Featured
-
-
Article
| Open Access Distinct pre-initiation steps in human mitochondrial translation
Distinct pre-initiation steps in human mitochondrial translation
Translation within mitochondria relies upon specialized mitoribosomes and initiation factors. Here the authors use Cryo-EM and single molecule approaches to describe the early steps leading to translation initiation in human mitochondria, and the roles of mitochondria-specific ribosomal proteins and initiation factors mtIF2 and mtIF3.
- Anas Khawaja
- , Yuzuru Itoh
- & Joanna Rorbach
-
Article
| Open Access Ribosome engineering reveals the importance of 5S rRNA autonomy for ribosome assembly
Ribosome engineering reveals the importance of 5S rRNA autonomy for ribosome assembly
Ribosomes of all organisms have retained 5S rRNA as an autonomous rRNA species. Here the authors engineer a bacterial strain with ribosomes that do not have free 5S rRNA, and carry structural analyses that suggest the evolutionary preservation of 5S rRNA as an independent molecule is based on its role in the dynamic process of ribosome biogenesis.
- Shijie Huang
- , Nikolay A. Aleksashin
- & Alexander S. Mankin
-
Article
| Open Access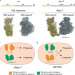 A fully orthogonal system for protein synthesis in bacterial cells
A fully orthogonal system for protein synthesis in bacterial cells
Ribosome engineering is an emerging powerful approach for synthetic protein synthesis. Here the authors invert the Ribo-T system, using the engineered ribosome to translate the proteome while the native ribosome translates specific mRNA.
- Nikolay A. Aleksashin
- , Teresa Szal
- & Alexander S. Mankin
-
Article
| Open Access Ribosome profiling at isoform level reveals evolutionary conserved impacts of differential splicing on the proteome
Ribosome profiling at isoform level reveals evolutionary conserved impacts of differential splicing on the proteome
Genes express multiple mRNA isoforms through alternative processing. Here the authors analyze ribosome profiling data with ORQAS (ORF quantification pipeline for alternative splicing) and report that 40–50% of the expressed mRNA isoforms are translated.
- Marina Reixachs-Solé
- , Jorge Ruiz-Orera
- & Eduardo Eyras
-
Article
| Open Access Mechanism of ribosome shutdown by RsfS in Staphylococcus aureus revealed by integrative structural biology approach
Mechanism of ribosome shutdown by RsfS in Staphylococcus aureus revealed by integrative structural biology approach
Upon transition to stationary phase or upon stress, bacteria limit protein synthesis through small inhibitory proteins that bind the ribosome. Here the authors decipher the interaction mode of the bacterial ribosome silencing factor (RsfS) at atomic details to provide an in depth view of how it shutdowns ribosomes.
- Iskander Khusainov
- , Bulat Fatkhullin
- & Marat Yusupov
-
Article
| Open Access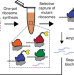 In vitro ribosome synthesis and evolution through ribosome display
In vitro ribosome synthesis and evolution through ribosome display
Directed evolution of the ribosome is challenging because the requirement of cell viability limits the mutations that can be made. Here the authors develop a platform for in vitro ribosome synthesis and evolution (RISE) to overcome these constraints.
- Michael J. Hammerling
- , Brian R. Fritz
- & Michael C. Jewett
-
Article
| Open Access MetAP-like Ebp1 occupies the human ribosomal tunnel exit and recruits flexible rRNA expansion segments
MetAP-like Ebp1 occupies the human ribosomal tunnel exit and recruits flexible rRNA expansion segments
The ErbB3 receptor binding protein Ebp1 binds to ribosomes and is linked to translational control. Here, the authors present the cryo-EM structure of human Ebp1 bound to a non-translating 80S ribosome and find that Ebp1 blocks the tunnel exit and recruits the rRNA expansion segment ES27L to the tunnel exit.
- Klemens Wild
- , Milan Aleksić
- & Irmgard Sinning
-
Article
| Open Access Identification of distinct maturation steps involved in human 40S ribosomal subunit biosynthesis
Identification of distinct maturation steps involved in human 40S ribosomal subunit biosynthesis
Ribosome synthesis is a complex multi-step process. Here the authors present a method that allows the efficient isolation and characterization of the preribosomal complexes formed along the entire ribosome synthesis pathway in human cells.
- Blanca Nieto
- , Sonia G. Gaspar
- & Mercedes Dosil
-
Article
| Open Access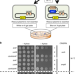 Release factor-dependent ribosome rescue by BrfA in the Gram-positive bacterium Bacillus subtilis
Release factor-dependent ribosome rescue by BrfA in the Gram-positive bacterium Bacillus subtilis
In bacteria, the conserved trans-translation system serves as the primary pathway of ribosome rescue, but many species can also use alternative rescue pathways. Here the authors report that in B. subtilis, the rescue factor BrfA binds to non-stop stalled ribosomes, recruits RF2 but not RF1, and induces transition of the ribosome into an open active conformation.
- Naomi Shimokawa-Chiba
- , Claudia Müller
- & Shinobu Chiba
-
Article
| Open Access Structure of ribosome-bound azole-modified peptide phazolicin rationalizes its species-specific mode of bacterial translation inhibition
Structure of ribosome-bound azole-modified peptide phazolicin rationalizes its species-specific mode of bacterial translation inhibition
The authors report the identification of phazolicin (PHZ) - a prokaryotic translation inhibitory peptide - and its structure in complex with the E. coli ribosome, delineating PHZ’s mode of action and suggesting a basis for its bacterial species-specific activity.
- Dmitrii Y. Travin
- , Zoe L. Watson
- & Konstantin Severinov
-
Article
| Open Access Engineered ribosomes with tethered subunits for expanding biological function
Engineered ribosomes with tethered subunits for expanding biological function
Ribo-T is a tethered ribosome complex capable of orthogonal ribosome-mRNA functionality, but has low activity. Here the authors evolve new tether designs that support faster growth and increased protein expression.
- Erik D. Carlson
- , Anne E. d’Aquino
- & Michael C. Jewett
-
Article
| Open Access Toxin-mediated ribosome stalling reprograms the Mycobacterium tuberculosis proteome
Toxin-mediated ribosome stalling reprograms the Mycobacterium tuberculosis proteome
MazF endoribonucleases are thought to arrest growth of Mycobacterium tuberculosis by global translation inhibition. Here, Barth et al. show that MazF-mt9 cleaves a specific tRNA, leading to ribosome stalling at AAA codons and thus selective mRNA degradation and changes in transcriptome and proteome.
- Valdir C. Barth
- , Ju-Mei Zeng
- & Nancy A. Woychik
-
Article
| Open Access Importance of potassium ions for ribosome structure and function revealed by long-wavelength X-ray diffraction
Importance of potassium ions for ribosome structure and function revealed by long-wavelength X-ray diffraction
Metal ions play essential roles in myriads of biological processes, from catalytic co-factors to supporting protein and nucleic acid structures. Here the authors use long-wavelength X-ray diffraction to locate hundreds of potassium ions taking part in the formation of rRNA tertiary structure, mediating rRNA–protein interactions and supporting ribosomal protein structures and function.
- Alexey Rozov
- , Iskander Khusainov
- & Gulnara Yusupova
-
Article
| Open Access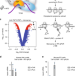 Co-translational assembly of mammalian nuclear multisubunit complexes
Co-translational assembly of mammalian nuclear multisubunit complexes
Genes encoding protein complex subunits are often dispersed in the genome of eukaryotes, raising the question how these protein complexes assemble. Here, the authors provide evidence that mammalian nuclear transcription complexes are formed co-translationally to ensure specific and functional interactions.
- Ivanka Kamenova
- , Pooja Mukherjee
- & László Tora
-
Article
| Open Access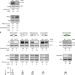 The exonuclease Xrn1 activates transcription and translation of mRNAs encoding membrane proteins
The exonuclease Xrn1 activates transcription and translation of mRNAs encoding membrane proteins
The exonuclease Xrn1 mediates crosstalk between transcription and mRNA decay in yeast. Here the authors demonstrate that Xrn1 promotes translation of mRNAs encoding membrane proteins, coupling transcription, translation, and mRNA decay.
- Bernat Blasco-Moreno
- , Leire de Campos-Mata
- & Juana Díez
-
Article
| Open Access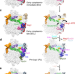 Tightly-orchestrated rearrangements govern catalytic center assembly of the ribosome
Tightly-orchestrated rearrangements govern catalytic center assembly of the ribosome
In eukaryotes, ribosome biogenesis culminates in the cytoplasm with the maturation of the peptidyl transfer center (PTC). Here the authors describe several structures of intermediates in late nuclear and cytoplasmic maturation of the large ribosomal subunit that reveal the tightly-choreographed sequence of protein and RNA rearrangements that lead to the completion of the PTC.
- Yi Zhou
- , Sharmishtha Musalgaonkar
- & David W. Taylor
-
Article
| Open Access Assembly and functionality of the ribosome with tethered subunits
Assembly and functionality of the ribosome with tethered subunits
The tethered ribosome system Ribo-T supports cell proliferation though at a reduced rate. Here the authors show this is due to slower ribosome assembly instead of reduced functionality.
- Nikolay A. Aleksashin
- , Margus Leppik
- & Alexander S. Mankin
-
Article
| Open Access Numerous cultivated and uncultivated viruses encode ribosomal proteins
Numerous cultivated and uncultivated viruses encode ribosomal proteins
Viruses can encode genes that regulate the host's translational machinery to their advantage. Here, the authors show that viruses encode ribosomal proteins that can be incorporated into the host’s ribosome and may affect translation.
- Carolina M. Mizuno
- , Charlotte Guyomar
- & Mart Krupovic
-
Article
| Open Access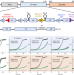 Cryo-EM structure of the essential ribosome assembly AAA-ATPase Rix7
Cryo-EM structure of the essential ribosome assembly AAA-ATPase Rix7
Rix7 is a type II AAA-ATPase that is required for the assembly of the large ribosomal subunit. Here the authors present the 4.5 Å cryo-EM structure of the Rix7 homohexamer with a polypeptide fragment bound in its central channel and provide insights into the function of Rix7 as a molecular unfoldase.
- Yu-Hua Lo
- , Mack Sobhany
- & Robin E. Stanley
-
Article
| Open Access Optimization of carbon and energy utilization through differential translational efficiency
Optimization of carbon and energy utilization through differential translational efficiency
Microorganisms must regulate allocation of resources in nutrient-limited conditions. Here, the authors combine Ribo-seq, RNA-seq and TSS-seq to study resource allocation in the acetogen C. ljungdahlii, and show that dynamic regulation of translational efficiency of metabolic pathways is critical.
- Mahmoud M. Al-Bassam
- , Ji-Nu Kim
- & Karsten Zengler
-
Article
| Open Access Cryo-EM structure of the hibernating Thermus thermophilus 100S ribosome reveals a protein-mediated dimerization mechanism
Cryo-EM structure of the hibernating Thermus thermophilus 100S ribosome reveals a protein-mediated dimerization mechanism
During stress conditions bacterial ribosomes dimerize and form inactive but stable hibernating 100S particles, a process that is facilitated by the hibernation-promoting factor (HPF). Here the authors analyze 100S dimer formation as a function of HPF protein concentration and present the Thermus thermophilus 100S ribosome cryo-EM structure.
- Rasmus Kock Flygaard
- , Niels Boegholm
- & Lasse B. Jenner
-
Article
| Open Access Suppressor mutations in Rpf2–Rrs1 or Rpl5 bypass the Cgr1 function for pre-ribosomal 5S RNP-rotation
Suppressor mutations in Rpf2–Rrs1 or Rpl5 bypass the Cgr1 function for pre-ribosomal 5S RNP-rotation
During biogenesis of the eukaryotic 60S ribosome, a large rotational movement of the 5S RNP is required to achieve its mature position. By analyzing extragenic suppressors of crg1—a key factor required for rotation—the authors provide mechanistic insight into a key step of ribosome biogenesis.
- Matthias Thoms
- , Valentin Mitterer
- & Ed Hurt
-
Article
| Open Access Visualization of translation termination intermediates trapped by the Apidaecin 137 peptide during RF3-mediated recycling of RF1
Visualization of translation termination intermediates trapped by the Apidaecin 137 peptide during RF3-mediated recycling of RF1
In bacteria, the process of translation termination is performed by three termination release factors RF1, RF2 and RF3. Here the authors provide detailed structural insights into the mechanism by which RF1 is dissociated from the ribosome by RF3 during termination.
- Michael Graf
- , Paul Huter
- & Daniel N. Wilson
-
Article
| Open Access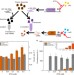 Dynamic allocation of orthogonal ribosomes facilitates uncoupling of co-expressed genes
Dynamic allocation of orthogonal ribosomes facilitates uncoupling of co-expressed genes
Competition between synthetic genetic circuits and host genes for shared resources can complicate circuit design and lead to failure. Here the authors demonstrate, mathematically and experimentally, the use of orthogonal ribosomes to decouple competing genes.
- Alexander P. S. Darlington
- , Juhyun Kim
- & Declan G. Bates
-
Article
| Open Access CUG initiation and frameshifting enable production of dipeptide repeat proteins from ALS/FTD C9ORF72 transcripts
CUG initiation and frameshifting enable production of dipeptide repeat proteins from ALS/FTD C9ORF72 transcripts
Repeat-associated non-AUG (RAN) translation contributes to the pathogenic mechanism of several microsatellite expansion diseases. Here the authors delineate the different steps involved in recruiting the ribosome to initiate G4C2 RAN translation to produce poly-Glycine Alanine, poly-Glycine Proline, and poly-Glycine Arginine repeats.
- Ricardos Tabet
- , Laure Schaeffer
- & Clotilde Lagier-Tourenne
-
Article
| Open Access Inhibition of Poly(A)-binding protein with a synthetic RNA mimic reduces pain sensitization in mice
Inhibition of Poly(A)-binding protein with a synthetic RNA mimic reduces pain sensitization in mice
Poly(A)-binding protein (PABP) is an RNA binding protein with translation function. Here, Barragán-Iglesias and colleagues devise an RNA mimic that inhibits PABP activity, and show that inhibitors can reduce animal’s pain response in vivo when injected locally.
- Paulino Barragán-Iglesias
- , Tzu-Fang Lou
- & Zachary T. Campbell
-
Article
| Open Access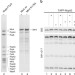 Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Excision of internal transcribed spacer 2 (ITS2) within eukaryotic pre-ribosomal RNA is essential for ribosome function. Here, the authors reconstitute the entire cycle of ITS2 processing in vitro using purified components, providing insights into the cleavage process and demonstrating that 26S pre-rRNA processing necessarily precedes 7S pre-rRNA processing.
- Lisa Fromm
- , Sebastian Falk
- & Ed Hurt
-
Article
| Open Access Atomic resolution snapshot of Leishmania ribosome inhibition by the aminoglycoside paromomycin
Atomic resolution snapshot of Leishmania ribosome inhibition by the aminoglycoside paromomycin
Leishmaniasis is a parasitic disease transmitted by the bite of infected sand flies. Here the authors describe an atomic resolution cryo-EM structure of the Leishmania ribosome in complex with the recently approved drug paromomycin (PAR) and highlight conserved elements in the drug binding pocket that mediate PAR deleterious effects on the parasite.
- Moran Shalev-Benami
- , Yan Zhang
- & Georgios Skiniotis
-
Article
| Open Access A conformational switch in initiation factor 2 controls the fidelity of translation initiation in bacteria
A conformational switch in initiation factor 2 controls the fidelity of translation initiation in bacteria
The GTP-bound form of initiation factor 2 (IF2) promotes translation initiation by accelerating 50S ribosomal subunit joining the 30S ribosomal initiation complex (30S IC). Here the authors use single-molecule FRET and ensemble rapid kinetic methods to uncover the mechanism behind IF2-mediated subunit joining.
- Kelvin Caban
- , Michael Pavlov
- & Ruben L. Gonzalez Jr
-
Article
| Open Access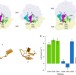 Origin of the omnipotence of eukaryotic release factor 1
Origin of the omnipotence of eukaryotic release factor 1
The eukaryotic release factor eRF1 is able to recognize the three stop codons UAA, UAG and UGA with high accuracy, while discriminating against near-cognate codons. Here the authors use molecular dynamic simulation to provide insight into the molecular basis behind the remarkable codon specificity of eRF1.
- Christoffer Lind
- , Ana Oliveira
- & Johan Åqvist
-
Article
| Open Access The Hsp70 homolog Ssb affects ribosome biogenesis via the TORC1-Sch9 signaling pathway
The Hsp70 homolog Ssb affects ribosome biogenesis via the TORC1-Sch9 signaling pathway
The yeast Hsp70 homolog Ssb is a chaperone that binds translating ribosomes where it is thought to function primarily by promoting nascent peptide folding. Here the authors find that the ribosome biogenesis defect associated with the loss of Ssb is attributable to a specific disruption in TORC1 signaling rather than defects in ribosomal protein folding.
- Kaivalya Mudholkar
- , Edith Fitzke
- & Sabine Rospert
-
Article
| Open Access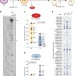 High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
Ribosome biogenesis is a dynamic process that involves the ordered assembly of ribosomal proteins and numerous RNA structural rearrangements. Here the authors apply ChemModSeq, a high-throughput RNA structure probing method, to quantitatively measure changes in RNA flexibility during the nucleolar stages of 60S assembly in yeast.
- Elena Burlacu
- , Fredrik Lackmann
- & Sander Granneman
-
Article
| Open Access A general mechanism of ribosome dimerization revealed by single-particle cryo-electron microscopy
A general mechanism of ribosome dimerization revealed by single-particle cryo-electron microscopy
When bacteria enter the stationary growth phase, protein translation is suppressed via the dimerization of 70S ribosomes into inactive complexes. Here the authors provide a structural basis for how the dual domain hibernation promotion factor promotes ribosome dimerization and hibernation in bacteria.
- Linda E. Franken
- , Gert T. Oostergetel
- & Albert Guskov
-
Article
| Open Access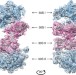 The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
Under conditions of nutrient limitation, bacterial ribosomes undergo dimerization, forming a 100S complex that is translationally inactive. Here the authors present the structural basis for formation of the 100S complexes in Gram-positive bacteria, shedding light on the mechanism of translation suppression by the ribosome-silencing factors.
- Donna Matzov
- , Shintaro Aibara
- & Ada E. Yonath
-
Article
| Open Access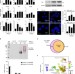 Tia1 dependent regulation of mRNA subcellular location and translation controls p53 expression in B cells
Tia1 dependent regulation of mRNA subcellular location and translation controls p53 expression in B cells
Sequestering mRNA in cytoplasmic stress granules is a mechanism for translational repression. Here the authors find that p53 mRNA, present in stress granules in activated B lymphocytes, is released upon DNA damage and is translated in a CAP-independent manner.
- Manuel D. Díaz-Muñoz
- , Vladimir Yu. Kiselev
- & Martin Turner
-
Article
| Open Access The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan
The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan
The transcription factor Gcn4 is known to regulate yeast amino acid synthesis. Here, the authors show that Gcn4 also acts as a repressor of protein biosynthesis in a range of conditions that enhance yeast lifespan, such as ribosomal protein knockout, calorie restriction or mTOR inhibition.
- Nitish Mittal
- , Joao C. Guimaraes
- & Mihaela Zavolan
-
Article
| Open Access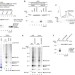 Ubiquitination of stalled ribosome triggers ribosome-associated quality control
Ubiquitination of stalled ribosome triggers ribosome-associated quality control
Several protein quality control mechanisms are in place to trigger the rapid degradation of aberrant polypeptides and mRNAs. Here the authors describe a mechanism of ribosome-mediated quality control that involves the ubiquitination of ribosomal proteins by the E3 ubiquitin ligase Hel2/RQT1.
- Yoshitaka Matsuo
- , Ken Ikeuchi
- & Toshifumi Inada
-
Article
| Open Access Protein-directed ribosomal frameshifting temporally regulates gene expression
Protein-directed ribosomal frameshifting temporally regulates gene expression
Programmed −1 ribosomal frameshifting (−1 PRF) is a mechanism whereby specific signals within mRNAs direct ribosomes to shift into an alternative reading frame. Here the authors describe a mechanism of −1 PRF that is temporally regulated by a viral protein over the course of the virus replicative cycle.
- Sawsan Napthine
- , Roger Ling
- & Andrew E. Firth
-
Article
| Open Access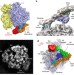 Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Membrane proteins are inserted co-transnationally through the association between ribosome, the signal recognition particle and its receptor, and the membrane-bound translocon. Here the authors present a cryo-EM reconstruction of this quaternary complex in the activated state and propose a model for signal sequence transfer to the translocon.
- Ahmad Jomaa
- , Yu-Hsien Hwang Fu
- & Nenad Ban
-
Article
| Open Access PTENβ is an alternatively translated isoform of PTEN that regulates rDNA transcription
PTENβ is an alternatively translated isoform of PTEN that regulates rDNA transcription
PTEN is a potent tumour suppressor involved in cell growth, proliferation and survival. Here the authors identify an N-terminal extended isoform of PTEN, termed PTENβ that negatively regulates ribosomal DNA transcription and cell proliferation, expanding the pleiotropic functions of the PTEN family.
- Hui Liang
- , Xi Chen
- & Yuxin Yin
-
Article
| Open Access Recurring RNA structural motifs underlie the mechanics of L1 stalk movement
Recurring RNA structural motifs underlie the mechanics of L1 stalk movement
Translocation of the tRNA on the ribosome is associated with large-scale molecular movements of the ribosomal L1 stalk. Here the authors identify the key determinants that allow these dramatic movements, and suggest they represent general strategies used to enable large-scale motions in functional RNAs.
- Srividya Mohan
- & Harry F Noller
-
Article
| Open Access ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
Nonsense-mediated mRNA decay (NMD) is a quality control pathway that recognizes and degrades transcripts harbouring nonsense mutations. Here the authors show that the ATPase activity of UPF1 mediates functional interactions between the NMD machinery and ribosomes required for efficient ribosome release at premature termination codons.
- Lucas D. Serdar
- , DaJuan L. Whiteside
- & Kristian E. Baker
-
Article
| Open Access Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
mRNA surveillance is essential to maintain homeostasis in eukaryotes and is activated by mRNAs lacking a stop codon. Here the authors describe a high resolution cryo-EM structure of a nonstop complex that shows how arrested ribosome recognition is achieved during Dom34-mediated mRNA surveillance.
- Tarek Hilal
- , Hiroshi Yamamoto
- & Christian M.T. Spahn
-
Article
| Open Access Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
The correct folding of proteins often requires the intervention molecular chaperones, which can occur co-translationally. Here the authors identify elements of yeast Ssb (Hsp70) that mediate ribosomal binding, and suggest a mechanism that directs efficient interaction of Ssb with the nascent chain.
- Marie A. Hanebuth
- , Roman Kityk
- & Elke Deuerling
-
Article
| Open Access Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
In yeast, the heterodimeric ribosome-associated complex (RAC) acts in concert with the Hsp70 protein Ssb, forming a unique chaperone triad. Here the authors use structural and biochemical approaches to shed light on how translation and folding are coupled in eukaryotes.
- Andrea Gumiero
- , Charlotte Conz
- & Irmgard Sinning
-
Article
| Open Access CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
Epithelial-to-mesenchymal transition is a key process in tumorigenesis but little is known about the molecular mechanism regulating such process at the translational level. Here, the authors identify a subset of mRNAs important for this process that are specifically modulated by the RNA-binding protein CELF1.
- Arindam Chaudhury
- , Shebna Cheema
- & Joel R. Neilson
-
Article
| Open Access Structure of the ribosome post-recycling complex probed by chemical cross-linking and mass spectrometry
Structure of the ribosome post-recycling complex probed by chemical cross-linking and mass spectrometry
Ribosome recycling orchestrated by ABCE1 connects translation termination and mRNA surveillance mechanisms with re-initiation. Using a cross-linking and mass spectrometry approach, Kiosze-Becker et al. provide new information on the large conformational rearrangements that occur during ribosome recycling.
- Kristin Kiosze-Becker
- , Alessandro Ori
- & Robert Tampé
-
Article
| Open Access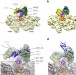 Cryo-EM study of start codon selection during archaeal translation initiation
Cryo-EM study of start codon selection during archaeal translation initiation
Initiation factor eIF2, common to eukaryotes and archaea, is a central actor in translation initiation. Here the authors describe two cryo-EM structures of archaeal 30S initiation complexes that provide a novel view of the central role that e/aIF2 plays in start codon selection.
- Pierre-Damien Coureux
- , Christine Lazennec-Schurdevin
- & Yves Mechulam
-
Article
| Open Access How EF-Tu can contribute to efficient proofreading of aa-tRNA by the ribosome
How EF-Tu can contribute to efficient proofreading of aa-tRNA by the ribosome
The translation of mRNA by the ribosome is governed by a series of large-scale conformational transitions. Here the authors use MD simulations to demonstrate how the rate of dissociation of elongation factor Tu affects the dynamics of tRNA accommodation and proofreading.
- Jeffrey K. Noel
- & Paul C. Whitford

 Structural snapshots of human pre-60S ribosomal particles before and after nuclear export
Structural snapshots of human pre-60S ribosomal particles before and after nuclear export