Article
|
Open Access
Featured
-
-
Article
| Open Access LARP7 family proteins have conserved function in telomerase assembly
LARP7 family proteins have conserved function in telomerase assembly
The telomerase holoenzyme is minimally composed of the reverse transcriptase and the RNA template. Here the authors identify Lar7 as a member of the full complex that helps to stabilise it and protect telomerase RNA from degradation.
- Laura C. Collopy
- , Tracy L. Ware
- & Kazunori Tomita
-
Article
| Open Access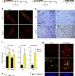 SOD3 improves the tumor response to chemotherapy by stabilizing endothelial HIF-2α
SOD3 improves the tumor response to chemotherapy by stabilizing endothelial HIF-2α
Tumour vasculature influences drug delivery. Here, the authors show that SOD3 re-expression enhances doxorubicin delivery and effects through normalization of tumour vasculature via the HIF-2a/VE-cadherin pathway.
- Emilia Mira
- , Lorena Carmona-Rodríguez
- & Santos Mañes
-
Article
| Open Access Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Cutaneous squamous cell carcinoma (cSCC) is a skin cancer that normally progresses from UV-induced actinic keratosis (AK). Here, the authors investigate the epigenomics of cSCC and highlight two distinct subclasses of AK and cSCC originating from distinct keratinocyte differentiation stages.
- Manuel Rodríguez-Paredes
- , Felix Bormann
- & Frank Lyko
-
Article
| Open Access Drosha drives the formation of DNA:RNA hybrids around DNA break sites to facilitate DNA repair
Drosha drives the formation of DNA:RNA hybrids around DNA break sites to facilitate DNA repair
The mechanism through which Drosha and Dicer affect DNA repair is not clear. Here the authors use a high-throughput approach to uncover the role of Drosha in promoting DNA:RNA hybrids at DNA damaged sites.
- Wei-Ting Lu
- , Ben R. Hawley
- & Martin Bushell
-
Article
| Open Access Senataxin resolves RNA:DNA hybrids forming at DNA double-strand breaks to prevent translocations
Senataxin resolves RNA:DNA hybrids forming at DNA double-strand breaks to prevent translocations
Recent studies suggest key roles of RNA in DNA double-strand breaks repair. Here the authors identify the helicase senataxin to be involved in DNA repair and resolve RNA:DNA hybrids forming at DNA double-strand breaks.
- Sarah Cohen
- , Nadine Puget
- & Gaëlle Legube
-
Article
| Open Access Cdk9 regulates a promoter-proximal checkpoint to modulate RNA polymerase II elongation rate in fission yeast
Cdk9 regulates a promoter-proximal checkpoint to modulate RNA polymerase II elongation rate in fission yeast
While post-translational modifications of the RNA polymerase II complex regulate gene expression, the exact mechanism remains unknown. Here, the authors use altered-specificity kinase mutations to examine the role of Cdk9, Mcs6/Cdk7, and Lsk1/Cdk12 on transcription at base-pair resolution in yeast.
- Gregory T. Booth
- , Pabitra K. Parua
- & John T. Lis
-
Article
| Open Access AhR and SHP regulate phosphatidylcholine and S-adenosylmethionine levels in the one-carbon cycle
AhR and SHP regulate phosphatidylcholine and S-adenosylmethionine levels in the one-carbon cycle
Methyl metabolites in the one-carbon cycle, such as phosphatidylcholines and S-adenosylmethionine, play a role in hepatic triglyceride regulation. Here Kim et al. show that AhR and SHP are both involved in the expression of several key enzymes of one-carbon metabolism, with the former regulating them early after feeding and the latter inhibiting AhR at later stages.
- Young-Chae Kim
- , Sunmi Seok
- & Jongsook Kim Kemper
-
Article
| Open Access A chiral selectivity relaxed paralog of DTD for proofreading tRNA mischarging in Animalia
A chiral selectivity relaxed paralog of DTD for proofreading tRNA mischarging in Animalia
The number of tRNA isodecoders has expanded significantly during evolution, which has resulted in ambiguity in tRNA selection by synthetases. Here the authors identify and characterize a dedicated proofreading factor that eliminates errors caused by ambiguity in tRNA selection by eukaryotic tRNA synthetases.
- Santosh Kumar Kuncha
- , Mohd Mazeed
- & Rajan Sankaranarayanan
-
Article
| Open Access Unraveling the determinants of microRNA mediated regulation using a massively parallel reporter assay
Unraveling the determinants of microRNA mediated regulation using a massively parallel reporter assay
MiRNAs are known regulators of gene expression. Here the authors perform a large-scale massively parallel reporter assay to investigate the effect of a large number of designed 3′ UTR sequences on reporter expression and asses how miRNA regulatory elements features affect miRNA mediated repression.
- Ilya Vainberg Slutskin
- , Shira Weingarten-Gabbay
- & Eran Segal
-
Article
| Open Access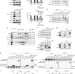 BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1
BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1
The polycomb group protein BMI1 is highly expressed in prostate cancer. Here, the authors demonstrate that BMI1 directly interacts with AR leading to increased AR signaling independently of PRC1 complex and that targeting BMI1 inhibits tumor growth of castration-resistant prostate cancer tumors.
- Sen Zhu
- , Dongyu Zhao
- & Qi Cao
-
Article
| Open Access Target identification of small molecules using large-scale CRISPR-Cas mutagenesis scanning of essential genes
Target identification of small molecules using large-scale CRISPR-Cas mutagenesis scanning of essential genes
Cancer therapy drugs are designed to target genetic vulnerabilities, but loss-of-function screens often fail to identify essential genes in drug mechanism studies. Here the authors demonstrate CRISPRres, which exploits in-frame variation generated by indel formation to discover gene-drug interactions.
- Jasper Edgar Neggers
- , Bert Kwanten
- & Dirk Daelemans
-
Article
| Open Access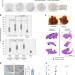 Hexokinase-2 depletion inhibits glycolysis and induces oxidative phosphorylation in hepatocellular carcinoma and sensitizes to metformin
Hexokinase-2 depletion inhibits glycolysis and induces oxidative phosphorylation in hepatocellular carcinoma and sensitizes to metformin
Hexokinase 2 (HK2) is selectively upregulated in hepatocellular carcinoma (HCC). Here the authors show that HK2 ablation decreases glycolysis and triggers oxidative phosphorylation (OXPHO) rendering HCC more susceptible to the OXPHO inhibitor metformin and to the FDA-approved drug sorafenib.
- Dannielle DeWaal
- , Veronique Nogueira
- & Nissim Hay
-
Article
| Open Access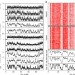 Single-cell replication profiling to measure stochastic variation in mammalian replication timing
Single-cell replication profiling to measure stochastic variation in mammalian replication timing
While DNA replication is temporally regulated during S-phase, variation in replication timing is not well understood. Here, the authors measure variation in replication timing using DNA copy number in single mouse ESCs and find stochastic variation to be independent of elements that regulate timing.
- Vishnu Dileep
- & David M. Gilbert
-
Article
| Open Access HTLV-1 Tax plugs and freezes UPF1 helicase leading to nonsense-mediated mRNA decay inhibition
HTLV-1 Tax plugs and freezes UPF1 helicase leading to nonsense-mediated mRNA decay inhibition
UPF1 is a central protein in nonsense-mediated mRNA decay (NMD), but contribution of its RNA processivity to NMD is unclear. Here, the authors show how the retroviral Tax protein interacts with and inhibits UPF1, and demonstrate that UPF1’s translocase activity contributes to NMD.
- Francesca Fiorini
- , Jean-Philippe Robin
- & Vincent Mocquet
-
Article
| Open Access Streamlined ex vivo and in vivo genome editing in mouse embryos using recombinant adeno-associated viruses
Streamlined ex vivo and in vivo genome editing in mouse embryos using recombinant adeno-associated viruses
CRISPR-Cas9 has been widely adopted for genetically manipulating rodents for scientific research. Here the authors transduce mouse embryos with CRISPR-Cas9 components using rAAVs in explant culture or in vivo to produce gene-edited animals.
- Yeonsoo Yoon
- , Dan Wang
- & Jaime A. Rivera-Pérez
-
Article
| Open Access Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
RNA-binding protein hnRNP A2/B1 is suggested to promote miRNA processing as a m6A 'reader'. Here, the authors determine crystal structures of RRM domains of hnRNP A2/B1 in complex with various RNA substrates and determine that hnRNP A2/B1 may function as an auxiliary factor in 'm6A switch' instead.
- Baixing Wu
- , Shichen Su
- & Jinbiao Ma
-
Article
| Open Access Kruppel-like factor 4-dependent Staufen1-mediated mRNA decay regulates cortical neurogenesis
Kruppel-like factor 4-dependent Staufen1-mediated mRNA decay regulates cortical neurogenesis
While being known as a transcription factor, Kruppel-like factor 4 (Klf4) may have other molecular functions. This study shows that Klf4 in neural progenitor cells regulate neurogenesis and self-renewal by interacting with RNA-binding protein Staufen1 and RNA helicase Ddx5/17 to control mRNA decay.
- Byoung-San Moon
- , Jinlun Bai
- & Wange Lu
-
Article
| Open Access Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
How the cardiac myocyte epigenome is rearranged during development, postnatal maturation and disease is not well understood. Here, the authors investigate the human cardiac myocyte epigenome during development and chronic heart failure and identify distinct epigenetic programs regulating these processes.
- Ralf Gilsbach
- , Martin Schwaderer
- & Lutz Hein
-
Article
| Open Access High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening
High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening
Cell-type transcriptome profiling greatly elucidate organismal development. Here, the authors report a spatiotemporally resolved comprehensive transcriptome analysis of tomato fruit ontogeny and suggest a new model of fruit maturation which initiates in internal tissues then radiates outwards.
- Yoshihito Shinozaki
- , Philippe Nicolas
- & Jocelyn K. C. Rose
-
Article
| Open Access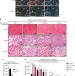 Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Satellite cells can differentiate both into myocytes and brown adipocytes. Here, the authors show that the histone demethylase Lsd1 prevents adipogenic differentiation of satellite cells by repressing expression of Glis1, and that its ablation changes satellite cell fate towards brown adipocytes and delays muscle regeneration in mice.
- Milica Tosic
- , Anita Allen
- & Roland Schüle
-
Article
| Open Access Impaired DNA damage response signaling by FUS-NLS mutations leads to neurodegeneration and FUS aggregate formation
Impaired DNA damage response signaling by FUS-NLS mutations leads to neurodegeneration and FUS aggregate formation
Abnormal cytoplasmic aggregates of FUS are a hallmark of some forms of amyotrophic lateral sclerosis (ALS). Here, using neurons derived from patients with FUS-ALS, the authors demonstrate that impairment of PARP-dependent DNA damage signaling is an event that occurs upstream of neurodegeneration and cytoplasmic aggregate formation in FUS-ALS.
- Marcel Naumann
- , Arun Pal
- & Andreas Hermann
-
Article
| Open Access Quantitative proteomics identifies redox switches for global translation modulation by mitochondrially produced reactive oxygen species
Quantitative proteomics identifies redox switches for global translation modulation by mitochondrially produced reactive oxygen species
The role of reactive oxygen species (ROS) in signalling and specific targets is not fully understood. Here the authors perform a global proteomic analysis to delineate the yeast redoxome and show that increased levels of intracellular ROS caused by dysfunctional mitochondria decrease global protein synthesis.
- Ulrike Topf
- , Ida Suppanz
- & Bettina Warscheid
-
Article
| Open Access Stress-dependent miR-980 regulation of Rbfox1/A2bp1 promotes ribonucleoprotein granule formation and cell survival
Stress-dependent miR-980 regulation of Rbfox1/A2bp1 promotes ribonucleoprotein granule formation and cell survival
Rbfox1, a pro-survival RNA-binding protein, is expressed in a complex manner and mediates diverse developmental processes. Here, the authors observe alternative splicing of Rbfox1 and stress-dependent regulation by miR-980 in Drosophila ovaries and Rbfox1 localisation in ribonucleoprotein granules in human cells.
- Mariya M. Kucherenko
- & Halyna R. Shcherbata
-
Article
| Open Access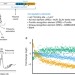 PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs
PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs
A large number of RNA binding proteins (RBPs) and miRNAs bind to the 3′ untranslated regions of mRNA, but methods to dissect their function and interactions are lacking. Here the authors introduce post-transcriptional regulatory element sequencing (PTRE-seq) to dissect sequence preferences, interactions and consequences of RBP and miRNA binding.
- Kyle A. Cottrell
- , Hemangi G. Chaudhari
- & Sergej Djuranovic
-
Article
| Open Access Binding of NUFIP2 to Roquin promotes recognition and regulation of ICOS mRNA
Binding of NUFIP2 to Roquin promotes recognition and regulation of ICOS mRNA
The RNA-binding proteins Roquin-1 and Roquin-2 are essential for immune cell function and postnatal survival in mice. Here, the authors identify NUFIP2 as a cofactor of Roquin; Roquin binds and stabilizes NUFIP2 in cells while NUFIP2 regulates Roquin mRNA target recognition.
- Nina Rehage
- , Elena Davydova
- & Vigo Heissmeyer
-
Article
| Open Access Enhancer-associated long non-coding RNA LEENE regulates endothelial nitric oxide synthase and endothelial function
Enhancer-associated long non-coding RNA LEENE regulates endothelial nitric oxide synthase and endothelial function
eNOS expression is dynamically regulated both transcriptionally and post-transcriptionally by various stimuli. Here the authors identify an enhancer-associated lncRNA (LEENE) that is co-regulated with, and enhances eNOS expression.
- Yifei Miao
- , Nassim E. Ajami
- & Zhen Chen
-
Article
| Open Access BLM helicase suppresses recombination at G-quadruplex motifs in transcribed genes
BLM helicase suppresses recombination at G-quadruplex motifs in transcribed genes
Bloom syndrome is characterized by high levels of sister chromatid exchanges (SCEs). Here, the authors use single-cell DNA template strand-sequencing to map SCEs in patient cells, and propose that the BLM helicase protects the genome against unwanted recombination at sites of G-quadruplex structures.
- Niek van Wietmarschen
- , Sarra Merzouk
- & Peter M. Lansdorp
-
Article
| Open Access A general and flexible method for signal extraction from single-cell RNA-seq data
A general and flexible method for signal extraction from single-cell RNA-seq data
Single-cell RNA sequencing (scRNA-seq) data provides information on transcriptomic heterogeneity within cell populations. Here, Risso et al develop ZINB-WaVE for low-dimensional representations of scRNA-seq data that account for zero inflation, over-dispersion, and the count nature of the data.
- Davide Risso
- , Fanny Perraudeau
- & Jean-Philippe Vert
-
Article
| Open Access Mechanically-sensitive miRNAs bias human mesenchymal stem cell fate via mTOR signalling
Mechanically-sensitive miRNAs bias human mesenchymal stem cell fate via mTOR signalling
Mesenchymal stem cell (MSC) fate can be mechanically regulated by substrate stiffness but this is difficult to control in a 3D hydrogel. Here the authors identify miRNAs that change expression in response to substrate stiffness and RhoA signalling and show that they can bias MSC fate in a 3D soft hydrogel.
- Jessica E. Frith
- , Gina D. Kusuma
- & Justin J. Cooper-White
-
Article
| Open Access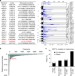 The protective role of DOT1L in UV-induced melanomagenesis
The protective role of DOT1L in UV-induced melanomagenesis
The interaction of DOT1L with MLL oncogenic fusion proteins has been implicated in leukemogenesis. Here, the authors show a contrasting role for DOT1L in protecting UVR-induced melanomagenesis by facilitating DNA repair through interaction with XPC.
- Bo Zhu
- , Shuyang Chen
- & Rutao Cui
-
Article
| Open Access The lncRNA GATA6-AS epigenetically regulates endothelial gene expression via interaction with LOXL2
The lncRNA GATA6-AS epigenetically regulates endothelial gene expression via interaction with LOXL2
LncRNAs influence endothelial cell function via a number of mechanisms. Here the authors show that the lncRNA GATA6-AS regulates endothelial gene expression through interaction with the nuclear deaminase LOXL2, with functional consequences on endothelial-mesenchymal transition and angiogenesis.
- Philipp Neumann
- , Nicolas Jaé
- & Stefanie Dimmeler
-
Article
| Open Access Dot1 regulates nucleosome dynamics by its inherent histone chaperone activity in yeast
Dot1 regulates nucleosome dynamics by its inherent histone chaperone activity in yeast
Dot1 is an evolutionarily conserved enzyme, responsible for histone H3K79 methylation in most eukaryotes. Here authors show that, in yeast, Dot1p has histone chaperone activity and regulates nucleosome dynamics via histone exchange in transcribed regions.
- Soyun Lee
- , Seunghee Oh
- & Daeyoup Lee
-
Article
| Open Access Single-molecule FRET reveals multiscale chromatin dynamics modulated by HP1α
Single-molecule FRET reveals multiscale chromatin dynamics modulated by HP1α
Chromatin fibers undergo continuous structural rearrangements but their dynamic architecture is poorly understood. Here, the authors use single-molecule FRET to determine the structural states and interconversion kinetics of chromatin fibers, monitoring their effector protein-dependent dynamic motions.
- Sinan Kilic
- , Suren Felekyan
- & Beat Fierz
-
Article
| Open Access High-resolution TADs reveal DNA sequences underlying genome organization in flies
High-resolution TADs reveal DNA sequences underlying genome organization in flies
Although topologically associating domains (TADs) have been extensively investigated, it is not clear to what extent DNA sequence contributes to their formation. Here the authors develop software to identify high-resolution TAD boundaries and reveal their relationship to underlying DNA motifs.
- Fidel Ramírez
- , Vivek Bhardwaj
- & Thomas Manke
-
Article
| Open Access USP48 restrains resection by site-specific cleavage of the BRCA1 ubiquitin mark from H2A
USP48 restrains resection by site-specific cleavage of the BRCA1 ubiquitin mark from H2A
BRCA1 ligase activity is tightly regulated to maintain genome stability and confer DNA double strand repair. Here the authors identify USP48 as a H2A deubiquitinating enzyme that acts as a BRCA1 E3 ligase antagonist and characterize its role during DNA repair.
- Michael Uckelmann
- , Ruth M. Densham
- & Joanna R. Morris
-
Article
| Open Access Sub-kb Hi-C in D. melanogaster reveals conserved characteristics of TADs between insect and mammalian cells
Sub-kb Hi-C in D. melanogaster reveals conserved characteristics of TADs between insect and mammalian cells
Topologically associating domain (TAD) boundaries in flies seem to be different from those in mammals. Here, the authors use Hi-C with sub-kb resolution to identify about 4000 TADs in flies, most demarcated by the insulator complexes BEAF-32/CP190 or BEAF-32/Chromator like CTCF/cohesin in mammals.
- Qi Wang
- , Qiu Sun
- & Zhifeng Shao
-
Article
| Open Access Structural insights into two distinct binding modules for Lys63-linked polyubiquitin chains in RNF168
Structural insights into two distinct binding modules for Lys63-linked polyubiquitin chains in RNF168
E3 ubiquitin ligase RNF168 is important for the repair of DNA double-strand breaks and recognizes ubiquitylated targets through two Ub-dependent DSB recruitment modules UDM1 and UDM2. Here the authors combine crystallography, cell biology and biochemical experiments to reveal how UDM1 and UDM2 interact with polyubiquitin chains.
- Tomio S. Takahashi
- , Yoshihiro Hirade
- & Shuya Fukai
-
Article
| Open Access CUG initiation and frameshifting enable production of dipeptide repeat proteins from ALS/FTD C9ORF72 transcripts
CUG initiation and frameshifting enable production of dipeptide repeat proteins from ALS/FTD C9ORF72 transcripts
Repeat-associated non-AUG (RAN) translation contributes to the pathogenic mechanism of several microsatellite expansion diseases. Here the authors delineate the different steps involved in recruiting the ribosome to initiate G4C2 RAN translation to produce poly-Glycine Alanine, poly-Glycine Proline, and poly-Glycine Arginine repeats.
- Ricardos Tabet
- , Laure Schaeffer
- & Clotilde Lagier-Tourenne
-
Article
| Open Access SIRT6-dependent cysteine monoubiquitination in the PRE-SET domain of Suv39h1 regulates the NF-κB pathway
SIRT6-dependent cysteine monoubiquitination in the PRE-SET domain of Suv39h1 regulates the NF-κB pathway
Sirtuins are involved in the regulation of responses to diverse types of cellular stress. Here the authors describe the SirT6-dependent cysteine monoubiquitination of the histone methyltransferase Suv39h1 as part of a regulatory circuit for the NF-κB pathway.
- Irene Santos-Barriopedro
- , Laia Bosch-Presegué
- & Alejandro Vaquero
-
Article
| Open Access Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases
Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases
Histone post-translational modifications are known key regulators of gene expression. Here, the authors characterize histone crotonylation at histone H3 lysine 18 in intestinal epithelia and find that it is a highly dynamic cell cycle regulated mark under the regulation of the HDAC deacetylases.
- Rachel Fellows
- , Jérémy Denizot
- & Patrick Varga-Weisz
-
Article
| Open Access TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice
TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice
Pachytene piRNAs are abundant piRNAs in mammalian adult testes but their biogenesis pathway is not fully understood. Here, the authors identify TDRD5 as a piRNA biogenesis factor in mice, showing that it binds piRNA precursors and promotes pachytene piRNA production from specific transcript regions.
- Deqiang Ding
- , Jiali Liu
- & Chen Chen
-
Article
| Open Access Integrated omics dissection of proteome dynamics during cardiac remodeling
Integrated omics dissection of proteome dynamics during cardiac remodeling
Transcriptome data provide only a partial picture of disease states. Here, via integration of transcript-, protein abundance and protein turnover data for a mouse model of cardiac hypertrophy, the authors uncover additional disease gene signatures, and show that turnover data sheds unique light on posttranslational regulation.
- Edward Lau
- , Quan Cao
- & Peipei Ping
-
Article
| Open Access Silencing of the lncRNA Zeb2-NAT facilitates reprogramming of aged fibroblasts and safeguards stem cell pluripotency
Silencing of the lncRNA Zeb2-NAT facilitates reprogramming of aged fibroblasts and safeguards stem cell pluripotency
The efficiency of somatic cell reprogramming is lowered by ageing. Here the authors show that the transcription factor Zeb2 and its long non-coding RNA Zeb2-NAT are expressed at high levels in older fibroblasts and their inhibition increases reprogramming efficiency.
- Bruno Bernardes de Jesus
- , Sérgio Pires Marinho
- & Maria Carmo-Fonseca
-
Article
| Open Access Structural analysis of mtEXO mitochondrial RNA degradosome reveals tight coupling of nuclease and helicase components
Structural analysis of mtEXO mitochondrial RNA degradosome reveals tight coupling of nuclease and helicase components
The mitochondrial RNA degradosome (mtEXO) plays an essential role in the regulation of mitochondrial gene expression and is composed of the 3′-to-5′ exoribonuclease Dss1 and the helicase Suv3. Here the authors present the RNA bound mtEXO crystal structure and give insights into its mechanism.
- Michal Razew
- , Zbigniew Warkocki
- & Marcin Nowotny
-
Article
| Open Access HoxC5 and miR-615-3p target newly evolved genomic regions to repress hTERT and inhibit tumorigenesis
HoxC5 and miR-615-3p target newly evolved genomic regions to repress hTERT and inhibit tumorigenesis
The expression of telomerase catalytic subunit hTERT is frequently upregulated in many cancers. Here, the authors show HoxC5 and miR-615-3p can negatively regulate hTERT to impede tumorigenesis by targeting the newly evolved cis-regulatory genomic elements of hTERT.
- TingDong Yan
- , Wen Fong Ooi
- & Shang Li
-
Article
| Open Access Targeting the CoREST complex with dual histone deacetylase and demethylase inhibitors
Targeting the CoREST complex with dual histone deacetylase and demethylase inhibitors
Alteration of the epigenetic landscape has been implicated in several disease processes, where targeting histone modifiers may have therapeutic applications. Here the authors report a bifunctional small molecule inhibitor that simultaneously targets the deacetylase (HDAC1) and demethylase (LSD1) activities of the CoREST complex.
- Jay H. Kalin
- , Muzhou Wu
- & Philip A. Cole
-
Article
| Open Access C9ORF72 GGGGCC repeat-associated non-AUG translation is upregulated by stress through eIF2α phosphorylation
C9ORF72 GGGGCC repeat-associated non-AUG translation is upregulated by stress through eIF2α phosphorylation
Hexanucleotide GGGGCC repeat expansion in C9ORF72 is the most frequent cause of both amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). Here the authors show that (GGGGCC) n translation can initiate without a 5′-cap, and this cap-independent translation is upregulated by stress mediated through eIF2α phosphorylation.
- Weiwei Cheng
- , Shaopeng Wang
- & Shuying Sun
-
Article
| Open Access RAD54 N-terminal domain is a DNA sensor that couples ATP hydrolysis with branch migration of Holliday junctions
RAD54 N-terminal domain is a DNA sensor that couples ATP hydrolysis with branch migration of Holliday junctions
RAD54 stimulates activity of the RAD51 recombinase and catalyzes branch migration of Holliday junctions during DNA repair and recombination. Here the authors show that the N-terminal domain of RAD54 mediates RAD54 oligomerization to promote branch migration, and is the target of phosphorylation that inhibits oligomerization and branch migration but not RAD51 stimulation.
- Nadish Goyal
- , Matthew J. Rossi
- & Alexander V. Mazin
-
Article
| Open Access LncRNA CAIF inhibits autophagy and attenuates myocardial infarction by blocking p53-mediated myocardin transcription
LncRNA CAIF inhibits autophagy and attenuates myocardial infarction by blocking p53-mediated myocardin transcription
Little is known about the role of long lncRNAs in autophagy. The authors identify lncCAIF, and show that it suppresses cardiac autophagy and attenuates myocardial infarction by targeting p53 -mediated transcription of myocardin.
- Cui-Yun Liu
- , Yu-Hui Zhang
- & Kun Wang
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Pof8 is a La-related protein and a constitutive component of telomerase in fission yeast
Pof8 is a La-related protein and a constitutive component of telomerase in fission yeast