein complexes. Here the authors provide evidence that RUVBL1/RUVBL2, subunits of that cochaperone complex, directly interact with ZNHIT2 to regulate assembly of U5 small ribonucleoprotein.
- Philippe Cloutier
- , Christian Poitras
- & Benoit Coulombe
-
Article
| Open Access Tailing and degradation of Argonaute-bound small RNAs protect the genome from uncontrolled RNAi
Tailing and degradation of Argonaute-bound small RNAs protect the genome from uncontrolled RNAi
While RNA interference is a highly conserved mechanism of gene regulation, how Argonaute-bound small RNAs are targeted for degradation is not well understood. Here the authors show that Cid14 and Cid16 target Argonaute-bound small RNAs for degradation and protect the genome from uncontrolled RNAi activity.
- Paola Pisacane
- & Mario Halic

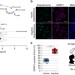 Dedicated surveillance mechanism controls G-quadruplex forming non-coding RNAs in human mitochondria
Dedicated surveillance mechanism controls G-quadruplex forming non-coding RNAs in human mitochondria