Abstract
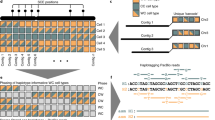
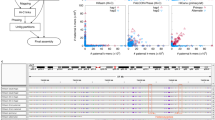
Haplotype information is essential to the complete description and interpretation of genomes1, genetic diversity2 and genetic ancestry3. Although individual human genome sequencing is increasingly routine4, nearly all such genomes are unresolved with respect to haplotype. Here we combine the throughput of massively parallel sequencing5 with the contiguity information provided by large-insert cloning6 to experimentally determine the haplotype-resolved genome of a South Asian individual. A single fosmid library was split into a modest number of pools, each providing ∼3% physical coverage of the diploid genome. Sequencing of each pool yielded reads overwhelmingly derived from only one homologous chromosome at any given location. These data were combined with whole-genome shotgun sequence to directly phase 94% of ascertained heterozygous single nucleotide polymorphisms (SNPs) into long haplotype blocks (N50 of 386 kilobases (kbp)). This method also facilitates the analysis of structural variation, for example, to anchor novel insertions7,8 to specific locations and haplotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
12 April 2011
In the version of this supplementary file originally posted online, Supplementary Figure 4a was not properly drawn. The error has been corrected in this file as of 12 April 2011.
References
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007).
International HapMap Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Green, R.E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Anonymous. Human genome: Genomes by the thousand. Nature 467, 1026–1027 (2010).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Kidd, J.M. et al. Mapping and sequencing of structural variation from eight human genomes. Nature 453, 56–64 (2008).
Li, R. et al. Building the sequence map of the human pan-genome. Nat. Biotechnol. 28, 57–63 (2010).
Kidd, J.M. et al. Characterization of missing human genome sequences and copy-number polymorphic insertions. Nat. Methods 7, 365–371 (2010).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
McKernan, K.J. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Schatz, M.C., Delcher, A.L. & Salzberg, S.L. Assembly of large genomes using second-generation sequencing. Genome Res. 20, 1165–1173 (2010).
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Roach, J.C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Reich, D., Thangaraj, K., Patterson, N., Price, A.L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high density in vitro transposition. Genome Biol. 11, R119 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Bansal, V. & Bafna, V. HapCUT: an efficient and accurate algorithm for the haplotype assembly problem. Bioinformatics 24, i153–i159 (2008).
Kim, J.H., Waterman, M.S. & Li, L.M. Diploid genome reconstruction of Ciona intestinalis and comparative analysis with Ciona savignyi. Genome Res. 17, 1101–1110 (2007).
Bansal, V., Halpern, A.L., Axelrod, N. & Bafna, V. An MCMC algorithm for haplotype assembly from whole-genome sequence data. Genome Res. 18, 1336–1346 (2008).
Alkan, C. et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 41, 1061–1067 (2009).
Hormozdiari, F. et al. Next-generation VariationHunter: combinatorial algorithms for transposon insertion discovery. Bioinformatics 26, i350–i357 (2010).
Zody, M.C. et al. Evolutionary toggling of the MAPT 17q21.31 inversion region. Nat. Genet. 40, 1076–1083 (2008).
Ng, S.B. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Ng, S.B. et al. Exome sequencing identifies the cause of a mendelian disorder. Nat. Genet. 42, 30–35 (2010).
Drysdale, C.M. et al. Complex promoter and coding region beta 2-adrenergic receptor haplotypes alter receptor expression and predict in vivo responsiveness. Proc. Natl. Acad. Sci. USA 97, 10483–10488 (2000).
Korbel, J.O. et al. Paired-end mapping reveals extensive structural variation in the human genome. Science 318, 420–426 (2007).
Ma, L. et al. Direct determination of molecular haplotypes by chromosome microdissection. Nat. Methods 7, 299–301 (2010).
Tycko, B. Allele-specific DNA methylation: beyond imprinting. Hum. Mol. Genet. 19, R210–R220 (2010).
Raymond, C.K. et al. Targeted, haplotype-resolved resequencing of long segments of the human genome. Genomics 86, 759–766 (2005).
Sudmant, P.H. et al. Diversity of human copy number variation and multicopy genes. Science 330, 641–646 (2010).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Acknowledgements
We thank C. Lee and M. Malig for technical assistance, J. Akey, T. O'Connor and P. Green for helpful discussions, D. Reich for ancestry information on NA20847, the U.W. Genome Sciences Genomics Resource Center (GS-GRC) for sequencing and the 1000 Genomes Project for early data release. This work was supported by National Institutes of Health grants AG039173 (J.B.H.) and HG002385 (E.E.E.), a National Science Foundation Graduate Research Fellowship (J.O.K.), a Natural Sciences and Engineering Research Council of Canada Fellowship (P.H.S.) and a fellowship from the Achievement Rewards for College Scientists Foundation (J.B.H.). E.E.E. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
The project was conceived and experiments planned by J.O.K., E.E.E. and J.S. J.O.K., A.P.M. and R.Q. carried out all experiments. J.O.K., A.A., J.B.H., R.P.P., P.H.S., S.B.N. and C.A. performed data analysis. J.O.K., A.P.M., A.A., J.B.H., R.P.P. and J.S. wrote the manuscript, and all authors reviewed it. All aspects of the study were supervised by J.S.
Corresponding authors
Ethics declarations
Competing interests
J.S. is a member of the science advisory boards of Tandem Technologies, Stratos Genomics, Good Start Genetics and Adaptive TCR. E.E.E. is on the scientific advisory board for Pacific Biosciences.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3,5, Supplementary Methods and Supplementary Figs. 1–7 (PDF 1756 kb)
Supplementary Table 4
Pan-genome and novel sequence anchoring. (XLS 1322 kb)
Rights and permissions
About this article
Cite this article
Kitzman, J., MacKenzie, A., Adey, A. et al. Haplotype-resolved genome sequencing of a Gujarati Indian individual. Nat Biotechnol 29, 59–63 (2011). https://doi.org/10.1038/nbt.1740
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1740
This article is cited by
-
Noninvasive prenatal diagnosis of monogenic disorders based on direct haplotype phasing through targeted linked-read sequencing
BMC Medical Genomics (2021)
-
Noninvasive prenatal testing of α-thalassemia and β-thalassemia through population-based parental haplotyping
Genome Medicine (2021)
-
Determination of complete chromosomal haplotypes by bulk DNA sequencing
Genome Biology (2021)
-
High-quality genome sequences of uncultured microbes by assembly of read clouds
Nature Biotechnology (2018)
-
Long range haplotyping of paired-homologous chromosomes by single-chromosome sequencing of a single cell
Scientific Reports (2018)