Abstract
Microbes associate in nature forming complex communities, but they are often studied in purified form. Here I show that neighbouring species enforce the re‐distribution of carbon and antimicrobial molecules, predictably changing drug efficacy with respect to standard laboratory assays. A simple mathematical model, validated experimentally using pairwise competition assays, suggests that differences in drug sensitivity between the competing species causes the re‐distribution of drug molecules without affecting carbon uptake. The re‐distribution of drug is even when species have similar drug sensitivity, reducing drug efficacy. But when their sensitivities differ the re‐distribution is uneven: The most sensitive species accumulates more drug molecules, increasing efficacy against it. Drug efficacy tests relying on samples with multiple species are considered unreliable and unpredictable, but study demonstrates that efficacy in these cases can be qualitatively predicted. It also suggests that living in communities can be beneficial even when all species compete for a single carbon source, as the relationship between cell density and drug required to inhibit their growth may be more complex than previously thought.
Similar content being viewed by others
Introduction
The notion of pure culture is fundamental in microbiology. The isolation and growth of individual species is justified by the seemingly unpredictability [1] of in vitro assays when cultures contain multiple species. For antimicrobial sensitivity tests, this means the same microbe can show different sensitivities to the same drugs, depending on whether the cultures contain one or multiple species [2,3,4]. But microbes live in communities in nature, even when they are mostly competing for resources [5]. The question is, therefore, whether co‐existing species can threaten drug efficacy in vivo with respect to standard sensitivity tests. Current data suggest they do [4, 6,7,8,9], notably reducing drug efficacy, and the result is that infections containing multiple species are more difficult to treat [10,11,12]—requiring alternatives to antibiotics altogether [10, 12]—and waters more difficult to treat [13]. This phenomenon has been associated with biofilm formation [14, 15], stochastic phenotypic variations of isogenic bacterial populations [14, 15], signalling molecules [14, 15] or enzymatic degradation of antimicrobials [14, 15]. But questions still remain about its predictability, and why, sometimes, drug efficacy seems to increase [6].
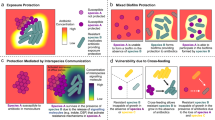
Below I present a model that provides a passive physical mechanism to explain and, perhaps more importantly, predict this phenomenon. The model relies on Fick’s first diffusion law and aims to understand how microbial growth changes the flow of carbon and antimicrobial molecules, and, thus, influence drug efficacy against all species exposed. Note that, when surrounded by neighbours, species attain lower densities within a community with respect to that in pure culture because carbon is shared between multiple species [16]. Now, the model predicts that carbon and antimicrobial molecules will distribute evenly among all species exposed if they have similar sensitivity to the antimicrobial. The result is relatively less drug molecules per cell of each species, and therefore lower drug efficacy. If one or more species are not sensitive to the antimicrobial, carbon molecules still distribute evenly as it is an active process [17, 18] but drug molecules are not: They flow back through diffusion from non‐sensitive species into the environment, re‐exposing those that are sensitive to the drug. Here, the model predicts relatively more drug per cell of sensitive species resulting thus in higher drug efficacy. These predictions were maintained across a range of parameter values for carbon uptake rate, carbon affinity, and biomass yield—fundamental components of the growth function in microbes [19].
To validate these predictions, I measured the efficacy of tetracycline against Escherichia coli Wyl in standard sensitivity assays using pure cultures, cultures with equal proportion of another microbe with similar drug‐sensitivity (Salmonella typhimurium), and cultures containing equal proportion of another microbe now tolerant to tetracycline (Escherichia coli GB(c)). Consistently with the theoretical predictions, inhibiting Wyl required more tetracycline—with respect to pure cultures—in the presence of equally sensitive neighbours and less tetracycline in the presence of drug‐tolerant neighbours.
Results
Consider n phenotypically distinct species competing for a limited resource, C, and exposed to a drug, A, supplied at concentration Ae (0) = A0, cast as the following model:
here \(\dot S_j\) and \(\dot A_j\) represent the density of individuals per unit volume from species j and their content of drug A over time, respectively, with initial conditions Sj(0) = Sj0, Aj(0) = 0, and C(0) = C0 > 0. The uptake of resource C by individuals of species j can be modelled as a saturating Monod function given that carbon transport is mediated by enzymes in nature [17, 18]. This function is proportional to the maximal uptake rate \(\overline {\mu _j}\),
where Kj is the half‐saturation parameter—the affinity of individuals from species j for the limited resource C is therefore given by 1/Kj. The growth rate of each species, at a given resource concentration is denoted by Gj(C): = Uj(C) · yj, where yj is the biomass yield per unit of resource in individuals from species j. Note that, for simplicity, I assume that yj does not vary between pure and mixed culture conditions. Biomass yield depends on resource availability [19] and competition for a common carbon source will likely change yj in all competing species, albeit these changes can be difficult to measure [20]. Their growth inhibition by drug A is described qualitatively by the Hill function [21].
Scaling the growth function Gj(S) similarly to bacteriostatic drugs [22]. This function is dimensionless and has two parameters. First, α is a dimensionless Hill coefficient which characterises the co‐operativity of the inhibition. And second, κj is the affinity of drug A for its target. It can be derived from the drug concentration required to halve the maximal growth rate, so that A50 = 1/κj → κj = 1/A50 [21]. For the sake of simplicity, I assumed that drug A diffuses from the environment into cells of species j, and vice versa, following Fick’s first diffusion law [23] with a diffusion coefficient \(\varphi _j\); and part of A being lost to chemical stability [24] at a rate d. This is not an unreasonable assumption given that antimicrobial molecules can indeed diffuse through cell membranes facilitated by membrane proteins [25]. Consequently, drug A is released in active form following the death of target and non‐target species.
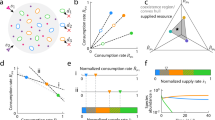
For my first computation I set the number of species j = 2, to facilitate later experimental validation, where I1(A) = I2(A) and G1(C) = G2(C). Thus, individuals from both species are sensitive to A and phenotypically identical. Given Eq. 3, the density of individuals from either species as pure cultures, after 24 h of incubation, declines with higher drug concentrations consistently with standard clinical protocols [26] (Fig. 1A). To allow experimental validation, I calculated the concentration of A inhibiting the growth of the pure cultures by 90% (IC90) as commonly used in clinic laboratories [27,28,29]. The drug sensitivity of each species depends on the values for the parameters K, \(\bar \mu\), and y of Eq. 2 (Fig. 1B–D, grey), with values that increase the density of individuals resulting in higher IC90 (details about the parameter values used can be found in Table 1). This is consistent with the inoculum effect [30] (Fig. S1), whereby sensitivity tests that use larger inocula also report higher minimum inhibitory concentrations, hence the standardisation of these clinical assays.
A Growth of species S1, with different parameter values (k1 \(\bar \mu\)1, and y1), after 24 h of growth in the presence of different antibiotic concentrations. I aggregated the resulting dose‐response profiles (blue) to create a density map from low predicted cell density (white) to high predicted cell density (black). B–D IC90, antibiotic concentration inhibiting 90% (IC90) the growth predicted without drug, resulting with different parameters values for the half‐saturation parameter k1 (B), maximal carbon up‐take \(\bar \mu\)1 (C), or biomass yield y1 (D) in eq. 1 when species S2 is drug‐sensitive. The IC90 for species S1 growing as pure cultures is shown in grey, and growing in mixed culture with S2 are shown in black. The parameter values for species S2 were fixed at a value noted by a black arrow on the y‐axis, followed by a dotted black line. E–G Change in IC90, as in Fig. (B–C), when the competing species S2 is not drug‐sensitive (resistant or tolerant). Parameter values used can be found in Table 1.
Figure S1 shows explicitly the theoretical relationship between IC90 and cell density of species S1 at the IC90 in pure culture, direct and non‐linear consistently with prior data [30,31,32]. For parameters \(\bar \mu\) and y, for example, the resulting change in IC90 with respect to the parameter values is nonmonotone, which means certain values for these parameters can maximise susceptibility for drug A. The neighbouring species shows similar, albeit not identical, changes in sensitivity (Fig. S2). This non‐linearity is caused by cultures not reaching the equilibrium, for all drug concentrations and parameter values tested, within the standard 24 h incubation times. If, in the model, cultures are allowed to grow for longer and reach equilibrium, the IC90–cell density profile can change (Fig. S1B). This phenomenon is exacerbated if both species grow in mixed culture conditions, where both become phenotypically more tolerant to drug A (Fig. 1B–D, black). If I were to target, say, individuals from species S1, doing so when the species is surrounded by S2 would require more drug. This is the case, for example, of pancreatic ductal adenocarcinoma treated with gemcitabine when bacteria grow within the tumour’s microenvironment [33]. More generally, genotypes analog to S1 should increase their drug tolerance when they are surrounded by similarly sensitive species. Despite the freedom in parameter values in the model, this prediction is robust to changes in biomass yield (yj) or maximal carbon uptake (\(\overline {\mu _j}\)) of species S2 (Fig. S3).
To test this hypothesis, I mixed equal proportions (cell/cell) of Escherichia coli Wyl and Salmonella typhimurium SL1344 in minimal media supplemented with different concentrations of tetracycline (see Methods). Note that the inoculum effect is not linear with respect to the size of inocula [31, 32] unless the difference between inocula is tenfold or more, which is not the case here. Tetracycline can diffuse passively into cells of both gram‐negative species [34], who also have similar sensitivity to this antibiotic: 0.232 ± 0.003 and 0.276 ± 0.016 μg/mL of tetracycline (mean IC90 ± 95% confidence, with n = 8 replicates, see Figs. S2 and S4). This approximates to I1(A) ≈ I2(A), as laid out by the theory above. The chromosome of E. coli Wyl carries yfp, gene encoding a yellow fluorescence protein (YFP), so I tracked its density in mixed culture conditions. Consistently with Eq. 1a–d, the bacterium was around 23% more tolerant to tetracycline when it grew in mixed culture with S. typhimurium (Mann–Whitney U‐test p = 1.554 × 10−4, ranksum = 36 with n = 8 replicates, Fig. 2A).
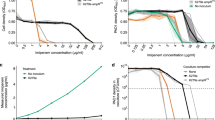
A, B Change in normalised density of Escherichia coli Wyl as a function of tetracycline concentration, when Wyl grows in mixed culture with tetracycline-sensitive Salmonella typhimurium (A) and tetracycline-resistant Escherichia coli GB(c) (B). The change in density of Wyl growing in mixed culture is shown in black, with grey showing the change in density in pure culture. The IC90 in each condition is shown as dots, red for mixed culture conditions and dark grey for pure culture, connected by a dotted line. Non-parametric, Mann–Whitney U-test between IC90s is shown in the inset. Raw data is shown as dots, whereas the boxes represent median (centre of the box), 25th, and 75th percentile of the dataset. The whiskers show the most extreme data points that are not outliers. C Theoretical change in S1-cell density with increasing antibiotic concentration in pure (grey) and mixed (black) culture conditions with neighbours that have different drug sensitivity. The plot represents the case where both species have different carbon uptake (\(\bar \mu\)j), and differences in IC90 are represented as shown in (A, B) for consistency. Insets in (A and B), and raw data for relative drug content can be found in Fig. S4.
Next, I explored in the model the case where individuals from both species have different sensitivities to drug A (I1(A) ≠ I2(A)). This scenario is akin to pathogens such as C. difficile growing alongside human cells [35] where the latter are unaffected by the drug (I2(A) ≈ 1). The model now predicts a subset of values for K, y, and \(\bar \mu\) that make S1 more sensitive to the drug in the presence of individuals from species S2 (Fig. 1E–G), whether S2 is not affected by drug A or is resistant to it for example through efflux pumps [36] (see Supplementary Text). To test this prediction, I mixed equal proportions (cell/cell) of two constructs of Escherichia coli with different sensitivities to tetracycline. One construct is Wyl, used above, who is sensitive to the antibiotic. The other construct is GB(c), harbouring a non‐transmissible plasmid carrying the gene tet(36) [37] and, therefore, resistant to the drug. Tetracycline binds to the bacterial ribosome, inhibiting protein synthesis [38], and tet(36) provides ribosomal protection against tetracycline [37] without degrading the antibiotic. The IC90 for this construct was 6.106 ± 0.272 μg/mL of tetracycline (mean IC90 ± 95% confidence with n = 8 replicates). Now, I1(A) « I2(A) satisfies the assumption above. The IC90 for E. coli Wyl was 0.232 ± 0.003 μg/mL of tetracycline as pure culture. Growing alongside drug‐resistant GB(c), however, it was 0.112 ± 0.003 μg/mL (Fig. 2B).
It is noteworthy to highlight the two processes at play. On the one hand, the growth of each species is limited due to competition for the same resource—also known as competitive suppression [16]. It could be conjectured that the observed variations in IC90 are caused by variations in growth, with respect to pure culture conditions, caused by competitive suppression. Indeed, E. coli Wyl grows less, with respect to pure culture conditions, in the presence of either competitor as Fig. S5 illustrates. Within this difference, Wyl grows more in the presence of tetracycline‐resistant E. coli GB(c), and yet, drug efficacy against Wyl was highest in the presence of this competitor. Now, the relative abundance of each species during the competition determines the flow and distribution of antimicrobial molecules given they diffuse passively following Fick’s law. Since glucose is actively transported into the cytoplasm, even against concentration gradients [17, 18], the model predicts virtually no effect on the carbon uptaken by each species as Fig. S6 illustrates. In other words, the carbon source C is depleted more rapidly when more species use it, limiting the growth of each competing species with respect to pure culture conditions.
On the other hand, the sensitivity to drug A determines the fraction of carbon that gets used by each species. When I1(A) ≈ I2(A), Eq. 1a–d suggest that individuals from both species accumulate similar amounts of drug A (Fig. S7A–C). This means that, analog to the carbon, drug A is depleted more rapidly, limiting the exposure of the target species S1. When combining both processes for carbon and drug molecules, the result is relatively fewer drug molecules per cell in both species with respect to that predicted in pure culture. However, when I1(A) ≠ I2(A), by virtue of different affinities of drug A for each species (κ1 ≠ κ2), the accumulation of drug molecules is uneven (Fig. S7D–I). Soon after the exposure to drug A, the species S2, with the least affinity for it, and the environment are in equilibrium. In other words, the number of drug molecules that diffuse into S2‐cells are the same that diffuse from these cells back to the environment. These molecules, in turns, are accumulated by the species with the most affinity for the drug. Here, integrating both processes results in more molecules per cell in the species with highest drug‐sensitivity (S1).
To verify this hypothesis, I estimated the content of tetracycline in E. coli Wyl by dividing the bacterium’s culture density, measured in relative fluorescence units to allow tracking in mixed culture conditions, by the concentration of tetracycline defining its IC90. The estimates closely resemble the theoretical predictions in Fig. 2C: E. coli Wyl contains ~20% less tetracycline growing next to Salmonella typhimurium and 65% more tetracycline growing alongside drug‐resistant GB(c) (Fig. 3).
A Difference in drug content per S1-cell at the IC90 between pure culture and mixed culture conditions. Positive and negative values denote more drug in S1-cells in pure and mixed culture conditions, respectively. Lack of difference is shown as a horizontal, dotted line. B Experimental estimation of the difference in relative cell content in the bacterium Escherichia coli Wyl. Raw data for Wyl is shown as red dots. Lack of difference is shown as a horizontal, black line and the 95% confidence of the lack of difference as a horizontal, dotted line.
Discussion
Why do microbes form communities? It was recently [5] suggested that bacteria rarely work together based on the biological interactions between them. But fundamental physics could draw a different picture, and help answer the question. If, as my study suggests, two species growing together can tolerate tetracycline better than they would otherwise do in isolation, it is not unreasonable to hypothesise that forming such communities can help diminish the effect of toxic molecules in nature. For example, to grow closer to an antibiotic‐producing microorganism and benefit from the carbon available or tolerate chemotherapies. Or, indeed, taking over a particular niche in case of drug‐tolerant microbes without producing specialised molecules such as bacteriocins [39]. All as a by‐product of a physical law.
It should be stressed that my theory is not specific to Escherichia coli and tetracycline, so I anticipate that other species and molecules obey the same principle described here. An example can be found in the interactions between tumours and bacteria, and the resulting tolerance to chemotherapies [8]; or between microbial pathogens [6]—albeit reliably predicting the outcome is still challenging [5]. Now, there are limitations in the model some of which are imposed by experimental conditions. For simplicity, the Fick’s term in Eq. 1b and c has no spatial derivative as it would be expected. However, during the experiment I shook the cultures to homogenise the distribution of tetracycline and nutrients. Here the aforementioned derivatives can be approximated as the difference in drug molecules between environment and cytoplasm. The model also assumes reliance on a single carbon source. In reality, however, I used two: Glucose as the main carbon source, and casamino acids. The latter can indeed be used as a carbon source when glucose is scarce [40]. The concentration I used (0.1% w/v) was low enough to enable the growth of the microorganisms without showing diauxic growth dynamics typical from cultures that are sustained on multiple carbon sources [41]. While speciation might indeed occur over time to avoid competition [42], is it really important to my conclusions?
The model that I present here is, like all models, wrong in some ways but also useful in others. The goal of my study is to demonstrate that growing alongside other microorganisms can predictably change the efficacy of a drug used against one—or against all—of them. It is, from the model standpoint, indifferent which carbon type is used to grow so long the relative frequency of the microbes surrounding the target is sufficiently high. Figure 1 shows the changes in IC90 are consistent regardless of the differences in growth parameters, akin to those existing between microorganisms using different carbon sources [43] or metabolic efficiencies [19]. Moreover, the model and experiment use two species mixed in the same proportion for the purpose of pinpointing the physical mechanism. In nature, communities are more complex and this phenomenon is likely to be more nuanced, if not even stronger, based on the ratio between the number of cells of the target species and its neighbours’.
My contribution with this model is to demonstrate that, contrary to current belief, the change in drug efficacy derived from the growth of other microorganisms is not arbitrary. There is, at least, one simple physical mechanism behind the changes in efficacy and, therefore, the changes can be qualitatively predicted—a necessary step if competition is to be harnessed [5] and used either in the clinic or treatment of pollutants in the soil.
Methods
Media and strains
The strains of Escherichia coli GB(c) and Wyl were a gift from Remy Chait and Roy Kishony, and non‐labelled Salmonella typhimurium SL1344 a gift from Markus Arnoldini and Martin Ackermann. Experiments were conducted in M9 minimal media supplemented with 0.4% glucose (Fisher Scientific #G/0500/53) and 0.1% casamino acids (Duchefa #C1301.0250), supplemented with tetracycline. M9 minimal media (50X) was made by mixing equal proportions (vol/vol) of two parts, part A and part B, and diluted accordingly to 1X. Part A (50X) contains 350 g/L of K2HPO4 (Sigma #P3786) and 100 g/L of KH2HPO4 (Sigma #P9791); whereas part B (50X) contains 29.4 g/L of trisodium citrate (Sigma #S1804), 50 g/L of (NH4)2SO4 (Sigma #A4418), and 10.45 g/L of MgSO4 (Fisher Scientific #M/1050/53). I made tetracycline stock solutions from powder stock (Duchefa #0150.0025) at 5 mg/mL in deionised water. Subsequent dilutions were made from this stock and kept at 4 °C.
Sensitivity assay
Using a 96‐pin replicator, I inoculated a 96‐well microtitre plate, containing 150 μL of media supplemented with 0–0.5 μg/mL of tetracycline (for E. coli Wyl and S. typhimurium) or 0–15 μg/mL (for E. coli GB(c)), with 1 μL from an overnight culture of each strain (~ 2·108 cells) to measure drug sensitivity in pure cultures. For sensitivity assays of Wyl in mixed culture conditions I inoculated the microtitre plate, containing 150 μg/mL of media supplemented with 0–0.5 μg/mL of tetracycline, with equal number of cells from two overnight cultures from Wyl and S. typhimurium using 8 technical replicates for each drug concentration. For the competition between Wyl and GB(c), however, I used 0–0.2 μg/mL to measure drug efficacy more reliably given the increased drug efficacy observed against Wyl. Figure S8 illustrates that mixing 1 μL of each species, therefore doubling the inoculum size with respect to pure culture conditions, or mixing 0.5 μL to maintain the same inoculum size has no impact on the sensitivity assays consistently with the nonlinear relationship reported between inoculum size and drug efficacy [31, 32].
I incubated the microtitre plate at 30 °C in a commercial spectrophotometer, shaking at 750 rpm to ensure uniform mixing, and measured the optical density of each well at 600 nm (OD600), yellow fluorescence for Wyl (YFP excitation at 505 nm, emission at 540 nm), and cyan fluorescence for GB(c) (CFP at 430 nm/480 nm) every 20 min for 24 h. I defined the minimum inhibitory concentration as the tetracycline concentration able to inhibit 90% of the growth observed in the absence of antibiotic after the 24 h incubation period.
Culture readings
Fluorescence protein genes were constitutively expressed with an approximately constant fluorescence to optical density ratio (Fig. S9). The number of colony forming units is positively correlated with optical density measured at 600 nm (OD600) (Fig. S10). Thus, I normalised fluorescence readings with respect to optical density readings, using the ratio optical density to fluorescence that I in pure culture conditions, to track the relative abundance of Wyl in mixed culture conditions. This metric is referred to as ‘Estimated optical density’ or ‘eOD’ in the main text. Time series dataset were blank corrected prior to calculating the minimum inhibitory concentration.
Data availability
References
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Disk Susceptibility Tests; Approved Standard—Eleventh Edition. CLSI document M02-A11 (ISBN 1-56238-781-2 [print]; ISBN 1-56238-782-0 [electronic]). Wayne, PA: Clinical and Laboratory Standards Institute; 2012.
Shahidi A, Ellner PD. Effect of mixed cultures on antibiotic susceptibility testing. Appl Microbiol. 1969;18:766–70.
Ellner PD, Johnson E. Unreliability of Direct Antibiotic Susceptibility Testing on Wound Exudates. Antimicrob Agents Chemother. 1976;9:355–6.
Galera‐Laporta L, Garcia‐Ojalvo J. Antithetic population response to antibiotics in a polybacterial community. Sci Adv. 2020;6:eaaz5108.
Palmer JD, Foster KR. Bacterial species rarely work together. Science. 2022;376:581–2.
Tampakakis E, Peleg AY, Mylonakis E. Interaction of Candida albicans with an Intestinal Pathogen, Salmonella enterica Serovar Typhimurium. Eukaryotic Cell. 2009;8:732–7.
Gudrun K, Ana Y, U FK, Charlotte W, T SS, Jennifer M, et al. Evolution of Resistance to a LastResort Antibiotic in Staphylococcus aureus via Bacterial Competition. Cell. 2015;158:1060–71.
Lehouritis P, Cummins J, Stanton M, Murphy CT, McCarthy FO, Reid G, et al. Local bacteria affect the efficacy of chemotherapeutic drugs. Sci Rep. 2015;5:1–12.
Lasa I, Solano C. Polymicrobial infections: do bacteria behave differently depending on their neighbours? Virulence. 2018;9:895–7.
Peters BM, Jabra‐Rizk MA, Graeme A, Costerton JW, Shirtliff ME. Polymicrobial interactions: impact on pathogenesis and human disease. Clin. Microbiol. Rev. 2012;25:193–213.
Simón‐Soro A, Mira A. Solving the etiology of dental caries. Trends Microbiol. 2015;23:76–82.
Buch PJ, Chai Y, Goluch ED. Treating polymicrobial infections in chronic diabetic wounds. Clin Microbiol Rev. 2019;32:e00091–18.
Vaccaro RF, Briggs MP, Carey CL, Ketchum BH. Viability of Escherichia coli in sea water. Am J Public Health Nations Health. 1950;40:1257–66.
Vega NM, Gore J. Collective antibiotic resistance: mechanisms and implications. Curr Opin Microbiol. 2014;21:28–34.
Bottery MJ, Pitchford JW, Friman VP. Ecology and evolution of antimicrobial resistance in bacterial communities. ISME J. 2021;15:939–48.
Grigg G. Competitive suppression and the detection of mutations in microbial populations. Aust J Biol Sci. 1958;11:69–84.
Buhr A, Flükiger K, Erni B. The glucose transporter of Escherichia coli. Overexpression, purification, and characterization of functional domains. J Biol Chem. 1994;269:23437–43.
Youk H, Van Oudenaarden A. Growth landscape formed by perception and import of glucose in yeast. Nature. 2009;462:875–9.
Reding‐Roman C, Hewlett M, Duxbury S, Gori F, Gudelj I, Beardmore R. The unconstrained evolution of fast and efficient antibiotic‐resistant bacterial genomes. Nat Ecol Evol. 2017;1:0050.
Ram Y, Dellus‐Gur E, Bibi M, Karkare K, Obolski U, Feldman MW, et al. Predicting microbial growth in a mixed culture from growth curve data. Proc Natl Acad Sci USA 2019;116:14698–707.
Klumpp S, Zhang Z, Hwa T. Growth rate‐dependent global effects on gene expression in bacteria. Cell. 2009;139:1366–75.
Pankey GA, Sabath LD. Clinical Relevance of Bacteriostatic versus Bactericidal Mechanisms of Action in the Treatment of Gram‐Positive Bacterial Infections. Clin Infect Dis. 2004;38:864–70.
Fick A. V. On liquid diffusion. Phil Mag. 1855;10:30–9.
Gómez‐Pacheco C, Sánchez‐Polo M, Rivera‐Utrilla J, López‐Peñalver J. Tetracycline degradation in aqueous phase by ultraviolet radiation. Chem Eng J. 2012;187:89–95.
Braun V, Bös C, Braun M, Killmann H. Outer membrane channels and active transporters for the uptake of antibiotics. J Infect Dis. 2001;183:S12–S16.
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Second Informational Supplement. CLSI document M100-S22 (ISBN 1-56238-785-5 [print]; ISBN 1-56238-786-3 [electronic]). Wayne, PA: Clinical and Laboratory Standards Institute; 2012.
Cedillo‐Rivera R, Munoz O. In‐vitro susceptibility of Giardia lamblia to albendazole, mebendazole and other chemotherapeutic agents. J Med Microbiol. 1992;37:221–4.
Cottrell ML, Hadzic T, Kashuba AD. Clinical pharmacokinetic, pharmacodynamic and drug‐interaction profile of the integrase inhibitor dolutegravir. Clin Pharmacokinet. 2013;52:981–94.
Baumgartner M, Bayer F, Pfrunder‐Cardozo KR, Buckling A, Hall AR. Resident microbial communities inhibit growth and antibiotic‐resistance evolution of Escherichia coli in human gut microbiome samples. PLoS Biol. 2020;18:e3000465.
Eng R, Smith S, Cherubin C. Inoculum effect of new beta‐lactam antibiotics on Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1984;26:42–7.
Alexander HK, MacLean RC. Stochastic bacterial population dynamics restrict the establishment of antibiotic resistance from single cells. Proc Natl Acad Sci USA 2020;117:19455–64.
Loffredo MR, Savini F, Bobone S, Casciaro B, Franzyk H, Mangoni ML, et al. Inoculum effect of antimicrobial peptides. Proc Natl Acad Sci USA. 2021;118:e2014364118.
Geller LT, Barzily‐Rokni M, Danino T, Jonas OH, Shental N, Nejman D, et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357:1156–60.
Grossman TH. Tetracycline antibiotics and resistance. Cold Spring Harb Perspect Med. 2016;6:a025387.
Burke KE, Lamont JT. Fecal Transplantation for Recurrent Clostridium difficile Infection in Older Adults: A Review. J Am Geriatr Soc. 2013;61:1394–8.
Blair JM, Bavro VN, Ricci V, Modi N, Cacciotto P, Kleinekathöfer U, et al. AcrB drug‐binding pocket substitution confers clinically relevant resistance and altered substrate specificity. Proc Natl Acad Sci USA 2015;112:3511–6.
Whittle G, Whitehead TR, Hamburger N, Shoemaker NB, Cotta MA, et al. Identification of a new ribosomal protection type of tetracycline resistance gene, tet(36), from swine manure pits. Appl Environ Microbiol. 2003;64:4151–8.
Epe B, Woolley P. The binding of 6‐demethylchlortetracycline to 70S, 50S and 30S ribosomal particles: a quantitative study by fluorescence anisotropy. EMBO J. 1984;3:121–6.
Dobson A, Cotter PD, Ross RP, Hill C. Bacteriocin Production: a Probiotic Trait? Appl Environ Microbiol. 2012;78:1–6.
Maser A, Peebo K, Vilu R, Nahku R. Amino acids are key substrates to Escherichia coli BW25113 for achieving high specific growth rate. Res Microbiol. 2020;171:185–93.
Monod J. The Growth of Bacterial Cultures. Annu Rev Microbiol. 1949;3:371–94.
Cooper VS. The origins of specialization: insights from bacteria held 25 years in captivity. PLoS Biol. 2014;12:e1001790.
Bren A, Park JO, Towbin BD, Dekel E, Rabinowitz JD, Alon U. Glucose becomes one of the worst carbon sources for E. coli on poor nitrogen sources due to suboptimal levels of cAMP. Sci Rep. 2016;6:1–10.
Acknowledgements
The author thanks Robert Beardmore, University of Exeter, for laboratory support in the early stages of this study.
Author information
Authors and Affiliations
Contributions
CR developed and implemented the mathematical model, designed and executed the experimental assays, the corresponding data analysis and visualisation, and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reding, C. Predicting the re‐distribution of antibiotic molecules caused by inter‐species interactions in microbial communities. ISME COMMUN. 2, 110 (2022). https://doi.org/10.1038/s43705-022-00186-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-022-00186-5