Featured
-
-
Article
| Open Access Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
The molecular events underlying the assembly and maturation of the early pre-60S particles during eukaryotic ribosome synthesis are not well understood. Here, the authors combine yeast genetics and biochemical experiments to characterise the functions of two important players of eukaryotic ribosome biogenesis, the box C/D snoRNP snR190 and the helicase Dbp7, which both interact. They show that the snR190 snoRNA acts as a RNA chaperone that assists the structuring of the 25S rRNA during the maturation of early pre-60S particles and that Dbp7 is important for facilitating remodeling events in the peptidyl transferase center region of the 25S rRNAs during the maturation of early pre-60S particles.
- Mariam Jaafar
- , Julia Contreras
- & Anthony K. Henras
-
Article
| Open Access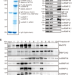 Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
MeCP2 mutations can cause Rett syndrome, a severe childhood neurological disorder. Here the authors show that MeCP2 mediates the higher-order assembly of a large splicing complex Rbfox/LASR, which is disrupted in the mouse models of Rett syndrome.
- Yan Jiang
- , Xing Fu
- & Jingyi Hui
-
Article
| Open Access HIV reprograms host m6Am RNA methylome by viral Vpr protein-mediated degradation of PCIF1
HIV reprograms host m6Am RNA methylome by viral Vpr protein-mediated degradation of PCIF1
m6Am is a modification of the 5′ end of mRNAs catalyzed by PCIF1. Here, Zhang et al. show that HIV infection induces a decrease in m6Am of cellular mRNAs through Vpr-mediated PCIF1 ubiquitination and degradation, resulting in increased HIV replication through regulation of host transcription factors.
- Qiong Zhang
- , Yuqi Kang
- & Tariq M. Rana
-
Article
| Open Access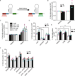 ADAR-mediated RNA editing of DNA:RNA hybrids is required for DNA double strand break repair
ADAR-mediated RNA editing of DNA:RNA hybrids is required for DNA double strand break repair
Different roles of specific RNA-related factors in DNA repair have now been reported. Here the authors reveal a role for RNA-editing by ADAR in DNA end resection following double strand break formation and a change in pattern of ADAR2-mediated A-to-I editing.
- Sonia Jimeno
- , Rosario Prados-Carvajal
- & Pablo Huertas
-
Article
| Open Access Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
The authors show that post-transcriptional regulation of the cilia-driven leftward flow target dand5 is central to symmetry breakage in frog, fish and mouse and is mediated by a 139 nt Bicc1 responsive element in the dand5 3′UTR, and they present evidence that Pkd2 regulates this Bicc1/dand5 module.
- Markus Maerker
- , Maike Getwan
- & Axel Schweickert
-
Article
| Open Access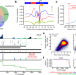 The landscape of alternative polyadenylation in single cells of the developing mouse embryo
The landscape of alternative polyadenylation in single cells of the developing mouse embryo
Alternative polyadenylation regulates localization, half-life and translation of mRNA isoforms. Here the authors investigate alternative polyadenylation using single cell RNA sequencing data from mouse embryos and identify 3’-UTR isoforms that are regulated across cell types and developmental time.
- Vikram Agarwal
- , Sereno Lopez-Darwin
- & Jay Shendure
-
Article
| Open Access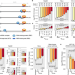 The regulatory impact of RNA-binding proteins on microRNA targeting
The regulatory impact of RNA-binding proteins on microRNA targeting
miRNAs are loaded into Argonaute protein and repress complementary mRNA targets. Here the authors show the unappreciated role of RNA binding proteins for efficient miRNA targeting and expand the current understanding of miRNA targeting.
- Sukjun Kim
- , Soyoung Kim
- & Daehyun Baek
-
Article
| Open Access Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
Global view on the metabolism of RNA poly(A) tails in yeast Saccharomyces cerevisiae
RNA polyadenosine tails are important for the export, translation and stability of mRNAs and play a role in non-coding RNA biogenesis. Here the authors measure yeast poly(A) tail lengths by direct RNA sequencing, revealing its dynamics in yeast exonuclease, deadenylase and poly(A) polymerase mutants.
- Agnieszka Tudek
- , Paweł S. Krawczyk
- & Andrzej Dziembowski
-
Article
| Open Access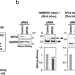 SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
The length distribution of human pre-mRNA introns is very extensive. The authors demonstrate that splicing in a subset of short introns is dependent on SPF45 (RBM17), which replaces authentic U2AF-heterodimer on the truncated poly-pyrimidine tracts and interacts with the U2 snRNP protein SF3b155.
- Kazuhiro Fukumura
- , Rei Yoshimoto
- & Akila Mayeda
-
Article
| Open Access Nuclear export and translation of circular repeat-containing intronic RNA in C9ORF72-ALS/FTD
Nuclear export and translation of circular repeat-containing intronic RNA in C9ORF72-ALS/FTD
Hexanucleotide repeat expansion in the intron 1 of the C9ORF72 gene can cause amyotrophic lateral sclerosis (ALS) and frontal temporal dementia (FTD). Here the authors use single molecule imaging to show nuclear export and translation of circular repeat-containing C9ORF72 intronic RNA.
- Shaopeng Wang
- , Malgorzata J. Latallo
- & Shuying Sun
-
Article
| Open Access A ligand-insensitive UNC5B splicing isoform regulates angiogenesis by promoting apoptosis
A ligand-insensitive UNC5B splicing isoform regulates angiogenesis by promoting apoptosis
UNC5B is a Netrin-1 receptor expressed in endothelial cells that in the absence of ligand induces apoptosis. Here the authors identify an UNC5B splicing isoform that is insensitive to the pro-survival ligand Netrin-1 and is required for apoptosis-dependent blood vessel development.
- Davide Pradella
- , Gianluca Deflorian
- & Claudia Ghigna
-
Article
| Open Access RETRACTED ARTICLE: AKT3-mediated IWS1 phosphorylation promotes the proliferation of EGFR-mutant lung adenocarcinomas through cell cycle-regulated U2AF2 RNA splicing
RETRACTED ARTICLE: AKT3-mediated IWS1 phosphorylation promotes the proliferation of EGFR-mutant lung adenocarcinomas through cell cycle-regulated U2AF2 RNA splicing
IWS1 regulates multiple steps in RNA metabolism, including RNA elongation and alternative RNA splicing. Here the authors show that AKT3 phosphorylates IWS1, which alters U2AF2 RNA splicing and promotes growth of lung adenocarcinomas via a Sororin/ERK-dependent pathway.
- Georgios I. Laliotis
- , Evangelia Chavdoula
- & Philip N. Tsichlis
-
Article
| Open Access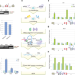 The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
We know that most splicing reactions take place co-transcriptionally, but how the transcription machinery facilitate splicing of introns is unknown. Here the authors show that the 5′ splice site remains associated with the transcription machinery during intron synthesis through U1 snRNP, providing a basis for the rapid splicing reaction of introns.
- Yodfat Leader
- , Galit Lev Maor
- & Gil Ast
-
Article
| Open Access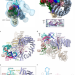 Structural basis of intron selection by U2 snRNP in the presence of covalent inhibitors
Structural basis of intron selection by U2 snRNP in the presence of covalent inhibitors
Chemical modulation of intron selection has emerged as a route for cancer therapy. Here, structures of the U2 snRNP’s SF3B module and of prespliceosome- both in complexes with splicing modulators- provide insight into the mechanisms of intron recognition and branch site inactivation.
- Constantin Cretu
- , Patricia Gee
- & Vladimir Pena
-
Article
| Open Access Therapeutic manipulation of IKBKAP mis-splicing with a small molecule to cure familial dysautonomia
Therapeutic manipulation of IKBKAP mis-splicing with a small molecule to cure familial dysautonomia
Familial dysautonomia is caused by splicing mutation of IKBKAP gene, which induces skipping of exon 20 and subsequent functional loss. Here, the authors report that a synthetic splice modulator RECTAS ameliorates pathogenic exon 20 skipping and shows therapeutic effects in cellular and animal models.
- Masahiko Ajiro
- , Tomonari Awaya
- & Masatoshi Hagiwara
-
Article
| Open Access A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion
Premature termination codons can cause early translation termination and lead to disease. Here the authors perform a screen to identify compounds with readthrough activity and show that these reduce eRF1 levels to suppress premature termination associated with cystic fibrosis.
- Jyoti Sharma
- , Ming Du
- & David M. Bedwell
-
Article
| Open Access Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
The Argonaute protein CSR-1 is essential for fertility and viability in C. elegans. Here the authors show that CSR-1A isoform associates preferentially with small RNAs mapping to spermatogenesis-specific genes while CSR-1B isoform binds small RNAs mapping to oogenesis-specific genes. Arginine methylation of CSR-1A promotes small RNA-binding specificity.
- Dieu An H. Nguyen
- & Carolyn M. Phillips
-
Article
| Open Access Fluid flow-induced left-right asymmetric decay of Dand5 mRNA in the mouse embryo requires a Bicc1-Ccr4 RNA degradation complex
Fluid flow-induced left-right asymmetric decay of Dand5 mRNA in the mouse embryo requires a Bicc1-Ccr4 RNA degradation complex
Questioning what regulates left-right asymmetry breaking in the mouse node: the authors identify a 200 bp stretch of the Dand5 3’UTR where Bicc1 binds, and Cnot proteins downstream of calcium flow regulate the post-transcriptional regulation of Dand5 by Bicc1.
- Katsura Minegishi
- , Benjamin Rothé
- & Hiroshi Hamada
-
Article
| Open Access Terminal uridyltransferase 7 regulates TLR4-triggered inflammation by controlling Regnase-1 mRNA uridylation and degradation
Terminal uridyltransferase 7 regulates TLR4-triggered inflammation by controlling Regnase-1 mRNA uridylation and degradation
Terminal uridyltransferase 7 (TUT7) adds U-tails on diverse RNAs to promote degradation. Here the authors show that TUT7 is induced upon LPS treatment in macrophages and promotes decay of Regnase-1, thereby regulating the expression of a subset of cytokines, including IL-6.
- Chia-Ching Lin
- , Yi-Ru Shen
- & Li-Chung Hsu
-
Article
| Open Access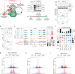 SMG5-SMG7 authorize nonsense-mediated mRNA decay by enabling SMG6 endonucleolytic activity
SMG5-SMG7 authorize nonsense-mediated mRNA decay by enabling SMG6 endonucleolytic activity
Degradation of nonsense mediated mRNA decay (NMD) substrates is carried out by two seemingly independent pathways, SMG6-mediated endonucleolytic cleavage and/or SMG5-SMG7-induced accelerated deadenylation. Here the authors show that SMG5-SMG7 maintain NMD activity by permitting SMG6 activation.
- Volker Boehm
- , Sabrina Kueckelmann
- & Niels H. Gehring
-
Article
| Open Access The human nucleoporin Tpr protects cells from RNA-mediated replication stress
The human nucleoporin Tpr protects cells from RNA-mediated replication stress
Tpr nucleoporin is known to be essential for nuclear transport and mitosis processes. Here the authors explore the link between Tpr and genome instability providing insights into the role of Tpr in safeguarding cells from RNA-mediated replication stress.
- Martin Kosar
- , Michele Giannattasio
- & Marco Foiani
-
Article
| Open Access Hakai is required for stabilization of core components of the m6A mRNA methylation machinery
Hakai is required for stabilization of core components of the m6A mRNA methylation machinery
The E3 ligase Hakai can interact with the m6A methylation machinery but its function is still unclear. Here, the authors show that Hakai is a conserved component of the m6A methyltransferase complex and provide functional and molecular insights into its role in regulating m6A levels in Drosophila.
- Praveen Bawankar
- , Tina Lence
- & Jean-Yves Roignant
-
Article
| Open Access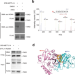 Arginine methylation of METTL14 promotes RNA N6-methyladenosine modification and endoderm differentiation of mouse embryonic stem cells
Arginine methylation of METTL14 promotes RNA N6-methyladenosine modification and endoderm differentiation of mouse embryonic stem cells
The methyltransferase complex of METTL3-METTL14-WTAP is responsible for m6A modification on RNA. Here the authors report that METTL14 arginine 255 (R255) is methylated by PRMT1 and this modification increases interaction of METTL3/METTL14 interaction with WTAP and substrate RNA, promoting m6A methylation activity of the complex.
- Xiaona Liu
- , Hailong Wang
- & Shan Xiao
-
Article
| Open Access TSSC4 is a component of U5 snRNP that promotes tri-snRNP formation
TSSC4 is a component of U5 snRNP that promotes tri-snRNP formation
The correct assembly and recycling of the multicomponent spliceosome remains largely elusive. Here, the authors show that a previously uncharacterized protein TSSC4 associates with de novo formed spliceosomal U5 snRNP as well as with a post-splicing U5-PRPF19 particle, and that TSSC4 is important for assembly of the splicing competent tri-snRNP.
- Klára Klimešová
- , Jitka Vojáčková
- & David Staněk
-
Article
| Open Access Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
22G-RNAs are single-stranded antisense small RNAs that are expressed in C. elegans germline. Here the authors show that CSR-1 dependent 22G-RNAs are produced in the cytosol on mRNAs actively engaged in translation and that codon usage of an mRNA regulates the biogenesis of CSR-1 dependent 22G-RNAs.
- Meetali Singh
- , Eric Cornes
- & Germano Cecere
-
Article
| Open Access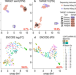 MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
Confounding factors on gene expression analysis can be analyzed by several existing tools. Here the authors develop an algorithm called MOCCASIN to correct the effect of known and unknown confounders on RNA splicing quantification.
- Barry Slaff
- , Caleb M. Radens
- & Yoseph Barash
-
Article
| Open Access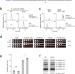 A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
Spliceosomal U6 snRNA is modified by m6A. Here the authors show that m6A modification on U6 snRNA promotes efficient recognition of the splice site through cooperating with U5 snRNA, indicating that U6 snRNA m6A allows the 5’ exon sequence variation.
- Yuma Ishigami
- , Takayuki Ohira
- & Tsutomu Suzuki
-
Article
| Open Access Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Yeast helicase Prp28 promotes the first step of spliceosome remodeling. By placing a photoactivatable unnatural amino acid in Prp28, the authors capture Prp28 in action revealing its dynamic interactions and cofactor Npl3.
- Fu-Lung Yeh
- , Shang-Lin Chang
- & Tien-Hsien Chang
-
Article
| Open Access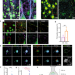 Synaptic FUS accumulation triggers early misregulation of synaptic RNAs in a mouse model of ALS
Synaptic FUS accumulation triggers early misregulation of synaptic RNAs in a mouse model of ALS
Mutations in the RNA-binding protein FUS contribute to ALS. Here the authors use CLIP-seq on synaptoneurosomes to identify proteins associated with synapse organization and plasticity that are differentially regulated in a knock-in ALS mouse model.
- Sonu Sahadevan
- , Katharina M. Hembach
- & Magdalini Polymenidou
-
Article
| Open Access The RNA binding protein FgRbp1 regulates specific pre-mRNA splicing via interacting with U2AF23 in Fusarium
The RNA binding protein FgRbp1 regulates specific pre-mRNA splicing via interacting with U2AF23 in Fusarium
Human RBM42 associates with the spliceosome complex. Here the authors show that the fungus counterpart of RBM42, FgRbp1 regulates splicing by interacting with FgU2AF23.
- Minhui Wang
- , Tianling Ma
- & Zhonghua Ma
-
Article
| Open Access Selectivity of mRNA degradation by autophagy in yeast
Selectivity of mRNA degradation by autophagy in yeast
Autophagic degradation of proteins and organelles is well studied, but the role of autophagy in RNA regulation is unclear. Here, the authors profile mRNAs targeted to the vacuole in yeast and observe autophagy-mediated mRNA degradation is not random but rather is a preferential process.
- Shiho Makino
- , Tomoko Kawamata
- & Yoshinori Ohsumi
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Role of Hakai in m6A modification pathway in Drosophila
Role of Hakai in m6A modification pathway in Drosophila
Drosophila m6A writer complex regulates alternative splicing of the Sex-lethal gene. Here the authors show that a potential E3 ligase Hakai interacts with the fly m6A writer complex and that m6A level is reduced in Hakai mutant flies.
- Yanhua Wang
- , Lifeng Zhang
- & Dong Yan
-
Article
| Open Access Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Arginine methyltransferases (PRMTs) are involved in the regulation of various physiological and pathological conditions. Using proteomics, the authors here profile the methylation substrates of PRMTs 4, 5 and 7 and characterize the roles of these enzymes in cancer-associated splicing regulation.
- Wen-juan Li
- , Yao-hui He
- & Wen Liu
-
Article
| Open Access A conserved role for the ALS-linked splicing factor SFPQ in repression of pathogenic cryptic last exons
A conserved role for the ALS-linked splicing factor SFPQ in repression of pathogenic cryptic last exons
SFPQ is a splicing factor and its mutations are associated to amyotrophic lateral sclerosis (ALS) patients. Here, the authors show that SFPQ represses the use of pathogenic cryptic last exons in zebrafish, mouse and human cells.
- Patricia M. Gordon
- , Fursham Hamid
- & Corinne Houart
-
Article
| Open Access R-loop resolution promotes co-transcriptional chromatin silencing
R-loop resolution promotes co-transcriptional chromatin silencing
Nascent non-coding RNA can mediate chromatin silencing, however mechanistically this process is poorly understood. Here the authors show that resolution of an R-loop during 3'-end processing of a plant antisense transcript recruits chromatin modifiers to promote chromatin silencing.
- Congyao Xu
- , Zhe Wu
- & Caroline Dean
-
Article
| Open Access ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells
ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells
One type of RNA editing involves ADAR-mediated conversion of adenosine to inosine. Here the authors show that ADAR1 nuclear isoform p110 regulates R loop formation and genome stability at telomeres in cancer cells.
- Yusuke Shiromoto
- , Masayuki Sakurai
- & Kazuko Nishikura
-
Article
| Open Access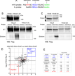 A neural m6A/Ythdf pathway is required for learning and memory in Drosophila
A neural m6A/Ythdf pathway is required for learning and memory in Drosophila
Epitranscriptomic modifications can regulate learning and memory. Here, the authors provide proteomic and functional analysis of N6-methyladenosine (m6A)-binding proteins in D. melanogaster and unveil behavioral and regulatory defects for m6A/Ythdf mutants.
- Lijuan Kan
- , Stanislav Ott
- & Eric C. Lai
-
Article
| Open Access The methyltransferase SETD2 couples transcription and splicing by engaging mRNA processing factors through its SHI domain
The methyltransferase SETD2 couples transcription and splicing by engaging mRNA processing factors through its SHI domain
The methylation of Histone 3 at Lysine 36 (H3K36) has been implicated in the regulation of transcription and coupled processes such as mRNA splicing. Here the authors show that the histone methyltransferase SETD2 interacts with hnRNP L to mediate the crosstalk between the transcription and splicing machineries.
- Saikat Bhattacharya
- , Michaella J. Levy
- & Jerry L. Workman
-
Article
| Open Access A choreography of centrosomal mRNAs reveals a conserved localization mechanism involving active polysome transport
A choreography of centrosomal mRNAs reveals a conserved localization mechanism involving active polysome transport
Centrosomes function as microtubule organizing centers where several mRNAs accumulate. By employing high-throughput single molecule FISH screening, the authors discover that 8 human mRNAs localize to centrosomes with unique cell cycle dependent patterns using an active polysome targeting mechanism.
- Adham Safieddine
- , Emeline Coleno
- & Edouard Bertrand
-
Article
| Open Access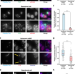 Exon junction complex dependent mRNA localization is linked to centrosome organization during ciliogenesis
Exon junction complex dependent mRNA localization is linked to centrosome organization during ciliogenesis
Exon junction complexes (EJCs) that mark untranslated mRNA are involved in transport, translation and nonsense-mediated mRNA decay. Here the authors show centrosomal localization of EJCs which appears to be required for both the localization of NIN mRNA around centrosomes and ciliogenesis.
- Oh Sung Kwon
- , Rahul Mishra
- & Hervé Le Hir
-
Article
| Open Access The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis
The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis
TUTase mediated uridylation of mRNA promotes degradation. Here, Scheer, de Almeida et al. show that Arabidopsis TUTase URT1 interacts directly with the translation inhibitor and decay factor DECAPPING5 and suppresses siRNA biogenesis by preventing accumulation of deadenylated mRNAs
- Hélène Scheer
- , Caroline de Almeida
- & Dominique Gagliardi
-
Article
| Open Access Interaction of 7SK with the Smn complex modulates snRNP production
Interaction of 7SK with the Smn complex modulates snRNP production
The noncoding RNA 7SK controls the transcription of mRNAs. Here, the authors show that the 7SK complex interacts with the Smn complex, suggesting crosstalk between transcription and snRNP assembly.
- Changhe Ji
- , Jakob Bader
- & Michael Briese
-
Article
| Open Access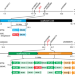 The genetic basis of cytoplasmic male sterility and fertility restoration in wheat
The genetic basis of cytoplasmic male sterility and fertility restoration in wheat
The development of hybrid wheat cultivars is hampered by the lack of an effective way to control male fertility in breeding lines. Here, the authors report the identification of two restorer-of-fertility genes Rf1 and Rf3 that can restore fertility of wheat plants carrying Triticum timopheevii-type cytoplasmic male sterility.
- Joanna Melonek
- , Jorge Duarte
- & Ian Small
-
Article
| Open Access C. elegans germ granules require both assembly and localized regulators for mRNA repression
C. elegans germ granules require both assembly and localized regulators for mRNA repression
Nematode P granules are cytoplasmic RNA–protein biomolecule condensates central to germ cell development. Here the authors show that dimerization of the PGL-1 scaffolding protein is crucial to granule formation and mRNA repression, and that the WAGO-1 Argonaute protein is a cofactor in repressing PGL-1 bound mRNAs.
- Scott Takeo Aoki
- , Tina R. Lynch
- & Judith Kimble
-
Article
| Open Access L1 retrotransposons exploit RNA m6A modification as an evolutionary driving force
L1 retrotransposons exploit RNA m6A modification as an evolutionary driving force
L1 is a group of active retrotransposons in humans. Here the authors show that m6A modifications on L1 RNA increase translation efficiency and retrotransposition in human cells. M6A motifs are more enriched in evolutionary young L1s.
- Sung-Yeon Hwang
- , Hyunchul Jung
- & Kwangseog Ahn
-
Article
| Open Access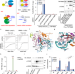 A scaffold lncRNA shapes the mitosis to meiosis switch
A scaffold lncRNA shapes the mitosis to meiosis switch
In fission yeast, the antagonistic RNA-binding proteins Mmi1 and Mei2 respectively promote and inhibit meiotic mRNA degradation during mitotic growth. Here the authors show that the lncRNA mamRNA scaffolds Mmi1 and Mei2 proteins to enable their mutual controls.
- Vedrana Andric
- , Alicia Nevers
- & Mathieu Rougemaille
-
Article
| Open Access Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Chromatin is known to regulate splicing by modulating recruitment of splicing factors. Using machine learning approaches, the authors have underlined a chromatin code for alternative splicing regulation that is conserved amongst cell lines.
- E. Agirre
- , A. J. Oldfield
- & R. F. Luco
-
Article
| Open Access Cryo-EM structures of the SARS-CoV-2 endoribonuclease Nsp15 reveal insight into nuclease specificity and dynamics
Cryo-EM structures of the SARS-CoV-2 endoribonuclease Nsp15 reveal insight into nuclease specificity and dynamics
Nsp15 is a uridine specific endoribonuclease present in all coronaviruses. Here, the authors determine the cryo-EM structures of SARS-CoV-2 Nsp15 in the apo and UTP-bound states, which together with biochemical experiments, mass spectrometry and molecular dynamics simulations provide insights into the catalytic mechanism of Nsp15 and its conformational dynamics.
- Monica C. Pillon
- , Meredith N. Frazier
- & Robin E. Stanley

 The RNA helicase Dbp7 promotes domain V/VI compaction and stabilization of inter-domain interactions during early 60S assembly
The RNA helicase Dbp7 promotes domain V/VI compaction and stabilization of inter-domain interactions during early 60S assembly