Featured
-
-
Article
| Open Access Chromatin state changes during neural development revealed by in vivo cell-type specific profiling
Chromatin state changes during neural development revealed by in vivo cell-type specific profiling
While transitions between active and repressive chromatin states are essential for differentiation, little is known regarding their role during development of the brain in Drosophila. Here, the authors investigate the large scale chromatin remodelling taking place during fly neural development.
- Owen J. Marshall
- & Andrea H. Brand
-
Article
| Open Access Double mimicry evades tRNA synthetase editing by toxic vegetable-sourced non-proteinogenic amino acid
Double mimicry evades tRNA synthetase editing by toxic vegetable-sourced non-proteinogenic amino acid
Non-proteinogenic (np) amino acids in the food chain present challenges for the human translation machinery. Here the authors show that, while AlaRS and ProRS activate toxic np azetidine-2-carboxylic acid (Aze) present in sugar beets and lilies, only the AlaRS editing system rejects Aze.
- Youngzee Song
- , Huihao Zhou
- & Paul Schimmel
-
Article
| Open Access Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains
Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains
Proximity-ligation methods like Hi-C map DNA-DNA interactions and reveal its organization into topologically associating domains (TADs). Here the authors describe PSYCHIC, a computational approach for analysing Hi-C data that allows the identification of promoter-enhancer interactions.
- Gil Ron
- , Yuval Globerson
- & Tommy Kaplan
-
Article
| Open Access Circadian clock regulates hepatic polyploidy by modulating Mkp1-Erk1/2 signaling pathway
Circadian clock regulates hepatic polyploidy by modulating Mkp1-Erk1/2 signaling pathway
Circadian clock regulates hepatic gene expression and functions. Here Chao et al. show that alteration of circadian clock genes by Period deletion induces polyploidy in hepatocytes due to impaired regulation of Erk signaling by mitogen-activated protein kinase phosphatase 1.
- Hsu-Wen Chao
- , Masao Doi
- & Hitoshi Okamura
-
Article
| Open Access Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis
Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis
Despite being an important cancer drug target, the role of epigenetic reader Brd4 in cell differentiation and development remains unclear. Here, the authors provide evidence that Brd4 plays an important role in adipogenesis and myogenesis by binding to active enhancers to regulate gene expression.
- Ji-Eun Lee
- , Young-Kwon Park
- & Kai Ge
-
Article
| Open Access Dynamic intramolecular regulation of the histone chaperone nucleoplasmin controls histone binding and release
Dynamic intramolecular regulation of the histone chaperone nucleoplasmin controls histone binding and release
The histone chaperone nucleoplasmin (Npm) stores histones H2A/H2B in the egg and embryo. Here, the authors use NMR to show that Npm’s intrinsically disordered tail domain controls histone binding at an acidic stretch, which is autoregulated through direct competition with its basic C-terminus.
- Christopher Warren
- , Tsutomu Matsui
- & David Shechter
-
Article
| Open Access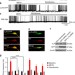 The sigma-1 receptor modulates methamphetamine dysregulation of dopamine neurotransmission
The sigma-1 receptor modulates methamphetamine dysregulation of dopamine neurotransmission
The dopamine transporter (DAT), a regulator of dopamine homeostasis in the brain, and sigma-1 receptor (σ1R), an endoplasmic reticulum membrane protein, are both implicated in drug addiction. In this work, the authors investigate how σ1R modulates DAT response to methamphetamine.
- Danielle O. Sambo
- , Min Lin
- & Habibeh Khoshbouei
-
Article
| Open Access RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
Juvenile myelomonocytic leukemia (JMML) is an aggressive disease with limited options for treatment. Here, the authors analyse the DNA methylome and mutational profile of JMML to define three subgroups with unique molecular and clinical characteristics.
- Daniel B. Lipka
- , Tania Witte
- & Christoph Plass
-
Article
| Open Access Catastrophic disassembly of actin filaments via Mical-mediated oxidation
Catastrophic disassembly of actin filaments via Mical-mediated oxidation
MICAL Redox enzymes post-translationally modify F-actin to promote its cellular destabilization. Here, the authors present a 3.9Å cryoEM structure of Mical-oxidized F-actin, showing its nucleotide-state dependent dynamic instability and susceptibility to cofilin-induced severing in the presence of inorganic phosphate.
- Elena E. Grintsevich
- , Peng Ge
- & Emil Reisler
-
Article
| Open Access RNA degradation by the plant RNA exosome involves both phosphorolytic and hydrolytic activities
RNA degradation by the plant RNA exosome involves both phosphorolytic and hydrolytic activities
The yeast and human RNA exosome is structurally related to prokaryotic phosphorylases but degrades RNA only via associated hydrolytic activities. Here the authors show that the RNA exosome of plants, and likely those of a few basal eukaryotes, combines phosphorolytic and hydrolytic activities to degrade RNA.
- Natalia Sikorska
- , Hélène Zuber
- & Dominique Gagliardi
-
Article
| Open Access Single-molecule imaging reveals multiple pathways for the recruitment of translesion polymerases after DNA damage
Single-molecule imaging reveals multiple pathways for the recruitment of translesion polymerases after DNA damage
Translesion synthesis (TLS) enables cells to tolerate damaged DNA encountered during replication. Here the authors use super-resolution photoactivation localization microscopy to reveal a lesion type dependent mechanism of recruitment of the TLS polymerase Pol IV following DNA damage.
- Elizabeth S. Thrall
- , James E. Kath
- & Joseph J. Loparo
-
Article
| Open Access DNA damage causes rapid accumulation of phosphoinositides for ATR signaling
DNA damage causes rapid accumulation of phosphoinositides for ATR signaling
Phosphoinositides are enriched in the nucleus and accumulate upon DNA damage but their role in responding to DNA damage is poorly defined. Here, the authors show that phosphoinositides rapidly accumulate at DNA damage sites and are required for ATR recruitment and subsequent Chk1 activation.
- Yu-Hsiu Wang
- , Anushya Hariharan
- & Michael P. Sheetz
-
Article
| Open Access In situ functional dissection of RNA cis-regulatory elements by multiplex CRISPR-Cas9 genome engineering
In situ functional dissection of RNA cis-regulatory elements by multiplex CRISPR-Cas9 genome engineering
RNA regulatory elements (RREs) are important post-transcriptional control features but studying them requires disrupting their activity without disturbing cellular homeostasis. Here the authors present GenERA, a CRISPR-Cas9 screening platform of in situ analysis of native RREs.
- Qianxin Wu
- , Quentin R. V. Ferry
- & Tudor A. Fulga
-
Article
| Open Access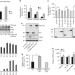 PI3Kδ activates E2F1 synthesis in response to mRNA translation stress
PI3Kδ activates E2F1 synthesis in response to mRNA translation stress
The oncogenic activity of EBNA1 protein is unknown; it contains a glycine and alanine repeat sequence (GAr) which regulates its own translation in cis. Here the authors show that GAr stimulates PI3Kδ-mediated induction of E2F1 translation, leading to c-Myc induction and stimulation of proliferation.
- Sivakumar Vadivel Gnanasundram
- , Slovénie Pyndiah
- & Robin Fåhraeus
-
Article
| Open Access SYK kinase mediates brown fat differentiation and activation
SYK kinase mediates brown fat differentiation and activation
Spleen protein tyrosine kinase (Syk) has so far been mainly studied in haematopoietic and immune cells. Here, the authors show that Syk also has a role in brown adipose tissue, where it regulates the formation of brown adipocytes and their thermogenic activation in response to β-adrenergic stimulation.
- Marko Knoll
- , Sally Winther
- & Harvey F. Lodish
-
Article
| Open Access Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Several families of natural compounds target core components of the pre-mRNA splicing machinery and display anti-tumor activity. Here the authors show that particular sequence features can be linked to drug response, and that drugs with very similar chemical structures display substantially different effects on splicing regulation.
- Luisa Vigevani
- , André Gohr
- & Juan Valcárcel
-
Article
| Open Access CTCF driven TERRA transcription facilitates completion of telomere DNA replication
CTCF driven TERRA transcription facilitates completion of telomere DNA replication
TERRA RNA is involved in maintaining stability during telomere repeat replication. Here the authors, by using CRISPR/Cas9, mutate CTCF-binding sites at start site of TERRA transcripts and find that subtelomeric CTCF facilitates telomeric DNA replication by promoting TERRA transcription.
- Kate Beishline
- , Olga Vladimirova
- & Paul M. Lieberman
-
Article
| Open Access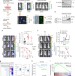 Breast cancer metastasis suppressor OTUD1 deubiquitinates SMAD7
Breast cancer metastasis suppressor OTUD1 deubiquitinates SMAD7
The activation of TGF-β signaling has been implicated in cancer metastasis. Here, the authors show that OTUD1 suppresses metastasis by antagonizing the TGF-β pathway via the deubiquitination of SMAD7, and its loss correlates with poor prognosis in breast cancer.
- Zhengkui Zhang
- , Yao Fan
- & Long Zhang
-
Article
| Open Access H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
SETDB1 is a histone methyltransferase that generates H3K9me3 marks in euchromatic regions. Here the authors show that the triple Tudor domain (3TD) of SETDB1 binds histone H3 tails containing K14 acetylation combined with K9 methylation, and that the K9me–K14ac modification defines a novel chromatin state enriched at SETDB1 binding sites.
- Renata Z. Jurkowska
- , Su Qin
- & Albert Jeltsch
-
Article
| Open Access Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors
Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors
Smad transcription factors are part of the TGF-β signal transduction pathways and are recruited to the genome by cell lineage-defining factors. Here, the authors identify specific Smad binding GC-rich motifs and provide structural information showing Smad3 and Smad4 bound to these motifs.
- Pau Martin-Malpartida
- , Marta Batet
- & Maria J. Macias
-
Article
| Open Access Proteomic analyses identify ARH3 as a serine mono-ADP-ribosylhydrolase
Proteomic analyses identify ARH3 as a serine mono-ADP-ribosylhydrolase
Protein ADP-ribosylation has emerged as a key post translational modification that regulates several stress responses. Here the authors characterize ARH3 as a major serine-specific mono–ADP-ribosylhydrolase and use a proteomics approach to identify the cellular targets of ARH3.
- Jeannette Abplanalp
- , Mario Leutert
- & Michael O. Hottiger
-
Article
| Open Access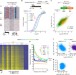 Global unleashing of transcription elongation waves in response to genotoxic stress restricts somatic mutation rate
Global unleashing of transcription elongation waves in response to genotoxic stress restricts somatic mutation rate
Precise orchestration of gene expression regulation upon DNA damage is essential for genome integrity. Here the authors identify a novel widespread stress-triggered defence mechanism that promotes rapid transcription-driven genomic surveillance thus limiting mutagenesis and shaping cancer genomes.
- Matthieu D. Lavigne
- , Dimitris Konstantopoulos
- & Maria Fousteri
-
Article
| Open Access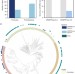 Inhibition of NHEJ repair by type II-A CRISPR-Cas systems in bacteria
Inhibition of NHEJ repair by type II-A CRISPR-Cas systems in bacteria
The double-strand breaks generated by CRISPR-Cas systems are the target of multiple DNA repair pathways. Here the authors find incompatibility between NHEJ and type II-A CRISPR-Cas systems due to Csn2 mediated inhibition of end-joining.
- Aude Bernheim
- , Alicia Calvo-Villamañán
- & David Bikard
-
Article
| Open Access Structure of outer membrane protein G in lipid bilayers
Structure of outer membrane protein G in lipid bilayers
Porins, like OmpG, are embedded in the outer membrane of bacteria and facilitate uptake and secretion of nutrients and ions. Here the authors present a protocol for solid state NMR structure determination of proteins larger than 25 kDa and use it to structurally characterize membrane embedded OmpG.
- Joren S. Retel
- , Andrew J. Nieuwkoop
- & Hartmut Oschkinat
-
Article
| Open Access A miR-327–FGF10–FGFR2-mediated autocrine signaling mechanism controls white fat browning
A miR-327–FGF10–FGFR2-mediated autocrine signaling mechanism controls white fat browning
White adipocytes can be stimulated to express thermogenic genes in a process known as beiging. Here, the authors show that miR-327 is downregulated during beiging, which releases FGF10 from inhibition and supports beige adipocyte formation via signaling through FGFR2.
- Carina Fischer
- , Takahiro Seki
- & Yihai Cao
-
Article
| Open Access Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis
Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis
Mutations in VHL have been linked to clear cell renal cancer, but the molecular mechanisms involved remain unclear. Here the authors generate a mouse model closely mimicking the human disease and show that VHL loss induces DNA replication stress that is rescued by the concomitant loss of PBRM1 permitting transformation.
- Judit Espana-Agusti
- , Anne Warren
- & Athena Matakidou
-
Article
| Open Access Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Most mammalian genes contain alternative polyadenylation sites. Here, the authors provide evidence that mRNA can be cleaved post-transcriptionally to generate mRNAs with shorter 3-´UTRs and stable autonomous uncapped 3´-UTR sequences.
- Yuval Malka
- , Avital Steiman-Shimony
- & Michael Berger
-
Article
| Open Access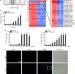 The LPS-inducible lncRNA Mirt2 is a negative regulator of inflammation
The LPS-inducible lncRNA Mirt2 is a negative regulator of inflammation
Excessive inflammation can be tissue destructive and contributes to auotinflammatory diseases and sepsis pathology. Here the authors show that the lncRNA Mirt2 is an endogenous negative feedback regulator of LPS-induced inflammation by limiting ubiquitination of TRAF6 and NF-κB activation.
- Meng Du
- , Lin Yuan
- & Kai Huang
-
Article
| Open Access Single-cell RNA-sequencing uncovers transcriptional states and fate decisions in haematopoiesis
Single-cell RNA-sequencing uncovers transcriptional states and fate decisions in haematopoiesis
Traditionally marker-based approaches are used to define haematopoietic cell type or state. Here, the authors use single-cell RNA-seq to establish a cellular hierarchy of lineage development in zebrafish haematopoiesis, and propose a refined model of developmental progression of haematopoietic cells.
- Emmanouil I. Athanasiadis
- , Jan G. Botthof
- & Ana Cvejic
-
Article
| Open Access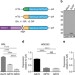 RAN translation at C9orf72-associated repeat expansions is selectively enhanced by the integrated stress response
RAN translation at C9orf72-associated repeat expansions is selectively enhanced by the integrated stress response
A nucleotide repeat expansion in C9orf72 is a common genetic cause of neurodegenerative disorders. Here, the authors provide insight into the molecular mechanism by which this repeat undergoes Repeat-Associated Non-AUG (RAN) translation, implicating the integrated stress response and eIF2α phosphorylation.
- Katelyn M. Green
- , M. Rebecca Glineburg
- & Peter K. Todd
-
Article
| Open Access The dynamic dimer structure of the chaperone Trigger Factor
The dynamic dimer structure of the chaperone Trigger Factor
The bacterial chaperone Trigger Factor (TF) is a dynamic protein and its dimer structure is unknown. Here the authors present a protocol combining NMR, computational and biophysical methods for the structural characterization of large dynamic protein complexes and show that TF forms a symmetric head-to-tail dimer.
- Leonor Morgado
- , Björn M. Burmann
- & Sebastian Hiller
-
Article
| Open Access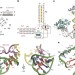 Crystal structure of an RNA-cleaving DNAzyme
Crystal structure of an RNA-cleaving DNAzyme
RNA-cleaving DNA enzymes are catalytic DNA that can cleave RNA in a sequence-specific manner. Here, the authors report three crystal structures of the 8–17 DNAzyme that include the pre-catalytic state of the RNA cleavage reaction, providing insight into the catalytic mechanism and may guide the rational design of DNAzymes.
- Hehua Liu
- , Xiang Yu
- & Jianhua Gan
-
Article
| Open Access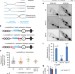 The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks
The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks
Terminally arrested replication forks are restarted through homologous recombination after processing single-stranded DNA gaps. Here the authors show that resection is regulated by the NHEJ factor Ku, helping to fine-tune recombination at forks.
- Ana Teixeira-Silva
- , Anissia Ait Saada
- & Sarah A. E. Lambert
-
Article
| Open Access Polη O-GlcNAcylation governs genome integrity during translesion DNA synthesis
Polη O-GlcNAcylation governs genome integrity during translesion DNA synthesis
Polη is a key player in translesion DNA synthesis. Here, the authors uncover that, in response to DNA damage, Polη undergoes O-GlcNAcylation at threonine 457 by O-GlcNAc transferase to facilitate the timely disassembly of Polη after DNA lesion bypass.
- Xiaolu Ma
- , Hongmei Liu
- & Caixia Guo
-
Article
| Open Access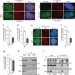 ATM and CDK2 control chromatin remodeler CSB to inhibit RIF1 in DSB repair pathway choice
ATM and CDK2 control chromatin remodeler CSB to inhibit RIF1 in DSB repair pathway choice
Cockayne syndrome group B protein (CSB) is a multifunctional chromatin remodeler involved in double-strand break repair. Here the authors investigate the molecular post-translational signals regulating CSB activity.
- Nicole L. Batenburg
- , John R. Walker
- & Xu-Dong Zhu
-
Article
| Open Access Alu-dependent RNA editing of GLI1 promotes malignant regeneration in multiple myeloma
Alu-dependent RNA editing of GLI1 promotes malignant regeneration in multiple myeloma
The treatment of multiple myeloma is challenging due to high relapse rates. Here the authors show that expression of ADAR1 correlates with poor patient outcomes, and that ADAR1-mediated editing of GLI1 is a mechanism relevant in the context of multiple myeloma progression and drug resistance.
- Elisa Lazzari
- , Phoebe K. Mondala
- & Catriona H. M. Jamieson
-
Article
| Open Access The transcript cleavage factor paralogue TFS4 is a potent RNA polymerase inhibitor
The transcript cleavage factor paralogue TFS4 is a potent RNA polymerase inhibitor
Transcript cleavage factors such as eukaryotic TFIIS assist the resumption of transcription following RNA pol II backtracking. Here the authors find that one of the Sulfolobus solfataricus TFIIS homolog—TFS4—has evolved into a potent RNA polymerase inhibitor potentially involved in antiviral defense.
- Thomas Fouqueau
- , Fabian Blombach
- & Finn Werner
-
Article
| Open Access BEX1 is an RNA-dependent mediator of cardiomyopathy
BEX1 is an RNA-dependent mediator of cardiomyopathy
Little is known about the changes in mRNA splicing, processing and stability that can alter gene expression during heart failure. Here, the authors show that BEX1 is induced during heart failure and is part of a ribonucleoprotein complex enhancing the expression and stability of proinflammatory genes.
- Federica Accornero
- , Tobias G. Schips
- & Jeffery D. Molkentin
-
Article
| Open Access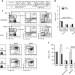 O-GlcNAcylation is required for B cell homeostasis and antibody responses
O-GlcNAcylation is required for B cell homeostasis and antibody responses
Post-translational modification has a variety of regulatory functions for important immune molecules. Here the authors use B-cell specific knockout mice to show how O-GlcNAcylation is required for functional B cell responses and humoral immunity.
- Jung-Lin Wu
- , Ming-Feng Chiang
- & Kuo-I Lin
-
Article
| Open Access Developmental YAPdeltaC determines adult pathology in a model of spinocerebellar ataxia type 1
Developmental YAPdeltaC determines adult pathology in a model of spinocerebellar ataxia type 1
Ataxin-1, linked to spinocerebellar ataxia type 1, is known to interact with the orphan nuclear receptor RORα. Here, Fujita and colleagues show that genetic supplementation of RORα-interacting protein YAPdeltaC during early development can rescue the adult pathologies of SCA1 mouse model.
- Kyota Fujita
- , Ying Mao
- & Hitoshi Okazawa
-
Article
| Open Access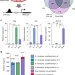 Synergistic gene expression during the acute phase response is characterized by transcription factor assisted loading
Synergistic gene expression during the acute phase response is characterized by transcription factor assisted loading
The cytokines IL-1β and IL-6 mediate the systemic acute phase response (APR). Here, the authors provide evidence that these cytokines lead to both synergistic and antagonistic gene expression during APR; synergistic induction occurs by assisted loading of STAT3 on chromatin by NF-κB.
- Ido Goldstein
- , Ville Paakinaho
- & Gordon L. Hager
-
Article
| Open Access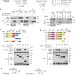 Parkin targets HIF-1α for ubiquitination and degradation to inhibit breast tumor progression
Parkin targets HIF-1α for ubiquitination and degradation to inhibit breast tumor progression
Parkin is an E3 ubiquitin ligase involved in Parkinson’s disease. Parkin has also been linked to cancer suppression but the mechanisms are unclear. Here the authors show that Parkin regulates HIF-1α through ubiquitin-dependent degradation, thus inhibiting metastasis of breast cancer cells.
- Juan Liu
- , Cen Zhang
- & Zhaohui Feng
-
Article
| Open Access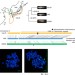 Centromere evolution and CpG methylation during vertebrate speciation
Centromere evolution and CpG methylation during vertebrate speciation
Centromeres and large-scale structural variants evolve and contribute to genome diversity during vertebrate speciation. Here Ichikawa et al perform de novo long-read genome assembly of three inbred medaka strains, and report long-range structure of centromeres and their methylation as well as correlation of structural variants with differential gene expression.
- Kazuki Ichikawa
- , Shingo Tomioka
- & Shinich Morishita
-
Article
| Open Access The chromatin remodeling factor ISW-1 integrates organismal responses against nuclear and mitochondrial stress
The chromatin remodeling factor ISW-1 integrates organismal responses against nuclear and mitochondrial stress
Changes in chromatin structure have been linked to organismal ageing. Here the authors show that altered histone expression and mitochondrial stress during C. elegans development result in chromatin changes and a cytosolic stress response that affects organismal longevity, and depends on HSF-1 and the chromatin remodeller, ISW-1.
- Olli Matilainen
- , Maroun S. Bou Sleiman
- & Johan Auwerx
-
Article
| Open Access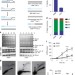 Break-induced replication promotes formation of lethal joint molecules dissolved by Srs2
Break-induced replication promotes formation of lethal joint molecules dissolved by Srs2
Break-induced replication (BIR) is a double-strand break repair pathway that can lead to genomic instability. Here the authors show that the absence of Srs2 helicase during BIR leads to uncontrolled binding of Rad51 to single-stranded DNA, which promotes the formation of toxic intermediates that need to be resolved by Mus81 or Yen1.
- Rajula Elango
- , Ziwei Sheng
- & Anna Malkova
-
Article
| Open Access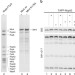 Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Excision of internal transcribed spacer 2 (ITS2) within eukaryotic pre-ribosomal RNA is essential for ribosome function. Here, the authors reconstitute the entire cycle of ITS2 processing in vitro using purified components, providing insights into the cleavage process and demonstrating that 26S pre-rRNA processing necessarily precedes 7S pre-rRNA processing.
- Lisa Fromm
- , Sebastian Falk
- & Ed Hurt
-
Article
| Open Access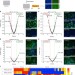 Cellular microRNA networks regulate host dependency of hepatitis C virus infection
Cellular microRNA networks regulate host dependency of hepatitis C virus infection
Using genome-wide miRNA mimic and hairpin inhibitor screens, Li et al. identify 31 miRNAs that either inhibit or promote hepatitis C virus (HCV) replication at different steps of the viral life cycle. Furthermore, human liver biopsies show that HCV down-regulates identified miRNAs with antiviral function.
- Qisheng Li
- , Brianna Lowey
- & T. Jake Liang
-
Article
| Open Access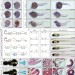 Evolutionary recruitment of flexible Esrp-dependent splicing programs into diverse embryonic morphogenetic processes
Evolutionary recruitment of flexible Esrp-dependent splicing programs into diverse embryonic morphogenetic processes
Epithelial-mesenchymal interplays are essential to many ontogenetic processes in vertebrates. Here Burguera et al. show diverse embryonic morphogenetic processes regulated by Epithelial Splicing Regulatory Protein (Esrp) in different deuterostome species.
- Demian Burguera
- , Yamile Marquez
- & Manuel Irimia
-
Article
| Open Access The STUbL RNF4 regulates protein group SUMOylation by targeting the SUMO conjugation machinery
The STUbL RNF4 regulates protein group SUMOylation by targeting the SUMO conjugation machinery
SUMO and ubiquitin are key signal transducers in several cellular processes including the DNA-damage response. Here the authors describe a method for selective enrichment of ubiquitin substrates for E3 ligases from complex cellular proteomes and identify the SUMO conjugation machinery as direct RNF4 substrates.
- Ramesh Kumar
- , Román González-Prieto
- & Alfred C. O. Vertegaal
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Inhibition of Poly(A)-binding protein with a synthetic RNA mimic reduces pain sensitization in mice
Inhibition of Poly(A)-binding protein with a synthetic RNA mimic reduces pain sensitization in mice