Article
|
Open Access
Featured
-
-
Article
| Open Access hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells
hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells
Coordinated regulation of alternative splicing is essential for germ cell development. Here, the authors report that hnRNPH1 interacts with alternative splicing factors PTBP2 and SRSF3 in the germline to regulate pre-mRNA alternative splicing.
- Shenglei Feng
- , Jinmei Li
- & Shuiqiao Yuan
-
Article
| Open Access Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
The protein gephyrin is involved in organizing synapses. Here, the authors show how different transcripts of gephyrin form and regulate inhibitory synapses.
- Raphaël Dos Reis
- , Etienne Kornobis
- & Eric Allemand
-
Article
| Open Access Dynamic mRNA degradome analyses indicate a role of histone H3K4 trimethylation in association with meiosis-coupled mRNA decay in oocyte aging
Dynamic mRNA degradome analyses indicate a role of histone H3K4 trimethylation in association with meiosis-coupled mRNA decay in oocyte aging
Developmental potential of oocytes decreases in women of advanced age. Here the authors observe impaired meiosis-coupled mRNA decay and reduced CXXC1-maintained histone H3K4 trimethylation in old oocytes of mouse and humans.
- Yun-Wen Wu
- , Sen Li
- & Qian-Qian Sha
-
Article
| Open Access Dynamic arrest and aging of biomolecular condensates are modulated by low-complexity domains, RNA and biochemical activity
Dynamic arrest and aging of biomolecular condensates are modulated by low-complexity domains, RNA and biochemical activity
Here the authors analyze material properties and aging of active phase-separated condensates by Differential Dynamic Microscopy. Arrested states are promoted by structured RNA. Low-complexity domains and biochemical reaction keep the droplets fluid-like.
- Miriam Linsenmeier
- , Maria Hondele
- & Paolo Arosio
-
Article
| Open Access Gene-specific nonsense-mediated mRNA decay targeting for cystic fibrosis therapy
Gene-specific nonsense-mediated mRNA decay targeting for cystic fibrosis therapy
The W1282X nonsense mutation in the CFTR gene causes cystic fibrosis by reducing its mRNA and functional protein levels. Here the authors developed antisense-oligonucleotide cocktails that restore CFTR protein function by gene-specific stabilization of CFTR mRNA.
- Young Jin Kim
- , Tomoki Nomakuchi
- & Adrian R. Krainer
-
Article
| Open Access RNase H2, mutated in Aicardi‐Goutières syndrome, resolves co-transcriptional R-loops to prevent DNA breaks and inflammation
RNase H2, mutated in Aicardi‐Goutières syndrome, resolves co-transcriptional R-loops to prevent DNA breaks and inflammation
RnaseH2 is mutated in severe neuro-inflammatory disorder Aicardi‐Goutières syndrome. Here the authors reveal that RNase H2 controls cellular R-loop homeostasis to promote transcription, genome integrity and prevent R-loop-associated inflammation.
- Agnese Cristini
- , Michael Tellier
- & Natalia Gromak
-
Article
| Open Access RNP components condense into repressive RNP granules in the aging brain
RNP components condense into repressive RNP granules in the aging brain
Aberrant RNA condensates are a hallmark of age-related neurodegenerative diseases. Here, the authors show that RNA condensation increases in aging Drosophila brains, triggering translation repression.
- Kavya Vinayan Pushpalatha
- , Mathilde Solyga
- & Florence Besse
-
Article
| Open Access RNA supply drives physiological granule assembly in neurons
RNA supply drives physiological granule assembly in neurons
RNA granules are important regulators of RNA metabolism. Here the authors report that RNA granules containing RNA helicase DDX6 disassemble during neuronal maturation.
- Karl E. Bauer
- , Niklas Bargenda
- & Michael A. Kiebler
-
Perspective
| Open Access Autophagy regulation by RNA alternative splicing and implications in human diseases
Autophagy regulation by RNA alternative splicing and implications in human diseases
Both alternative splicing and autophagy are core cell biological processes, but where they intersect has received little attention. Here, the authors reflect on recent connections identified between these pathways and consider their impact on human disease.
- Patricia González-Rodríguez
- , Daniel J. Klionsky
- & Bertrand Joseph
-
Article
| Open Access Deep learning modeling m6A deposition reveals the importance of downstream cis-element sequences
Deep learning modeling m6A deposition reveals the importance of downstream cis-element sequences
The site specificity of m6A mRNA modification is a fundamental RNA biology question. Here, Luo et al., develop the iM6A, a deep learning method to model this process, showing that its site-specificity is determined by the primary nucleotide sequence.
- Zhiyuan Luo
- , Jiacheng Zhang
- & Shengdong Ke
-
Article
| Open Access Enhancers regulate 3′ end processing activity to control expression of alternative 3′UTR isoforms
Enhancers regulate 3′ end processing activity to control expression of alternative 3′UTR isoforms
Transcriptional enhancers are known to regulate transcript production. Here, the authors show that transcription factors that bind to enhancers also regulate polyadenylation site cleavage activity, thus changing expression of alternative 3’ UTRs.
- Buki Kwon
- , Mervin M. Fansler
- & Christine Mayr
-
Article
| Open Access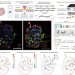 Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
Spatial epitranscriptomics reveals A-to-I editome specific to cancer stem cell microniches
The spatial context of epitranscriptomic features in the tumour microenvironment remains poorly understood. Here, a method for transcriptomic and epitranscriptomic analysis of immunofluorescence-stained tissue, Select-seq, is applied to stem cell-like microniches in triple negative breast cancer.
- Amos C. Lee
- , Yongju Lee
- & Sunghoon Kwon
-
Article
| Open Access Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Stop codon recoding is a unique mechanism for selenoprotein biogenesis. Here, the authors report regulatory systems for selenoprotein expression during skeletal muscle differentiation by regulating A-to-I RNA editing and stop codon recoding.
- Yuta Noda
- , Shunpei Okada
- & Tsutomu Suzuki
-
Article
| Open Access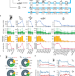 Developmental mRNA m5C landscape and regulatory innovations of massive m5C modification of maternal mRNAs in animals
Developmental mRNA m5C landscape and regulatory innovations of massive m5C modification of maternal mRNAs in animals
mRNAs are known to be decorated with m5C at a low-to-medium level. Here, the authors generate atlases of mRNA m5C during animal development in 6 species and identify convergent and unexpected massive methylation of maternal mRNAs by NSUN2 and NSUN6.
- Jianheng Liu
- , Tao Huang
- & Rui Zhang
-
Article
| Open Access Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
RNA modifications represent a critical aspect of RNA biology that is not well suited to sequencing methods. Here, the authors provide a software tool for automated analysis of RNA tandem mass spectra with full support of modifications, isotope labelling, and control of false discovery rate.
- Luigi D’Ascenzo
- , Anna M. Popova
- & James R. Williamson
-
Article
| Open Access Crystal structures and insights into precursor tRNA 5’-end processing by prokaryotic minimal protein-only RNase P
Crystal structures and insights into precursor tRNA 5’-end processing by prokaryotic minimal protein-only RNase P
HARP are member of protein-only RNase P, which catalyzes pre-tRNA 5’-end processing and maturation. Here, the authors present crystal structure and provide mechanistic insights into pre-tRNA binding and cleavage by HARP proteins.
- Yangyang Li
- , Shichen Su
- & Jianhua Gan
-
Article
| Open Access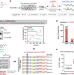 TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
RNA modifications are important regulators of RNA biology. Here we report N1-methyladenosine (m1A) enrichment on 22-nucleotide tRNA fragments and its effect on gene-silencing. Higher level of m1A in bladder cancer is accompanied by gene dysregulation in unfolded protein response.
- Zhangli Su
- , Ida Monshaugen
- & Anindya Dutta
-
Article
| Open Access Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Processing bodies (PBs) are higher order RNA-protein complexes that determine mRNA fate. Here, the authors show Sbp1 is responsible for PB disassembly and pinpoint a repeats of arginine and glycine (RGG) motif in Sbp1 that play a key role.
- Raju Roy
- , Gitartha Das
- & Purusharth I. Rajyaguru
-
Article
| Open Access Multilayered control of splicing regulatory networks by DAP3 leads to widespread alternative splicing changes in cancer
Multilayered control of splicing regulatory networks by DAP3 leads to widespread alternative splicing changes in cancer
RNA binding proteins (RBPs) can participate in regulatory networks to control alternative splicing. Here the authors show that DAP3 functions as an RBP splicing modulator via two mechanisms, and that its overexpression leads to mis-splicing events in cancers.
- Jian Han
- , Omer An
- & Leilei Chen
-
Article
| Open Access Clueless/CLUH regulates mitochondrial fission by promoting recruitment of Drp1 to mitochondria
Clueless/CLUH regulates mitochondrial fission by promoting recruitment of Drp1 to mitochondria
Drp1 is the master regulator of mitochondrial fission, which has important impact on cellular functions. Here, Yang et al identified evolutionarily conserved proteins Clueless and its homolog CLUH as key regulators of Drp1 that function via translation of Drp1 receptors MiD49 and Mff.
- Huan Yang
- , Caroline Sibilla
- & Ming Guo
-
Article
| Open Access Pumilio protects Xbp1 mRNA from regulated Ire1-dependent decay
Pumilio protects Xbp1 mRNA from regulated Ire1-dependent decay
In Drosophila, ER-targeted mRNAs are degraded by Ire1-dependent pathway. Here the authors report that the fly mRNA binding protein Pumilio is phosphorylated by Ire1 and binds to Xbp1 mRNA, protecting it from the non-canonical endoribonuclease activity of Ire1.
- Fátima Cairrão
- , Cristiana C. Santos
- & Pedro M. Domingos
-
Article
| Open Access Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
The authors develop an RNA sequencing-based platform, PERSIST-seq, to simultaneously delineate in-cell mRNA stability, ribosome load, and in-solution stability of a diverse mRNA library to derive design principles for improved mRNA therapeutics.
- Kathrin Leppek
- , Gun Woo Byeon
- & Rhiju Das
-
Article
| Open Access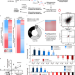 ADARs act as potent regulators of circular transcriptome in cancer
ADARs act as potent regulators of circular transcriptome in cancer
RNA editing and circRNAs are involved in tumorigenesis. Here the authors report that ADARs regulate the circular transcriptome in a bidirectional manner through and beyond their editing function in multiple cancer cells.
- Haoqing Shen
- , Omer An
- & Leilei Chen
-
Article
| Open Access An RNA-binding protein acts as a major post-transcriptional modulator in Bacillus anthracis
An RNA-binding protein acts as a major post-transcriptional modulator in Bacillus anthracis
HitRS is a two-component system that responds to cell envelope damage in the bacterium Bacillus anthracis. Here, the authors identify an RNA-binding protein that regulates HitRS function by modulating the stability of the hitRS mRNA. In addition, the protein binds to over 70 RNAs and affects the expression of genes involved in multiple cellular processes.
- Hualiang Pi
- , Andy Weiss
- & Eric P. Skaar
-
Article
| Open Access WTAP-mediated m6A modification of lncRNA NORAD promotes intervertebral disc degeneration
WTAP-mediated m6A modification of lncRNA NORAD promotes intervertebral disc degeneration
Intervertebral disc degeneration (IVDD) is the leading cause of low back pain and Nucleus pulposus cell senescence contributes a lot to its progression. Here, the authors reveal WTAP-mediated m6A modification of lncRNA NORAD promotes IVDD.
- Gaocai Li
- , Liang Ma
- & Cao Yang
-
Article
| Open Access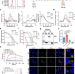 RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
Understanding the mechanisms of SARS-CoV-2 infection is important to control the pandemic. Here the authors show the biological and pathological role of RNA G-quadruplex structure in both SARS-CoV-2 genome and host factors, particularly TMPRSS2.
- Geng Liu
- , Wenya Du
- & Xianghui Fu
-
Article
| Open Access LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3/ATG8 plays an essential role in autophagy. Here the authors show that LC3B exhibits RNA-binding ability and induces rapid degradation of target mRNAs via autophagic activation, highlighting the interplay between autophagy and RNA biology.
- Hyun Jung Hwang
- , Hongseok Ha
- & Yoon Ki Kim
-
Article
| Open Access The aberrant upregulation of exon 10-inclusive SREK1 through SRSF10 acts as an oncogenic driver in human hepatocellular carcinoma
The aberrant upregulation of exon 10-inclusive SREK1 through SRSF10 acts as an oncogenic driver in human hepatocellular carcinoma
Alternative splicing is dysregulated in hepatocellular carcinoma. Here, the authors investigate the role of the splice variant of Splicing Regulatory Glutamic Acid and Lysine Rich Protein 1 (SREK1) and its upstream regulator, Serine/arginine-rich splicing factor 10 (SRSF10) in sustaining the oncogenic signal.
- Cunjie Chang
- , Muthukumar Rajasekaran
- & Jianxiang Chen
-
Article
| Open Access Functional genomics of RAP proteins and their role in mitoribosome regulation in Plasmodium falciparum
Functional genomics of RAP proteins and their role in mitoribosome regulation in Plasmodium falciparum
The function of RNA-binding domain abundant in Apicomplexans (RAP) protein family members is largely unknown. Here, using high-throughput functional genomics, including metabolomics, Hollin et al. characterize two RAP proteins that are essential for Plasmodium falciparum survival and control mitochondrial rRNAs.
- Thomas Hollin
- , Steven Abel
- & Karine G. Le Roch
-
Article
| Open Access The methyl phosphate capping enzyme Bmc1/Bin3 is a stable component of the fission yeast telomerase holoenzyme
The methyl phosphate capping enzyme Bmc1/Bin3 is a stable component of the fission yeast telomerase holoenzyme
There exists a remarkable evolutionary breadth of telomerase holoenzyme components among eukaryotes. Here the authors report that the fission yeast methyl phosphate capping enzyme Bin3/Bmc1 regulates telomerase activity by promoting holoenzyme formation.
- Jennifer Porat
- , Moaine El Baidouri
- & Mark A. Bayfield
-
Article
| Open Access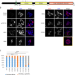 Sirtuin-1 sensitive lysine-136 acetylation drives phase separation and pathological aggregation of TDP-43
Sirtuin-1 sensitive lysine-136 acetylation drives phase separation and pathological aggregation of TDP-43
TDP-43 is a nucleic acid binding protein, whose insoluble aggregates are neuropathological hallmarks of specific subsets of patients with amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Post-translational modifications and acetylation of TDP-43 impact its interaction with RNA, its localization in the cells, and are linked to disease. Using antibodies generated against TDP-43 lysine acetylation sites, sirtuin-1 was found to potently deacetylate amber suppressed [acK136]TDP-43 and reduce its aggregation propensity. Thus, distinct lysine acetylations modulate nuclear import, RNA binding as well as phase separation and aggregation of TDP-43, suggesting regulatory mechanisms for TDP-43 pathogenesis.
- Jorge Garcia Morato
- , Friederike Hans
- & Philipp J. Kahle
-
Article
| Open Access Arabidopsis RBV is a conserved WD40 repeat protein that promotes microRNA biogenesis and ARGONAUTE1 loading
Arabidopsis RBV is a conserved WD40 repeat protein that promotes microRNA biogenesis and ARGONAUTE1 loading
MicroRNAs regulate gene expression through RNA cleavage or translation repression. Here the authors show that RBV, an evolutionarily conserved WD40 domain protein, acts to promote MIR transcription, pri-miRNA processing and miRNA loading into AGO1.
- Chao Liang
- , Qiang Cai
- & Xuemei Chen
-
Article
| Open Access CMTr cap-adjacent 2′-O-ribose mRNA methyltransferases are required for reward learning and mRNA localization to synapses
CMTr cap-adjacent 2′-O-ribose mRNA methyltransferases are required for reward learning and mRNA localization to synapses
The two cap methyltransferases (CMTrs) redundantly methylate riboses of first cap adjacent nucleotides in messenger RNAs in Drosophila. Here, CMTrs are required for reward learning and localization of untranslated messenger RNAs to synapses.
- Irmgard U. Haussmann
- , Yanying Wu
- & Matthias Soller
-
Article
| Open Access A multi-factor trafficking site on the spliceosome remodeling enzyme BRR2 recruits C9ORF78 to regulate alternative splicing
A multi-factor trafficking site on the spliceosome remodeling enzyme BRR2 recruits C9ORF78 to regulate alternative splicing
Bergfort et al. use biochemistry, cryoEM, structure-guided mutagenesis, transcriptomics and proteomics to reveal how the intrinsically unstructured C9ORF78 protein binds BRR2 and PRPF8, regulating cassette exons and alternative 3’-splice sites.
- Alexandra Bergfort
- , Marco Preußner
- & Markus C. Wahl
-
Article
| Open Access Two zinc finger proteins with functions in m6A writing interact with HAKAI
Two zinc finger proteins with functions in m6A writing interact with HAKAI
The components of m6A writer and their interactions are still far from fully understood. Here, the authors identify two HAKAI-interacting zinc finger proteins, HIZ1 and HIZ2, as components of the Arabidopsis m6A writer complex, and show that hiz2 mutant plants have an 85% reduction in m6A abundance and severe developmental defects.
- Mi Zhang
- , Zsuzsanna Bodi
- & Rupert G. Fray
-
Article
| Open Access A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
Rebelo-Guiomar et al. unveil late stage assembly intermediates of the human mitochondrial ribosome by inactivating the methyltransferase MRM2 in cells. Absence of MRM2 impairs organismal homeostasis, while its catalytic activity is dispensable for mitoribosomal biogenesis.
- Pedro Rebelo-Guiomar
- , Simone Pellegrino
- & Michal Minczuk
-
Article
| Open Access CPEB1 directs muscle stem cell activation by reprogramming the translational landscape
CPEB1 directs muscle stem cell activation by reprogramming the translational landscape
Skeletal muscle stem cells are actively maintained in quiescence, but can activate quickly upon extrinsic stimulation. Here the authors show that CPEB1 promotes muscle stem cell activation by reprogramming the translational landscape.
- Wenshu Zeng
- , Lu Yue
- & Tom H. Cheung
-
Article
| Open Access Xrn1 is a deNADding enzyme modulating mitochondrial NAD-capped RNA
Xrn1 is a deNADding enzyme modulating mitochondrial NAD-capped RNA
The cytoplasmic Xrn1 protein has long been established as the predominate 5′ to 3′ exoribonuclease that cleaves RNAs with an unprotected 5′ monophosphate end. Here the authors demonstrate Xrn1 can also degrade RNAs harboring the noncanonical nicotinamide adenine diphosphate (NAD) 5′ cap by removing the NAD cap and degrading the RNA.
- Sunny Sharma
- , Jun Yang
- & Megerditch Kiledjian
-
Article
| Open Access Parallel functional assessment of m6A sites in human endodermal differentiation with base editor screens
Parallel functional assessment of m6A sites in human endodermal differentiation with base editor screens
N6-methyladenosine (m6A) plays important role in lineage specifications of embryonic stem cells, but its role at specific sites has not been assessed. Here the authors develop an adenine editor-based strategy, and systematically identify functional m6A sites that control lineage decisions in human embryonic stem cells.
- Weisheng Cheng
- , Fang Liu
- & Jinkai Wang
-
Article
| Open Access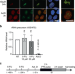 Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Different mechanisms have been reported to explain resistance to chemotherapy in cancer. Here, the authors show that the chemotherapeutic drug 5-fluorouracil alters the function of ribosomes to promote pro-survival gene translation leading to chemotherapy resistance.
- Gabriel Therizols
- , Zeina Bash-Imam
- & Jean-Jacques Diaz
-
Article
| Open Access Proximity labeling identifies a repertoire of site-specific R-loop modulators
Proximity labeling identifies a repertoire of site-specific R-loop modulators
R-loops are three-stranded nucleic acid structures that contribute to genome instability and accumulate in neurological diseases. Here the authors identify R-loop proximal factors, which are enriched for zinc finger and homeodomain proteins, including activity-dependent neuroprotective protein (ADNP). ADNP plays a role in R-loop resolution and loss-of-function leads to R-loop accumulation.
- Qingqing Yan
- , Phillip Wulfridge
- & Kavitha Sarma
-
Article
| Open Access Small molecule splicing modifiers with systemic HTT-lowering activity
Small molecule splicing modifiers with systemic HTT-lowering activity
Here the authors describe the discovery of a class of small molecule splicing modifiers which are orally bioavailable, cross the blood-brain barrier, and lower levels of huntingtin in a mouse model of Huntington’s disease (HD).
- Anuradha Bhattacharyya
- , Christopher R. Trotta
- & Stuart W. Peltz
-
Article
| Open Access Cellular variability of nonsense-mediated mRNA decay
Cellular variability of nonsense-mediated mRNA decay
Here the author developed a single-cell reporter system to identify cell-to-cell variability of nonsense-mediated mRNA decay (NMD). This approach provides a sensitive tool to investigate cellular heterogeneity of NMD in various physiological conditions.
- Hanae Sato
- & Robert H. Singer
-
Article
| Open Access Dynamic regulation of N6,2′-O-dimethyladenosine (m6Am) in obesity
Dynamic regulation of N6,2′-O-dimethyladenosine (m6Am) in obesity
m6A and m6Am are two prevalent mRNA modifications which are target for removal by the fat mass and obesity gene FTO. Here the authors capture the differential profile of these two modifications in the liver of obese mice and identify dynamic translation regulation by the m6Am modification.
- Moshe Shay Ben-Haim
- , Yishay Pinto
- & Gideon Rechavi
-
Article
| Open Access RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
The CCR4-NOT complex is a key regulator of eukaryotic mRNA deadenylation. Here the authors identified the E3 ubiquitin ligase RNF219 as an acetylation-regulated cofactor of the complex, which inhibits mRNA decay through a direct interaction with NOT9.
- Fabian Poetz
- , Joshua Corbo
- & Georg Stoecklin
-
Article
| Open Access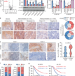 Nicotine-mediated OTUD3 downregulation inhibits VEGF-C mRNA decay to promote lymphatic metastasis of human esophageal cancer
Nicotine-mediated OTUD3 downregulation inhibits VEGF-C mRNA decay to promote lymphatic metastasis of human esophageal cancer
The mechanism of how VEGF-C is regulated to promote lymphatic metastasis of oesophageal cancer is unclear. Here the authors show that nicotine prevents the mRNA decay of VEGF-C mRNA and promotes lymphatic metastasis by downregulating deubiquitinase OTUD3 and RNA binding protein ZFP36.
- Meng Wang
- , Yue Li
- & Libing Song
-
Article
| Open Access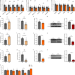 Fat mass and obesity-associated protein regulates RNA methylation associated with depression-like behavior in mice
Fat mass and obesity-associated protein regulates RNA methylation associated with depression-like behavior in mice
Post-transcriptional modification of RNA can contribute to regulating behavior. Here, the authors show that modulating the expression of Fto results in epitranscriptomic changes in the mouse hippocampus associated with depression-like behavior.
- Shu Liu
- , Jianbo Xiu
- & Qi Xu
-
Article
| Open Access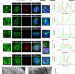 DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
In yeast, TFIP11 and DHX15 promote the disassembly of spliceosome complex after splicing is completed. Here the authors show that human TFIP11 functions independently of DHX15 and is required for U6 snRNA 2’-O-methylation and U4/U6.U5 tri-snRNP assembly.
- Amandine Duchemin
- , Tina O’Grady
- & Denis Mottet
-
Article
| Open Access The RNA-binding protein HuR is required for maintenance of the germinal centre response
The RNA-binding protein HuR is required for maintenance of the germinal centre response
Germinal centre (GC) responses may require RNA binding proteins (RBP) for post-transcriptional gene regulation. Here the authors show the RBP HuR supports GCs by promoting Myc and Myc-dependent transcription to enhance antigen-specific GC B cell selection and production of high affinity antibodies.
- Ines C. Osma-Garcia
- , Dunja Capitan-Sobrino
- & Manuel D. Diaz-Muñoz

 Limited effects of m6A modification on mRNA partitioning into stress granules
Limited effects of m6A modification on mRNA partitioning into stress granules