Abstract
N6-methyladenosine (m6A) is an abundant, dynamic mRNA modification that regulates key steps of cellular mRNA metabolism. m6A in the mRNA coding regions inhibits translation elongation. Here, we show how m6A modulates decoding in the bacterial translation system using a combination of rapid kinetics, smFRET and single-particle cryo-EM. We show that, while the modification does not impair the initial binding of aminoacyl-tRNA to the ribosome, in the presence of m6A fewer ribosomes complete the decoding process due to the lower stability of the complexes and enhanced tRNA drop-off. The mRNA codon adopts a π-stacked codon conformation that is remodeled upon aminoacyl-tRNA binding. m6A does not exclude canonical codon-anticodon geometry, but favors alternative more dynamic conformations that are rejected by the ribosome. These results highlight how modifications outside the Watson-Crick edge can still interfere with codon-anticodon base pairing and complex recognition by the ribosome, thereby modulating the translational efficiency of modified mRNAs.
Similar content being viewed by others
Introduction
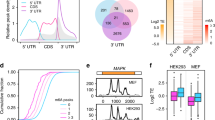
Post-transcriptional modifications of messenger RNA (mRNA) modulate key steps in mRNA metabolism, including mRNA stability and localization1, nuclear export2, exon-intron architecture3, and alternative splicing4. N6-methyladenosine (m6A) is one of the most abundant internal mRNA modification5 found in both coding and untranslated regions of mRNAs in all three domains of life with an average of three m6A per mRNA transcript6,7. The modification sites are located within a conserved consensus sequence of RRACH (R = A/G, H = U/A/C; UGCCAG in Escherichia coli), recognized by dedicated m6A methytransferases (writers) and demethylases (erasers)8,9. The activity of the catalytic subunit m6A-METTL writer complex (MAC) is stimulated by the regulatory subunit m6A-METTL-associated complex (MACOM)10. Reader proteins (e.g. eIF3, YTH domain proteins, HNRNP proteins in eukaryotes) recognize m6A and its consensus sequence, imparting a highly localized and regulated action11,12. The prevalence and conservation of m6A modification and of the machinery that installs, reads, and removes the modification suggests its important role in regulating mRNA dynamics and gene expression at the posttranscriptional level. Indeed, recent studies link m6A modification to a wide range of important biological processes such as neural development and differentiation13, spermatogenesis14, cell growth and cancer15. However, in most cases the exact regulatory pathway is unknown.
The majority of m6A sites are mapped within mRNA coding regions5,6,16, where the modification can modulate codon reading by aminoacyl-tRNA (aa-tRNA) in the A site of the ribosome. Most of the work on the mechanism by which m6A affects decoding has been carried out in the E. coli translation system; given the evolutionary conservation of the decoding mechanism17,18, similar mechanisms are likely to operate in bacteria and eukaryotes. Although the modification is not located on the Watson–Crick edge of the nucleotide, it decreases translation efficiency19, slows down the decoding process20, and alters the ribosome selectivity for the cognate aa-tRNA (i.e. fully matching the codon)20,21. Single-molecule experiments using Förster Resonance Energy Transfer (smFRET) that measure the duration of the decoding step (up to and including the peptide bond formation step) and tRNA–mRNA translocation showed that decoding is slowed down by m6A modification at the first position of the AAA codon (m6AAA) (15-fold) and Am6AA (8-fold) and to a smaller extent by AAm6A, Cm6AG and CCm6A modification (2.5-fold)20. Rapid kinetics measurements of GTP hydrolysis suggested that m6A reduced the initial selection capacity of the ribosome20,21. In addition, m6AAA caused a 1.5-fold excess hydrolysis of GTP per peptidyl transfer reaction, indicating that m6A induces excessive proofreading of cognate aa-tRNA20. Extrapolations of the kcat/KM values led the authors to suggest that m6A dramatically (10-fold) decreases the association rate (ka) of the ternary complex EF-Tu–GTP–aa-tRNA (TC) to the ribosome prior to codon recognition21. The postulated effect would imply that the modification induces a conformational state of the ribosome that is refractory to TC binding; however, structural data in support of this hypothesis is lacking. Thus, while experiments clearly show that the m6A modification has an effect on translation, the exact mechanism of how m6A regulates individual steps of the decoding cycle remain elusive.
In this study, we systematically analyze the effect of m6A at all three AAA codon positions on different steps of the decoding cycle by Lys-tRNALys using a combination of ensemble kinetics, smFRET, and single-particle cryo-EM techniques. In contrast to previous suggestions, we show that the rates of forward reactions of initial binding and codon recognition are largely unaffected, whereas the respective dissociation steps are faster in the presence of the modification, which explains the lower effective rate of GTP hydrolysis. Furthermore, the presence of m6A increases the transition time from codon recognition to post-decoding phase, which—together with an increased tRNA drop-off – results in a significantly fewer ribosomes reaching the translocation step. We show that the m6A effect depends on the position, the nature of the codon and on the chemical modifications of the tRNA anticodon. High-resolution cryo-EM structures reveal how the ribosome accommodates m6A modifications at different codon positions and suggest that the AAA codon adopts a structured conformation in the A site independently of the modification. In summary, the most dramatic effect of m6A modification appears to involve destabilization of the codon-anticodon interaction, which results in tRNA drop-off at all stages of decoding and explains the inhibitory effect of the modification on translation.
Results
Effect of m6A on the elemental steps of decoding
We monitored the consecutive decoding steps of an AAA codon modified at any of the three codon position by its cognate Lys-tRNALys using a fully-reconstituted in vitro translation system from E. coli22. Using the AAA codon enables to monitor the codon position effect of m6A within a single codon. In the first step of decoding (Fig. 1a), the ternary complex EF-Tu–GTP–Lys-tRNALys (TC) binds to the ribosome in a codon-independent manner (characterized by the rate constants k1 and k−1), thereby initiating codon reading17. In the next step, codon recognition (k2 and k−2) induces domain closure of the SSU and triggers GTPase activation of EF-Tu (k3) upon EF-Tu docking at the GTPase-activating center in the large ribosomal subunit (LSU), resulting in GTP hydrolysis (kGTP)17,23,24,25. The release of the reaction product Pi (k4;26) induces the transition of EF-Tu from the GTP- to the GDP-bound conformation, allowing aa-tRNA accommodation in the A site of the LSU (k5) and peptide bond formation (kpep), whereas EF-Tu is released from the ribosome (k6). Alternatively, aa-tRNA can by rejected from the ribosome (k7)17. Additional rejection steps, for example after GTP hydrolysis but prior to dissociation of aa-tRNA from EF-Tu are likely24, but are accounted for in the global rejection rate k7. Notably, all rejection rates are very low for the cognate Lys-tRNALys (22), which results in the efficient decoding and peptide bond formation.
a Kinetic scheme of decoding. SSU, yellow; LSU, light blue. E, P, and A indicate the tRNA binding sites of the ribosome22,27. b Fluorescence change reporting codon-independent initial binding (IB) of TC to the ribosome. The A-site codon AAA is unmodified (black), m6AAA (red), Am6AA (green) or AAm6A (blue). Each time course is an average of 5-6 technical replicates. The kapp values from exponential fitting are 70 ± 1 s−1 (AAA), 75 ± 1 s−1 (m6AAA), 65 ± 1 s−1 (Am6AA), and 61 ± 1 s−1 (AAm6A) for the major phase (80% of the fluorescence change). Control was obtained by mixing TC with buffer (gray). c Fluorescence change of EF-Tu–GTP–[14C]Lys-tRNALys(Prf16/17) reporting A-site codon recognition (CR). Each time course is an average of 5-6 technical replicates. The kapp values of the major reaction (fluorescence increase) are 33 ± 1 s−1 (AAA), 25 ± 1 s−1 (m6AAA), 32 ± 1 s−1 (Am6AA), and 37 ± 1 s−1 (AAm6A). Controls were obtained by mixing TC with buffer (gray). d Reaction times (τ) of GTP hydrolysis (τGTP, open bars) and peptide bond formation (τpep, closed bars) from exponential fitting of the respective time courses (Supplementary Fig. 1a). Error bars indicate SEM of the fit generated from three independent experiments (N = 3). P values are calculated using two-tailed unpaired Student’s t-test in comparison to unmodified AAA codon (*P < 0.05, **P < 0.005). For τGTP, P = 0.0011 for m6AAA, P = 0.0496 for Am6AA and P = 0.0419 for AAm6A. For τpep, P = 0.0019 for m6AAA, P = 0.0008 for Am6AA and P = 0.0023 for AAm6A. For all panels of Fig. 1, TC (Lys or Phe) (0.3 µM) was mixed with IC (0.9 µM). Source data are provided as a Source Data file.
To quantify the m6A effect, we prepared ribosome initiation complexes (IC) with f[3H]Met-tRNAfMet bound to the AUG codon in the P site and an AAA (or m6AAA, Am6AA, or AAm6A) codon in the A site. The IC was mixed with TC that contained reporters suitable to monitor each step of decoding using ensemble rapid kinetics22,27. To test the effect of m6A modification on the initial binding step prior to codon recognition, we used an established assay with a fluorescence reporter group proflavin attached to the tRNAPhe elbow region (position 16/17)28. The reaction does not proceed beyond initial binding, because tRNAPhe (anticodon AAG) does not match the AAA codon in the A site. Interaction of EF-Tu–GTP–Phe-tRNAPhe(Prf16/17) with the IC leads to a rapid fluorescence increase that is largely independent of the mRNA modification, with the apparent rate constant kapp1 in the range of 60–75 s−1 and very similar endpoints for all four codons (Fig. 1b). Although the apparent rate constant includes both the association (k1[IC]) and dissociation (k−1) terms, these results exclude the possibility that the association rate constant (k1 or ka) is decreased 10-fold by m6A modification as previously suggested21. Rather, the effect of the modification on the initial binding is small, if any.
Next, we monitored the codon recognition step using Lys-tRNALys labeled by proflavin at positions 16/17 (tRNALys(Prf16/17))22. After initial binding to the IC, Lys-tRNALys recognizes its cognate AAA codon, leading to a fluorescence increase when EF-Tu–GTP–[14C]Lys-tRNALys(Prf16/17) is mixed with IC (Fig. 1c). The apparent rate of codon recognition (kapp2) is essentially independent of the m6A modification, showing only a small variations from approximately 33 s−1 with AAA to 25-37 s−1 with m6A-containing codons and a minor effect on the amplitude of the signal change, consistent with a notion that the effect of m6A on initial binding and codon recognition is small. In contrast, GTP hydrolysis and peptide bond formation are strongly affected (shown as reaction times τGTP and τpep for better comparison; Fig. 1d and Supplementary Fig. 1a), in agreement with previous reports20. With m6AAA codon, τGTP was 8-fold and τpep was 40-fold higher than with AAA, indicating a significant slowing down of the decoding step. The effects of modifications at the 2nd and 3rd position were somewhat lower, decreasing τGTP by about 5-fold (Am6AA) or 2.4-fold (AAm6A) and τpep by 10-fold (Am6AA) or 5-fold (AAm6A) compared to unmodified AAA. We also tested the effect of the m6A modification on translocation using tripeptide formation as a proxy for successful movement of the ribosome along the mRNA (Supplementary Fig. 1b). Ribosome complexes that completed decoding and contain tRNAfMet in the P site and f[3H]Met-[14C]Lys-tRNALys in the A site were mixed with EF-G to induce mRNA–tRNA translocation and a TC with Phe-tRNAPhe that binds to the next codon resulting in synthesis of f[3H]Met-[14C]Lys-Phe (MKF) peptide. Comparison of the time courses of MKF synthesis shows that m6A at the 1st codon position slows down translocation by about 2-fold, whereas the 2nd- and 3rd-position modifications have no significant effect (Supplementary Fig. 1b).
Context effects
To test whether the codon position in the mRNA attenuates the m6A effect, we compared the rates of AAA and m6AAA decoding at the 2nd (Lys2) and 4th (Lys4) positions in the mRNA. In the latter case, we translated a 4-codon sequence for Met-Val-Phe-Lys (MVFK) with unmodified AAA or m6AAA codon. We observe a 10-fold decrease in the kpep value of MVFK formation with m6AAA relative to AAA codon, compared to the 40-fold effect observed when the codons are in the 2nd position (Fig. 2a). The m6A effect on Lys4 is very similar to that observed by measuring ribosome rotation (15-fold)20, which is a proxy for peptide bond formation. However, taking into account that incorporation of Lys4 is slower than of Lys2 due to additional time needed for MVF synthesis, the actual delay in decoding of the modified Lys4 codon (approximately 16 s; Fig. 2a) is even longer than on Lys2 (3 s; Fig. 1d). These results suggest that the m6A effect can be attenuated by the mRNA context, in agreement with the previous results20.
The coding sequences show the unmodified and first-position modified A-site AAA or ACC codon (bold). a Time courses of f[3H]Met-Val-Phe-[14C]Lys (Lys4 mRNA, open circles) and f[3H]Met-[14C]Lys (Lys2 mRNA, closed circles) formation obtained by mixing IC (0.9 μM) with the AAA (black) or m6AAA (red) with TC (0.3 μM) containing Val-tRNAVal, Phe-tRNAPhe, and [14C]Lys-tRNALys and EF-G (4 μM) or [14C]Lys-tRNALys respectively. Smooth black lines represent one-exponential fits. The reaction time τ calculated from exponential fitting is 0.08 s for AAA vs 3.3 s for m6AAA at Lys2 and 2 s for AAA vs 18 s for m6AAA at Lys4. Thus, the delay in decoding due to the modification is ~3 s at Lys2 and ~16 s for Lys4. b Dipeptide formation with Lys TC or Thr TC (0.3 μM) upon binding to IC (0.9 μM). f[3H]Met-[14C]Lys (closed circles) or f[3H]Met-[14C]Thr (open circles) dipeptides were separated by HPLC and quantified by scintillation counting. For both panels of this figure, each time course is the mean of three independent experiments with error bars representing standard deviation (N = 3). Source data are provided as a Source Data file.
m6A causes translation delays not only on modified AAA, but also on Cm6AG (coding for Gln) and CCm6A (Pro) codons20, whereas the effect on the ACC (Thr) codon was not tested. Notably, ACC is part of the RRACH consensus sequence that is likely to be methylated in vivo. We measured the effect of m6A on the kinetics of dipeptide formation (MK or MT) with IC programmed by the respective mRNAs and TC (Lys or Thr). Our results show that the effect on the ACC codon is smaller than on AAA, 5-fold vs. 40-fold, respectively (Fig. 2b). Thus, also the codon/amino acid identity modulates the translation delay caused by the m6A modification.
Interplay with tRNA modifications
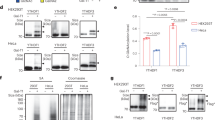
In the cell, not only mRNA, but also tRNAs can be modified, which prompted us to test how modifications at the tRNA anticodon affect reading of m6A-modified codons. We made use of the known detailed kinetic mechanism of AAA decoding by yeast tRNALys containing mcm5s2 modification at U34 and the well-documented contribution of the s2U34 to decoding22. The mcm5s2 modification at U34 is introduced by the ELP29 and URM1 pathways30. Deletion of the URM1 gene in yeast results in synthesis of Δs2U34 tRNALys, which carries all other modifications but lacks s2U3431, as verified by ((N-acrylolamino)phenyl)mercuric chloride (APM) gel retardation of tRNA (Methods). The hypomodified tRNA shows an increased propensity to dissociate from the ribosome (manifested in the increased k−2 and k7 dissociation rate constants), slower rearrangement after GTP hydrolysis (k4) and slower accommodation into the peptidyl transferase center (k5)22. To test how a fully modified mcm5s2 Lys-tRNALys and Δs2U34 Lys-tRNALys affect m6A decoding, we monitored fMet-[14C]Lys formation with IC containing a unmodified or m6-modified AAA codon in the A site (Fig. 3). The time course of peptide bond formation with the fully modified mcm5s2 Lys-tRNALys on the AAA codon is biphasic (Fig. 3a), which reflects heterogeneous pathways towards aa-tRNA accommodation depending on the timing of its release from EF-Tu (k5a and k5b, respectively)22. Removal of the s2U34 modification dramatically reduces the rate and end level of the reaction (Fig. 3), in agreement with the earlier report22. With m6A modification at any codon position, the rate of peptide bond formation with mcm5s2 Lys-tRNALys is very low, 30–80-fold lower than on an unmodified AAA codon (Fig. 3). Furthermore, the end level is reduced, indicating that a large fraction of Lys-tRNALys dissociates from the ribosome before entering the reaction. The effect is even more dramatic when the m6A modification on the mRNA is combined with the Δs2U34 Lys-tRNALys, in particular on m6AAA and Am6AA codons. These data suggest that m6A modification enhances rejection of mcm5s2 Lys-tRNALys and the lack of s2U34 tRNA modification further destabilizes the complexes, thereby practically abolishing translation.
Time courses of fMet-[14C]Lys formation obtained by mixing TC containing yeast [14C]Lys-tRNALys (0.3 μM) with IC (0.9 μM). [14C]Lys-tRNALys is fully modified with mcm5s2U34 (closed circles) or hypomodified containing mcm5U34 (open circles). Each curve is the mean of three independent experiments with error bars representing standard deviation (N = 3). Smooth black lines represent single or double exponential fits. fMet-[14C]Lys dipeptides were separated by HPLC and quantified by scintillation counting. The τpep values of the fast reaction with fully modified mcm5s2U34 [14C]Lys-tRNALys are approximately 10–20 s−1 irrespective of the modification, the reactions with mcm5U34 tRNA are too inefficient to obtain reliable values. Source data are provided as a Source Data file.
The mechanism of decoding inhibition by m6A
While there is a clear effect of m6 modification on GTP hydrolysis and dipeptide formation, the model explaining these effects by a slower association rate of the TC to the ribosome21 is clearly not supported by our experimental data. This prompted us to examine the mechanism of decoding in more detail using smFRET approaches32,33,34,35 comparing AAA and m6AAA codons, which gave the largest rate differences. smFRET experiments are designed to monitor binding and unbinding of TC to and from the ribosome, as well as movement of aa-tRNA through the ribosome, accommodation and the dynamics of the peptidyl-tRNA after peptide bond formation32,33. TC binding to the ribosome was monitored by smFRET between the ribosomal protein L11 labeled with FRET donor Cy3 (L11-Cy3) and Lys-tRNALys labeled with FRET acceptor Cy5 (Lys-tRNALys(Cy5)) (Fig. 4a). L11-Cy3 IC was immobilized on coverslips through 5′-biotinylated mRNAs carrying AAA or m6AAA codons in the A site. After injection of TC with Lys-tRNALys(Cy5) into the flow chamber, we observed four types of traces. The percentages of ribosomes following each of type of trajectories are strikingly different for AAA and m6AAA (Fig. 4b). Traces that show rapid (>30 s−1) appearance of FRET signal followed by a rapid loss of FRET reflect initial binding complex formation and dissociation prior to codon recognition (Fig. 4b–d and Supplementary Fig. 2a). Such traces comprise only a small percentage of complexes on the AAA codon (8 ± 2%), but increase to 37 ± 1% (P = 0.0015) on m6AAA. The dissociation rate (k−1) of these traces is higher with m6AAA, indicating that m6A promotes rejection of ternary complexes from the initial binding complex (Fig. 4b, Supplementary Fig. 2b and Table 1).
a Schematic of the smFRET experiment. 70S IC (light gray) is labeled with Cy3 (green star) at the protein L11 and Lys-tRNALys is labeled with Cy5 (red star). b Relative abundance of the different types of traces during decoding of AAA (black) or m6AAA (gray) codon. “Rejected” denotes the fraction of complexes that dissociate from the ribosome after initial binding (IB; c, d). “Decoding” pertains to traces that show the characteristic codon reading (CR) phase followed by accommodation and ensuing fluctuations between A/A, A/P, and A/P* states (e, f). “Post-decoding” are the complexes in which Lys-tRNALys is found in an A/A, A/P or A/P* state at the start of the recording (g, h). “Stalled” denotes the fraction of ribosomes that do not progress past the CR state (Supplementary Fig. 2). Shown are the mean values with error bars representing standard deviation from N = 3 independent experiments. ** indicates statistical significance with P < 0.01 using a two-tailed Welch’s t-test without adjustment for multiple comparisons. P = 0.0015 for rejected traces, P = 0.0038 for post-decoding traces, P = 0.0565 for decoding traces. Open circles are the values in the individual experiments. c Contour plot showing the distribution of FRET values after synchronization during initial binding and dissociation of Lys-tRNALys(Cy5) from ICs carrying the AAA codon in the A site. Traces are synchronized to the first transition with FRET > 0. N, number of traces. d same as c on the m6AAA codon. e Contour plot showing the distribution of FRET values during the full decoding trajectory on the AAA codon. FRET 0.96 ± 0.03 in the CR state derived from mean ± s.d. of the µ values of the Gaussian fitting of N = 3 independent experiments. f same as e on the m6AAA codon, FRET 0.98 ± 0.01. g Contour plot showing the distribution of FRET values in the post-decoding complexes carrying fMet-Lys-tRNALys(Cy5) on the AAA codon. FRET population distribution reveals two states with FRET 0.79 ± 0.01 (A/A, A/P) and 0.58 ± 0.02 (A/P*)35,64,65,66. h Same as g on m6AAA codon. A/A, A/P, FRET 0.80 ± 0.01; A/P*, FRET 0.61 ± 0.01. Data are from N = 3 independent experiments. Source data are provided as a Source Data file.
Two other types of traces correspond to ribosome complexes where tRNA is successfully accommodated in the A site. In those traces that show a full decoding trajectory, rapid (>30 s−1) codon reading results in appearance of high FRET 0.9, followed by the progression to lower FRET states (step assignment is from33). The latter corresponds to tRNA movement from codon reading towards the accommodation in the A site and peptide bond formation. After peptide bond formation, fMet-Lys-tRNALys(Cy5) starts to fluctuate between A/A, A/P (FRET 0.8) and A/P* (FRET 0.6) states (Fig. 4e, f and Supplementary Fig. 2c). The percentage and the decay rate of the codon reading state (kCR→forward) are not strongly affected by the m6 modification (Fig. 4b, Supplementary Fig. 2d and Table 1), suggesting that the codon reading state does not accumulate during the reaction. However, the transition rate kCR→A/P* from the codon reading (FRET 0.9) to the post-decoding A/P* state (FRET 0.6) is reduced 2-fold in the presence of m6A (Supplementary Fig. 2c, d and Table 1).
A significant fraction of ribosome complexes reached the post-decoding state already at the starting point of imaging, with fMet-Lys-tRNALys(Cy5) fluctuating between FRET 0.8 (A/A, A/P) and 0.6 (A/P*) states (Fig. 4g, h and Supplementary Fig. 2e). Here, we observe no significant differences in the transition rates between A/A, A/P (FRET 0.8) and A/P* (FRET 0.6) states (k0.8→0.6 and k0.6→0.8) in the presence of m6A, indicating that m6A has no effect on the tRNALys dynamics in classical and hybrid states (Supplementary Fig. 2f and Table 1). However, the percentage of ribosomes showing classical to hybrid fluctuations is reduced in the presence of m6A (P = 0.0038, Fig. 4b), consistent with the notion that a fraction of ribosomes has lost the tRNA before entering the post-decoding phase. Lastly, we also observed a small fraction of traces that did not progress beyond the codon reading state, but the percentage of these traces was not affected by the presence of m6A (Fig. 4b and Supplementary Fig. 2g–i). In conclusion, m6A increases the dissociation of TC during initial binding and codon reading, as well as delays the transition to the post-decoding state after GTP hydrolysis.
To further investigate the effect of m6A on the stability of codon-anticodon duplex in the A site, we repeated the experiments using EF-Tu(H84A), a GTPase-deficient EF-Tu mutant (Fig. 5a). Steps prior to GTP hydrolysis, i.e. initial binding, codon recognition and GTPase activation, are not affected by the mutation, but GTP hydrolysis and the subsequent steps of tRNA accommodation and peptide bond formation are abolished36. In the presence of EF-Tu(H84A), the TC stalled in the codon reading state represents the majority of the population (78 ± 3% for AAA and 72 ± 1% for m6AAA) (Fig. 5b), in agreement with previous studies using non-hydrolysable GTP analogs20,33. For both AAA and m6AAA, FRET population distribution revealed a single FRET 0.9 codon reading state that did not progress towards accommodation (Fig. 5c,d). The decay rate of the long-lived codon reading state reports the dissociation of the ternary complex from the ribosome and acts as a proxy for codon-anticodon stability22,27. In the presence of m6A, the decay rate (k−2) is 3.2-fold higher that on the unmodified AAA codon (Fig. 5d, inset), indicating that m6A destabilizes the codon-anticodon interactions during codon reading prior to GTP hydrolysis. This leads to rapid dissociation from the ribosome and thus further accounts for low efficiency of tRNA accommodation.
a Schematic of the smFRET experiment monitoring decoding attempts by the GTPase-deficient TC. CR, codon reading. b Relative abundance of smFRET traces during decoding of AAA (black) or m6AAA (gray) codons by EF-Tu(H84A). Shown are the mean values and error bars represent standard deviation from N = 2 independent experiments. Open circles are the values in the individual experiments. c Contour plot showing the distribution of FRET values after synchronization during CR (FRET 0.97 ± 0.01) without progression to accommodation of Lys-tRNALys(Cy5) on AAA codon. Data are from N = 2 independent experiments. d Same as c for the m6AAA codon (FRET 0.96 ± 0.01). Inset shows normalized dwell-time histograms of EF-Tu(H84A)–GTP–Lys-tRNALys(Cy5) dissociation (k−2) during decoding of AAA (closed circles) and m6AAA (open circles) codon (Table 1). Normalization was performed by division by the maximum value of the histogram. Source data are provided as a Source Data file.
Cryo-EM captures dipeptidyl-tRNALys bound to m6A-modified codons
Next, we obtained the structures of ribosome complexes with tRNALys in the A site using single-particle cryo-EM. The complexes were prepared by mixing EF-Tu–GTP–Lys-tRNALys with IC containing fMet-tRNAfMet in the P site and mRNAs with either AAA, m6AAA, Am6AA, or AAm6A in the A site. We collected four equivalent datasets (Supplementary Table 1) and sorted the particles according to the tRNA occupancy of the E, P, and A sites (Supplementary Fig. 3). In addition to a fraction of unreacted IC, we found ribosomes that completed the decoding process and contained tRNAfMet in the P site and fMet-Lys-tRNALys in the A site. The latter complexes could be further classified into (i) those where tRNAs were in classical (P/P, A/A) states with ribosomes in a non-rotated conformation and (ii) those with tRNAs in hybrid (P/E, A/P) states with the ribosomal subunit rotated relative to each other. The particle distribution was dramatically different between samples with unmodified AAA and m6A-modified codons (Fig. 6a). Specifically, 48% (m6AAA), 52% (Am6AA), or 46% (AAm6A) of ribosomes failed to progress from the IC towards decoding and peptide bond formation, compared to only 4% of ICs with unmodified AAA codon. These differences support the notion that a sizable fraction of Lys-tRNALys that attempts to bind to its cognate codon is rejected during the decoding process.
a From top to bottom: AAA, m6AAA, Am6AA, AAm6A (top left: schematic view, right: map colored by the RNA chain. LSU is shown in light blue, SSU in yellow, mRNA in red, P-site tRNA in cyan and A-site tRNA in dark blue). The pie charts show relative particle distributions of different states. Lys codons carrying an m6A modification lead to a lower proportion of ribosomes progressing towards decoding (orange slice). The maps are shown as surface representation. Top middle panel, close-up view of the P-site and A-site tRNAs and mRNA. Top right panel, close-up view of mRNA–tRNA interactions. b Base pairing at the 1st, 2nd and 3rd codon position. Left, AAA codon; right, the respective m6A-modified pair. The m6 methyl group (magenta) is shown in an anti conformation. The measured distances between the conventional hydrogen bond donor and acceptor indicated with a dashed line are expressed in Å. The maps are shown as mesh with contour rmsLevel 4.
The particles with tRNAs in classical states were used to reconstruct maps at 2.3 Å (AAA), 2.9 Å (m6AAA), 2.8 Å (Am6AA), and 2.6 Å (AAm6A) global resolutions (GSFSC with 0.143 cut-off; Fig. 6a), respectively. The high quality of the maps in the proximity of the decoding region (Supplementary Fig. 4) allowed the direct visualization of the m6A modification at different codon positions (Supplementary Fig. 5). Furthermore, the fMet-Lys moieties in the peptidyl transferase center (Supplementary Fig. 6), as well as all known tRNAfMet and tRNALys modifications (e.g. Cm32, mnm5U34) could be assigned. However, detailed comparisons did not reveal any pronounced structural differences between the models with and without m6A (Fig. 6b). Of note, we positioned the m6A in the anti conformation, which enables hydrogen bonding in all three codon-anticodon pairs. Even at the high resolution obtained, the cryo-EM densities do not unambiguously exclude the syn conformation. However, fitting m6A in a syn conformation precludes hydrogen bonding and results in an increased distance of the N6 atom to the O4 atom from the cognate U in the codon by ~0.5 Å (Supplementary Fig. 7). Thus, once the A-site aa-tRNA is bound, we cannot detect any major structural differences between unmodified and m6A-modified codons.
The AAA codon is structured in an unoccupied A site
To further elucidate the structural basis of slower translational rates of ribosomes decoding the m6A-modified Lys codon, we separately analyzed ICs from the same datasets and reconstructed maps at 2.4 Å (m6AAA), 2.2 Å (Am6AA), and 2.1 Å (AAm6A) overall resolution (Supplementary Fig. 8). Because the dataset for the IC with unmodified AAA codon was too small to obtain a structure at a similar resolution range (Fig. 7a), we collected two additional IC datasets with either an AAA or AAm6A codon in the A site (Fig. 7b). The complexes were prepared in the same way as described above, but without adding Lys-tRNALys; the final overall resolution of both structures was 2.0 Å (Supplementary Fig. 9 and 10). Despite the high quality of the maps, particularly in the decoding region (with local resolution between 2.4–2.7 Å), the density for the m6 modification was not well resolved (Supplementary Fig. 11). Surprisingly, in all structures the AAA codon was in a π-stacking conformation (Fig. 7c), which—to our knowledge—was not identified with any other codon sequence in the well-resolved IC structures available so far. The π-stacking was stabilized by the interactions with the decoding center of the ribosome, in particular 16S rRNA nucleotides A1493 and C1397. Residue A1493 in helix 44 (h44) flips out of its helical arrangement and is involved in stacking with A1 of the AAA codon. The stacking array extends beyond the A-site codon to the next nucleotide (U4) from the downstream codon UUC and with C1397 of 16S rRNA. Another key residue in the decoding center, A1492, remains stacked within an internal loop of h44 of the IC, whereas G530 is pushed away from the mRNA and does not appear to be engaged in the codon stabilization (Fig. 7c).
a From top to bottom: AAA, m6AAA, Am6AA, AAm6A. Left panel: schematic view; right panel: map colored by the RNA chain. LSU is shown in blue, SSU in yellow, mRNA in red, P-site tRNA in cyan. The A-site codon is indicated with dashed circles. b Structures of unreacted ICs (i.e. prepared without addition of Lys-tRNALys, see text) with AAA mRNA (left) and Am6AA mRNA (right). Maps are shown in surface representation. c Close-up view of interactions between AAA or AAm6A A-site codon (red), and rRNA (yellow) in the decoding region of the unreacted ICs (top). The corresponding models are also shown from different perspectives (bottom). In the unreacted AAA (left) and AAm6A ICs (middle) the nucleotides form a π-stacking array, whereas the equivalent nucleotides in the post-decoding complex (P/P A/A) appear unstacked (right). d Repositioning of the 16S rRNA nucleotides A1493, A1492, G530 and C1397. The conformational rearrangements during the transition from IC (yellow) to the post-decoding complex (P/P A/A; gray) are shown in reference to the mRNA codon.
Comparison between the unpaired AAA codon in IC and the post-decoding complex with fMet-Lys-tRNALys in the A site shows that the positioning of the 16S rRNA nucleotides C1397, A1492, A1493, and G530 changes between the two states (Fig. 7d). Upon decoding, the base array of A1-U4 mRNA becomes unstacked. The C1397 base rotates by approximately 90° with respect to the U4. While, in the IC, A1493 is involved in the stabilization of the A-site codon, in the decoding complex, A1492, A1493, and G530 monitor the correct Watson–Crick geometry of the codon-anticodon complex in the A site, consistent with previous findings37,38. A1493 and A1492 are flipped-out of the 16S rRNA stacking conformation and together with G530 interact with the minor grove of the codon-anticodon mini helix. Of note, the positions of these rRNA and mRNA nucleotides are identical in the m6A-modified and the AAA datasets (Fig. 7c), showing that m6A does not affect monitoring of the codon-anticodon complex by the ribosome.
Discussion
The results of this work, together with previous reports20,21, show that m6A modification on AAA and other A-containing codons, such as Glu, Gln, Pro, and Thr, leads to inefficient decoding in a codon context-dependent manner. We show that, in contrast to previous suggestions21, m6A does not affect the association rate of the TC to the ribosome and provide detailed insights into how m6A modifications modulate the decoding process during translation elongation. Prior to decoding, the AAA codon adopts a π-stacking arrangement involving the 3’-adjacent U in the mRNA and stabilized by A1493 and C1397 of 16S rRNA (Fig. 7). While the codon-independent initial binding of the cognate TC-Lys to the AAA-programmed ribosomes is rapid and independent of the m6A modification (Figs. 1 and 3), the resulting complexes tend to dissociate more rapidly when the modification is present (Table 1; Figs. 4 and 5). As a result, fewer ribosomes proceed towards rapid codon recognition (Fig. 4). The presence of m6A reduces the stability of the codon recognition complex (Table 1) and slows down GTP hydrolysis and accommodation compared to the unmodified codon (Fig. 1 and refs. 20,21). The delays in the forward decoding steps together with the increased tRNALys drop-off from the ribosome result in fewer ribosomes that complete decoding (Figs. 4 and 6). On those ribosomes that reached the post-decoding state, the π-stacking arrangement of the codon is resolved, and the codon-anticodon complex adopts a canonical Watson-Crick geometry stabilized by the residues of 16S rRNA independent of m6A modification (Fig. 6).
Comparison of the structural and biophysical data provides insights into the dynamics of AAA decoding and the role of the m6A modification. The observed π-stacking codon arrangement is similar to that found in yeast ribosomes and is stabilized by homologous nucleotides (Supplementary Fig. 12 and refs. 39,40), indicating evolutionary conservation of the recognition mechanism. However, our data show that a single unmodified AAA codon is efficiently decoded despite the unfavorable codon conformation (Figs. 1 and 6), indicating that the structuring of the A-site codon alone cannot explain translational stalling on extended poly(A) tracks, as previously suggested39,40. Also the presence of an AAAA sequence in the yeast structure instead of an AAAU in our structures cannot explain the difference, as AAAAAA sequences are translated efficiently39. Concerning the effect of the m6A modification, previous NMR data suggested that m6A stabilizes stacking compared to the unmodified base41. A stronger codon stacking in a conformation unfavorable for codon reading may explain why a sizable fraction of TCs that scan the ribosome during the initial binding and codon reading are rapidly rejected from the ribosome prior to forming a codon-anticodon complex, as indicated by smFRET experiments (Fig. 4b–d).
The rates of the codon recognition on those ribosomes that proceed towards decoding are not much changed by the m6A modification (Fig. 1c). This observation favors the conformational selection model, which assumes that the π-stacking and unstacked arrangements are in dynamic equilibrium on the ribosome. If the TC attempting to read the A site codon encounters the stacked conformation, the respective attempt is likely to fail, resulting in the dissociation of the TC prior to codon recognition. In contrast, when TC encounters an unstacked conformation, it can rapidly form a codon-anticodon complex and proceed towards tRNA accommodation. At this stage, also the switch from syn to anti conformation of the m6 group may play a role, as the optimal codon-anticodon pairing is accomplished by rotating away from its energetically preferred syn geometry on the Watson-Crick face to the higher-energy anti conformation41. The m6 modification is poorly resolved in the cryo-EM structures with the vacant A site, despite the high quality of the 70S IC structures (Fig. 7a, b). Given that the nucleotides are clearly resolved, the weak density for the modification (Supplementary Fig. 11) likely reflects its rotational dynamics. Notably, the density for the additional methyl group is better resolved in the post-decoding complexes (Supplementary Fig. 5) indicating that tRNA binding to the codon restricts m6 dynamics. Fitting the density with m6A in anti conformation yielded a perfect Watson–Crick geometry of base pairing (Supplementary Fig. 7), whereas using the syn conformation results in a shift of the entire nucleotide. The unfavorable anti conformation is partially compensated by hydrogen bonding and Watson–Crick pairing with U resulting in the net destabilization by 0.5–1.7 kcal/mol for a single methyl substitution42. The fluctuations between syn and anti conformation of m6A in combination with the steric hindrance of the codon-anticodon pairing in the energetically more favorable syn conformation may cause the destabilization of the codon-anticodon complex observed in smFRET measurements (Table 1). The altered dynamics of the codon-anticodon complex may also affect the propensity of the SSU to form a closed global conformation that is essential for the activation of GTP hydrolysis25,43,44, which may provide a potential mechanistic explanation for the observed slowing down of GTP hydrolysis.
The accommodation of tRNALys is also affected by m6A, in particular for the m6AAA codon, which is manifested by a further delay of peptide bond formation after GTP hydrolysis (Fig. 1d). smFRET experiments reveal the tendency for a delayed entry into the FRET 0.6 state upon the formation of tRNA hybrid states after peptide bond formation (Table 1). These data suggest that m6A enriches for conformations that enhance tRNA dissociation and disfavor the forward steps towards completion of the decoding step. However, the complexes that completed decoding are found in the canonical tRNA conformation regardless of the modification (Fig. 6). Thus, the ribosome rectifies the codon-anticodon dynamics to select the correct codon-anticodon geometry and the m6A modification affects this process, potentially by stabilizing otherwise unfavorable codon-anticodon conformations. Similarly, the ribosome selectively stabilizes G-U mismatches in a tautomeric form that is unfavorable in solution, but is favored by the ribosome, because it allows pairing with Watson–Crick geometry42,45, thus underscoring the generality of the ribosome rectification mechanism.
The dynamics of the codon-anticodon complex is further modulated by the modifications in the tRNA anticodon (Fig. 3). With yeast tRNALys, the presence of m6A had a strong effect not only on the rate of peptide bond formation, but also on the end level of reaction, which indicates enhanced drop-off of tRNALys from the ribosome during decoding. Similarly to the reaction with E. coli tRNALys, the most dramatic effect was observed with m6AAA and the smallest with AAm6A. Removal of the s2 modification from yeast mcm5s2U34 essentially inhibits the reaction (Fig. 3), underscoring the complex interplay between the dynamics of the codon and the properties of the tRNA anticodon for the outcome of tRNA selection. Taken together, our work combining ensemble kinetics, smFRET and cryo-EM methods provides detailed mechanistic insights into how m6A modification modulates decoding.
Methods
70S ribosomes, EF-Tu and f[3H]Met-tRNAfMet from E. coli were prepared as described in previous studies27,46,47. Unmodified and m6A-modified mRNA was purchased from IDT (Iowa, USA). Native Lys-tRNALys, Lys-tRNALys(Prf16/17) and Phe-tRNAPhe from E. coli and yeast Phe-tRNAPhe(Prf16/17) were prepared as described22,48. Yeast mcm5s2U34 and mcm5U34 Lys-tRNALys were prepared from wild-type S288C and ΔURM1 yeast strains, respectively31, and the s2 modification was quantified as described22. The extent of s2 modification was verified by ((N-acrylolamino)phenyl)mercuric chloride (APM) gel retardation of tRNA and was 70–90% for native tRNALys and 2% for urm1Δ tRNALys. The following mRNAs (unmodified and m6A-modified at different codon positions of AAA or ACC codon) were purchased from IDT (Iowa, USA). The coding region (starting with AUG) is separated from the 5’UTR by space and the m6 residue is introduced in one of the underlined A positions.
5′-GGCAAGGAGGUAAAUA AUGAAAUUCGUUAC-3′
5′-GGCAAGGAGGUAAAUA AUGACCUUCCGCCUCUCUCUC-3′
5′-GGCAAGGAGGUAAAUA AUGGUGUUCAAACUGCGCCUCUCUCUC-3′
For the smFRET experiments, the following 5′-biotinylated mRNA (unmodified and m6A-modified) was used.
5′-Biotin-CAACCUAAAACUUACACACCCGGCAAGGAGGUAAAUA AUGAAAUUC AUUACCUAA-3′
EF-Tu–GTP–[14C]Lys-tRNALys (Prf16/17) (TC-Lys) was prepared by incubating EF-Tu (75 µM), GTP (1 mM), PEP (3 mM), DTT (1 mM), pyruvate kinase (1% v/v), tRNALys(Prf16/17) (15 µM), ATP (3 mM), L-[14C]lysine (22.5 µM) and Lys-tRNA synthetase (2% v/v) in buffer A containing 50 mM Tris-HCl pH 7.5, 70 mM NH4Cl, 7 mM MgCl2 and 1 mM DTT. Assembled TC-Lys was purified by gel filtration on two Superdex 75 HR columns operated in tandem (GE Healthcare) in buffer A. EF-Tu–GTP–[14C]Phe-tRNAPhe(Prf16/17) and EF-Tu–[γ-32P]GTP–[14C]Lys-tRNALys were prepared and purified in similar ways.
IC was prepared from 70S ribosomes (2 µM), mRNAs as indicated (6 µM), f[3H]Met-tRNAfMet (3 µM), IF1, IF2, IF3 (3 µM each), DTT (1 mM) and GTP (1 mM) in buffer TAKM7 (50 mM Tris-HCl pH 7.5, 30 mM KCl, 70 mM NH4Cl and 7 mM MgCl2) for 30 min at 37 °C. IC was purified by centrifugation through 1.1 M sucrose cushion in TAKM7 for 2 h at 4 °C and 259,000 × g in Beckman Optima Max-XP ultracentrifuge. After centrifugation, the pellets were dissolved in TAKM7 and quantified by scintillation counting.
Rapid kinetics
All fluorescence stopped-flow experiments were performed in buffer A at 20 °C. Prf fluorescence was excited at 463 nm and measured after passing through a KV500 long pass filter (Schott). Experiments were performed by mixing equal volumes of IC (0.9 µM) with TC-Lys or TC-Phe (0.3 µM) as indicated and monitoring the time course of fluorescence change. Relative fluorescence was calculated by division of all fluorescence values by the value at time 0.
To monitor GTP hydrolysis, peptide bond formation, and mRNA–tRNA translocation following peptide bond formation, equal volumes of IC (0.9 µM) and respective TC (0.3 µM) were mixed in a quench-flow apparatus for time ranging from milliseconds to seconds in TAKM7 at 37 °C. For GTP hydrolysis, the reaction was quenched with 50% formic acid. Intact [γ-32P]GTP and γ-32Pi were separated by TLC in 0.5 M KH2PO427. The TLC plates were analyzed using phosphorimaging in Typhoon FLA9500 (GE Healthcare). For peptide bond formation and translocation, reactions at specific time points were quenched with 0.5 M KOH and the peptides released by alkaline hydrolysis at 37 °C. Peptide samples were neutralized by adding one fifth volume of 100% glacial acetic acid and analyzed by reversed-phase HPLC (LiChroSpher 100 RP-8 HPLC column, Merck) using 0–65% acetonitrile gradient in 0.1% trifluoroacetic acid. The HPLC fractions were quantified by double label radioactivity counting49. The data were normalized to an interval from 0 to 1 and exponential fitting of the data was performed with GraphPad Prism.
smFRET experiments
Ribosomes labeled at protein L11 and Lys-tRNALys labeled with Cy5 at the 3-amino-3-carboxypropyl group at uridine 47 were prepared as described32,35. IC was prepared by incubating 70S(L11-Cy3) with a 3-fold excess of IFs, mRNA and f[3H]Met-tRNAfMet and GTP (1 mM) for 30 min at 37 °C and purified by centrifugation through sucrose cushion (1.1 M) in TAKM21 (50 mM Tris-HCl pH 7.5, 70 mM NH4Cl, 30 mM KCl, 21 mM MgCl2). The pellet was dissolved in TAKM7. TC was prepared by incubating 3-fold excess EF-Tu (or EF-Tu(H84A) mutant) with GTP (1 mM), phosphoenolpyruvate (3 mM), and pyruvate kinase (0.5%) for 15 min at 37 °C and subsequent addition of Lys-tRNALys(Cy5).
Biotin-coated glass objective slides and coverslips were prepared as described35. Reaction chambers were incubated with TAKM7 containing putrescine (8 mM), spermidine (1 mM), BSA (10 mg/ml) and neutravidin (1 μM; Thermo Scientific) for 5 min at room temperature. Neutravidin was washed by adding the same buffer containing putrescine (8 mM), spermidine (1 mM) and BSA (1 mg/ml). Purified IC(L11-Cy3) were diluted to 1 nM in TAKM7 containing putrescine (8 mM), spermidine (1 mM) and added to the reaction chambers for 2 min at room temperature. Imaging started after addition of TAKM7 containing putrescine (8 mM), spermidine (1 mM), protocatechuic acid (2.5 mM), 50 nM Pseudomonas protocatechuate-3,4-dioxygenase (50 nM), 6-hydroxy-2,5,7,8-tetramethylchromane-2-carboxylic acid (1 mM) methylviologen (1 mM; Sigma-Aldrich), GTP (1 mM) and TC(Lys-tRNALys(Cy5)) (5 nM).
TIRF imaging was performed at 22 °C on an IX 81 inverted microscope using a PLAPON 60 × 1.45 numerical aperture objective (Olympus). Cy3 was excited using a 561 nm solid-state laser operated at 25 mW and images were recorded with an electron multiplying CCD (charge-coupled device) camera (CCD-C9100-13, Hamamatsu) at a rate of 30.3 frames/s. Color channels were separated by projecting donor and acceptor emission on different parts of the CCD chip using an image splitter (dual view micro imager DV2, Photometrics), filter specifications HQ 605/40, HQ 680/30 (Chroma Technology).
Fluorescence time courses for Cy3 and Cy5 were extracted using custom-made MATLAB (MathWorks) software according to published protocols32,35. A semi-automated algorithm (MATLAB) was used to select single fluorophores showing anticorrelated fluorescence intensities and single-step photobleaching. Cy3 bleed-through into the Cy5 channel was corrected using an experimentally determined coefficient of 0.13. FRET efficiency was calculated as the ratio of the measured emission fluorescence intensities, FICy5/(FICy3 + FICy5). Trajectories were truncated to remove photobleaching and photoblinking events. The set of all FRET traces for a given complex was compiled in a histogram, which was fitted to a sum of Gaussian functions. MATLAB code using an unconstrained nonlinear minimization procedure (fminsearch, MATLAB, R2011b) yields mean values and standard deviation for the distribution of FRET states. Two-dimensional contour plots were generated from raw time-resolved FRET trajectories using a custom-made software. smFRET trajectories were fitted by Hidden Markov model using the vbFRET software package (http://vbfret.sourceforge.net/)50 to generate the idealized trajectories. FRET changes in idealized trajectories that were smaller than the s.d. of the Gaussian distribution of the FRET states were not considered transitions because they could not be not distinguished from the noise. Dwell times of different FRET states were calculated from idealized trajectories. The dwell-time distribution was fitted to an exponential function, y = y0 + Ae−t/τ to calculate the decay rate (k = 1/τ). GraphPad prism 8 software was used for the representation of smFRET data and fits of the data.
Cryo-EM grid preparation
IC was prepared and purified as described above except that non-radioactive fMet-tRNAfMet was used. TC was prepared from EF-Tu (1.5 µM), EF-Ts (0.02 mM), Lys-tRNALys (0.3 µM), GTP (1 mM), phosphoenolpyruvate (3 mM), pyruvate kinase (1%), DTT (1 mM) in buffer TAKM7. IC and TC were mixed in a 1:1 molar ratio, diluted to the final concentration of approximately A260~20 (absorbance at 260 nm 10 mm path) and A280~10 (absorbance at 280 nm 10 mm path), and incubated on ice for 60 s. The “unreacted” IC complexes contained all the above mentioned components except for Lys-tRNALys. Approximately 3 μL of the solution was applied onto freshly glow-discharged TEM grids (Quantifoil R2/1, Cu 200 mesh) and plunge-frozen into liquid ethane by a Vitrobot Mark IV (Thermo Fisher Scientific) using the following parameters: humidity 100%, temperature 4 °C, blot total 1,wait time 0, blot force 0, blot time 2 s, drain time 0 s.
Cryo-EM single-particle reconstruction
Cryo-EM datasets were collected at National Cryo-EM Centre SOLARIS (Kraków, Poland). The datasets of IC + TC complexes with AAA, m6AAA, Am6AA, and AAm6A codons in the A site, as well as AAA unreacted-IC and AAm6A unreacted-IC contained 8435, 9366, 7172, 7207, 10115, and 10276 movies, respectively (40 frames each). The movies were acquired using Titan Krios G3i microscope (Thermo Fisher Scientific) operated at 300 kV accelerating voltage, magnification of 105k, and corresponding pixel size of 0.86 Å/px. A K3 direct electron detector used for data collection was fitted with BioQuantum Imaging Filter (Gatan) using 20 eV slit. The K3 detector was operated in a counting mode. Imaged areas were exposed to 40 e−/Å2 total dose (corresponding to ~16 e−/px/s dose rate measured in vacuum). The frame stacks were obtained using under-focus optical conditions with a defocus range of −2.1 to −0.9 µm and 0.3 µm steps. The collected datasets were analyzed with cryoSPARC v3.3.051. Firstly, patch motion correction and patch CTF estimation were performed. Next, approximately 500 particles were picked manually. The acquired sets of particles were subjected to 2D classification and used in the generation of preliminary classes for template picking. The application of a template picker and 2D classification of the datasets resulted in correspondingly 689123, 1113602, 732439, 774481, 1413299, and 1282719 particles, respectively (Supplementary Figs. 3 and 9). These sets were used for Heterogenous Refinement and particles from the classes corresponding to well-defined 70 S ribosomes served as an input for focused 3D classifications. The 3D Classification was performed in cryoSPARC using soft masks corresponding to E-, P- and A-site tRNA generated in Relion 3.152. The unreacted 70S IC particles did not have any discernible density in the E and A sites, as expected for the IC. In the absence of Lys-tRNALys, the fraction of particles with ordered P-site fMet-tRNAfMet reached 78% (AAA) and 80% (AAm6A) in the two datasets, demonstrating the comparable quality of IC preparations. The final particle stacks were unbinned for local motion correction53 in cryoSPARC. During final 3D Homogenous Refinements, the particles were subjected to Defocus Refinement, Global CTF Refinement, and Ewald Sphere Correction to generate high-resolution maps. Local map resolution was calculated using cryoSPARC. Prior to model fitting, the combined half-maps were sharpened with DeepEMhancer54.
Molecular modeling
The initial atomic model was assembled by combining core elements of E. coli 70S ribosome structures solved by cryo-EM at 2.0 Å (PDB: 7K0055) and 2.54 Å resolution (PDB: 6XZB56). The initial model of tRNAfMet was sourced from a 3.2 Å ribosome cryo-EM structure (PDB: 6WDD24) and tRNALys was isolated from a 3.6 Å ribosome cryo-EM reconstruction (PDB: 5JTE57). Other structural elements, such as mRNA, were built manually into the map using Coot58. Following rigid body fitting using ChimeraX59, the atomic coordinates were flexibly fit with Namdinator60 and Isolde61. The models were real-space refined in Phenix62. The final coordinates were validated using MolProbity63 and the model statistics are presented in Supplementary Table 1. The cryo-EM maps and atomic models were displayed using ChimeraX version 1.2.5.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The micrographs, cryo-EM densities, and atomic models generated in this study have been deposited in the Electron Microscopy Public Image Archive (EMPIAR), the Electron Microscopy Data Bank (EMDB), and the Protein Data Bank (PDB) under the following accession codes: (1) AAA EMPIAR-11287; AAA IC EMD-16031 and PDB ID 8BGH; AAA P/P A/A EMD-16015 and PDB ID 8BF7. (2) m6AAA EMPIAR-11290; m6AAA IC EMD-16065 and PDB ID 8BHP; m6AAA P/P A/A EMD-16062 and PDB ID 8BHN. (3) Am6AA EMPIAR-11289; Am6AA IC EMD 16059 and PDB ID 8BHL; Am6AA P/P A/A EMD-16057 and PDB ID 8BHJ. (4) AAm6A EMPIAR-11288; AAm6A IC EMD-16029 and PDB ID 8BGE; AAm6A P/P A/A EMD-16047 and PDB ID 8BH4. (5) AAA unreacted-IC EMPIAR-11291, EMD-16081 and PDB ID 8BIL. (6) AAm6A unreacted-IC EMPIAR-11292, EMD-16082 and PDB ID 8BIM. Ensemble kinetics data are provided in the Source Data file. Processed smFRET data are provided in the Source Data file. Original images of smFRET experiments are available upon request due to their large size and the lack of a relevant public database. Source data are provided with this paper.
Code availability
The codes used to analyze data in this study are available from the corresponding authors upon request.
References
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Lesbirel, S. et al. The m(6)A-methylase complex recruits TREX and regulates mRNA export. Sci. Rep. 8, 13827 (2018).
Uzonyi, A. et al. Exclusion of m6A from splice-site proximal regions by the exon junction complex dictates m6A topologies and mRNA stability. Mol. Cell 83, 237–251 (2022).
Adhikari, S., Xiao, W., Zhao, Y. L. & Yang, Y. G. m(6)A: Signaling for mRNA splicing. RNA Biol. 13, 756–759 (2016).
Deng, X. et al. Widespread occurrence of N6-methyladenosine in bacterial mRNA. Nucleic Acids Res. 43, 6557–6567 (2015).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Wang, K., Peng, J. & Yi, C. The m(6)A consensus motif provides a paradigm of epitranscriptomic studies. Biochemistry 60, 3410–3412 (2021).
Wei, C. M. & Moss, B. Nucleotide sequences at the N6-methyladenosine sites of HeLa cell messenger ribonucleic acid. Biochemistry 16, 1672–1676 (1977).
Su, S. et al. Cryo-EM structures of human m(6)A writer complexes. Cell Res. 32, 982–994 (2022).
Patil, D. P., Pickering, B. F. & Jaffrey, S. R. Reading m(6)A in the transcriptome: m(6)A-binding proteins. Trends Cell Biol. 28, 113–127 (2018).
Zhen, D. et al. m(6)A Reader: epitranscriptome target prediction and functional characterization of N (6)-methyladenosine (m(6)A) readers. Front. Cell Dev. Biol. 8, 741 (2020).
Yen, Y. P. & Chen, J. A. The m(6)A epitranscriptome on neural development and degeneration. J. Biomed. Sci. 28, 40 (2021).
Liu, S. et al. Role of RNA N6-methyladenosine modification in male infertility and genital system tumors. Front. Cell Dev. Biol. 9, 676364 (2021).
Wang, T., Kong, S., Tao, M. & Ju, S. The potential role of RNA N6-methyladenosine in Cancer progression. Mol. Cancer 19, 88 (2020).
Mao, Y. et al. m(6)A in mRNA coding regions promotes translation via the RNA helicase-containing YTHDC2. Nat. Commun. 10, 5332 (2019).
Rodnina, M. V. Translation in prokaryotes. Cold Spring Harb. Perspect. Biol. 10, 1–21 (2018).
Rodnina, M. V. Decoding and recoding of mRNA sequences by the ribosome. Annu. Rev. Biophys. 52, 161–182 (2023).
Hoernes, T. P. et al. Nucleotide modifications within bacterial messenger RNAs regulate their translation and are able to rewire the genetic code. Nucleic Acids Res. 44, 852–862 (2016).
Choi, J. et al. N(6)-methyladenosine in mRNA disrupts tRNA selection and translation-elongation dynamics. Nat. Struct. Mol. Biol. 23, 110–115 (2016).
Ieong, K. W., Indrisiunaite, G., Prabhakar, A., Puglisi, J. D. & Ehrenberg, M. N 6-Methyladenosines in mRNAs reduce the accuracy of codon reading by transfer RNAs and peptide release factors. Nucleic Acids Res. 49, 2684–2699 (2021).
Ranjan, N. & Rodnina, M. V. Thio-Modification of tRNA at the Wobble Position as Regulator of the Kinetics of Decoding and Translocation on the Ribosome. J. Am. Chem. Soc. 139, 5857–5864 (2017).
Rodnina, M. V., Fischer, N., Maracci, C. & Stark, H. Ribosome dynamics during decoding. Philos. Trans. R. Soc. Lond. B Biol. Sci. 372, 1–10 (2017).
Loveland, A. B., Demo, G. & Korostelev, A. A. Cryo-EM of elongating ribosome with EF-Tu*GTP elucidates tRNA proofreading. Nature 584, 640–645 (2020).
Fischer, N. et al. The pathway to GTPase activation of elongation factor SelB on the ribosome. Nature 540, 80–85 (2016).
Kothe, U. & Rodnina, M. V. Delayed release of inorganic phosphate from elongation factor Tu following GTP hydrolysis on the ribosome. Biochemistry 45, 12767–12774 (2006).
Gromadski, K. B. & Rodnina, M. V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell 13, 191–200 (2004).
Rodnina, M. V., Pape, T., Fricke, R., Kuhn, L. & Wintermeyer, W. Initial binding of the elongation factor Tu.GTP.aminoacyl-tRNA complex preceding codon recognition on the ribosome. J. Biol. Chem. 271, 646–652 (1996).
Huang, B., Johansson, M. J. & Bystrom, A. S. An early step in wobble uridine tRNA modification requires the Elongator complex. RNA 11, 424–436 (2005).
Leidel, S. et al. Ubiquitin-related modifier Urm1 acts as a sulphur carrier in thiolation of eukaryotic transfer RNA. Nature 458, 228–232 (2009).
Rezgui, V. A. et al. tRNA tKUUU, tQUUG, and tEUUC wobble position modifications fine-tune protein translation by promoting ribosome A-site binding. Proc. Natl Acad. Sci. USA 110, 12289–12294 (2013).
Adio, S. et al. Fluctuations between multiple EF-G-induced chimeric tRNA states during translocation on the ribosome. Nat. Commun. 6, 7442 (2015).
Geggier, P. et al. Conformational sampling of aminoacyl-tRNA during selection on the bacterial ribosome. J. Mol. Biol. 399, 576–595 (2010).
Chen, C. et al. Single-molecule fluorescence measurements of ribosomal translocation dynamics. Mol. Cell 42, 367–377 (2011).
Poulis, P., Patel, A., Rodnina, M. V. & Adio, S. Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting. Nat. Commun. 13, 4231 (2022).
Daviter, T., Wieden, H. J. & Rodnina, M. V. Essential role of histidine 84 in elongation factor Tu for the chemical step of GTP hydrolysis on the ribosome. J. Mol. Biol. 332, 689–699 (2003).
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30 S ribosomal subunit. Science 292, 897–902 (2001).
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012).
Chandrasekaran, V. et al. Mechanism of ribosome stalling during translation of a poly(A) tail. Nat. Struct. Mol. Biol. 26, 1132–1140 (2019).
Tesina, P. et al. Molecular mechanism of translational stalling by inhibitory codon combinations and poly(A) tracts. EMBO J. 39, e103365 (2020).
Roost, C. et al. Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J. Am. Chem. Soc. 137, 2107–2115 (2015).
Rozov, A. et al. Tautomeric G*U pairs within the molecular ribosomal grip and fidelity of decoding in bacteria. Nucleic Acids Res. 46, 7425–7435 (2018).
Loveland, A. B., Demo, G., Grigorieff, N. & Korostelev, A. A. Ensemble cryo-EM elucidates the mechanism of translation fidelity. Nature 546, 113–117 (2017).
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002).
Ogle, J. M. & Ramakrishnan, V. Structural insights into translational fidelity. Annu. Rev. Biochem. 74, 129–177 (2005).
Gromadski, K. B., Daviter, T. & Rodnina, M. V. A uniform response to mismatches in codon-anticodon complexes ensures ribosomal fidelity. Mol. Cell 21, 369–377 (2006).
Rodnina, M. V., Fricke, R., Kuhn, L. & Wintermeyer, W. Codon-dependent conformational change of elongation factor Tu preceding GTP hydrolysis on the ribosome. EMBO J. 14, 2613–2619 (1995).
Wintermeyer, W. & Zachau, H. G. Fluorescent derivatives of yeast tRNAPhe. Eur. J. Biochem. 98, 465–475 (1979).
Wohlgemuth, I., Brenner, S., Beringer, M. & Rodnina, M. V. Modulation of the rate of peptidyl transfer on the ribosome by the nature of substrates. J. Biol. Chem. 283, 32229–32235 (2008).
Bronson, J. E., Fei, J., Hofman, J. M., Gonzalez, R. L. Jr. & Wiggins, C. H. Learning rates and states from biophysical time series: a Bayesian approach to model selection and single-molecule FRET data. Biophys. J. 97, 3196–3205 (2009).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Watson, Z. L. et al. Structure of the bacterial ribosome at 2A resolution. Elife 9, e60482 (2020).
Pichkur, E. B. et al. Insights into the improved macrolide inhibitory activity from the high-resolution cryo-EM structure of dirithromycin bound to the E. coli 70S ribosome. RNA 26, 715–723 (2020).
Arenz, S. et al. A combined cryo-EM and molecular dynamics approach reveals the mechanism of ErmBL-mediated translation arrest. Nat. Commun. 7, 12026 (2016).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Kidmose, R. T. et al. Namdinator - automatic molecular dynamics flexible fitting of structural models into cryo-EM and crystallography experimental maps. IUCrJ 6, 526–531 (2019).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Struct. Biol. 74, 519–530 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Petrychenko, V. et al. Structural mechanism of GTPase-powered ribosome-tRNA movement. Nat. Commun. 12, 5933 (2021).
Carbone, C. E. et al. Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP. Nat. Commun. 12, 7236 (2021).
Rundlet, E. J. et al. Structural basis of early translocation events on the ribosome. Nature 595, 741–745 (2021).
Acknowledgements
We thank M. Jaciuk, G. Ważny and P. Indyka for helpful discussions and O. Geintzer, V. Herold, T. Hübner, F. Hummel, S. Kappler, M. Klein, C. Kothe, A. Pfeiffer, T. Steiger and M. Zimmermann for expert technical assistance. The work was supported by the German Research Council (Deutsche Forschungsgemeinschaft) (Priority Programme SPP1784 to N.R. and M.V.R., Germany’s Excellence Strategy - EXC 2067/1- 390729940 and Leibniz Prize to M.V.R.) and the European Research Council (ERC) (grant agreement No 101001394- tRNAslation to S.G.). We thank the MCB structural biology core facility (supported by the TEAM TECH CORE FACILITY/2017-4/6 grant from Foundation for Polish Science, S.G.) for providing instruments and support. This publication was developed under the provision of the Polish Ministry of Education and Science project: “Support for research and development with the use of research infrastructure of the National Synchrotron Radiation Centre SOLARIS” under contract nr 1/SOL/2021/2. We acknowledge SOLARIS Centre for the access to the Krios microscope, where the measurements were performed. This research was supported in part by PLGrid Infrastructure (Academic Computer Centre Cyfronet AGH).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
S.J. performed biochemical and rapid kinetics ensemble experiments and data analysis. P.P. designed and performed smFRET experiments and analyzed the data. L.K. prepared most cryo-EM samples, analyzed the datasets and build atomic models with the support of I.K. and S.G. L.K., I.K. and S.G. analyzed the structures and prepared the structural figures. M.G. prepared initial cryo-EM samples, collected datasets and analyzed the data. M.R. collected all cryo-EM datasets, performed initial quality assessment and organized data storage. N.R., S.G. and M.V.R. conceptualized and supervised the research. S.J., L.K., P.P., S.G. and M.V.R. wrote the manuscript with contributions of all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Olivier Namy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jain, S., Koziej, L., Poulis, P. et al. Modulation of translational decoding by m6A modification of mRNA. Nat Commun 14, 4784 (2023). https://doi.org/10.1038/s41467-023-40422-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-40422-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.