Abstract
Stem cell loss causes tissue deterioration associated with aging. The accumulation of genomic and oxidative stress-induced DNA damage is an intrinsic cue for stem cell loss1,2; however, whether there is an external microenvironmental cue that triggers stem cell loss remains unclear. Here we report that the involution of skin vasculature causes dermal stiffening that augments the differentiation and hemidesmosome fragility of interfollicular epidermal stem cells (IFESCs) in aged mouse skin. Aging-related IFESC dysregulation occurs in plantar and tail skin, and is correlated with prolonged calcium influx, which is contributed by the mechanoresponsive ion channel Piezo1 (ref. 3). Epidermal deletion of Piezo1 ameliorated IFESC dysregulation in aged skin, whereas Piezo1 activation augmented IFESC differentiation and hemidesmosome fragility in young mice. The dermis stiffened with age, which was accompanied by dermal vasculature atrophy. Conversely, induction of the dermal vasculature softened the dermis and ameliorated IFESC dysregulation in aged skin. Single-cell RNA sequencing of dermal fibroblasts identified an aging-associated anti-angiogenetic secretory molecule, pentraxin 3 (ref. 4), which caused dermal sclerotization and IFESC dysregulation in aged skin. Our findings show that the vasculature softens the microenvironment for stem cell maintenance and provide a potential mechanobiology-based therapeutic strategy against skin disorders in aging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The RNA-seq and scRNA-seq data were deposited in the Gene Expression Omnibus (GSE171035) and DNA Data Bank of Japan (DRA009978), respectively. Source data are provided with this study. All data needed to evaluate the conclusions in the study are present in the article and/or Supplementary Materials. All data are also available from the corresponding author upon reasonable request.
References
Behrens, A., van Deursen, J. M., Rudolph, K. L. & Schumacher, B. Impact of genomic damage and ageing on stem cell function. Nat. Cell Biol. 16, 201–207 (2014).
Vitale, I. et al. DNA damage in stem cells. Mol Cell. 66, 306–319 (2017).
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010).
Garlanda, C. et al. Non-redundant role of the long pentraxin PTX3 in anti-fungal innate immune response. Nature 420, 182–186 (2002).
Kurban, R. S. & Bhawan, J. Histologic changes in skin associated with aging. J. Dermatol. Surg. Oncol. 16, 908–914 (1990).
Ashcroft, G. S. et al. Estrogen accelerates cutaneous wound healing associated with an increase in TGF-β1 levels. Nat. Med. 11, 1209–1215 (1997).
Gosain, A. & DiPietro, L. A. Aging and wound healing. World J. Surg. 28, 321–326 (2004).
Liu, N. et al. Stem cell competition orchestrates skin homeostasis and ageing. Nature 568, 344–350 (2019).
Watanabe, M. et al. Type XVII collagen coordinates proliferation in the interfollicular epidermis. eLife 6, e26635 (2017).
Vázquez, F., Palacios, S., Alemañ, N. & Guerrero, F. Changes of the basement membrane and type IV collagen in human skin during aging. Maturitas 25, 209–215 (1996).
Bikle, D. D., Xie, Z. & Tu, C. L. Calcium regulation of keratinocyte differentiation. Expert Rev. Endocrinol. Metab. 7, 461–472 (2012).
Song, J. G. et al. Structural insights into Ca2+-calmodulin regulation of plectin 1a-integrin β4 interaction in hemidesmosomes. Structure 23, 558–570 (2015).
Wells, R. G. The role of matrix stiffness in regulating cell behavior. Hepatology 47, 1394–1400 (2008).
Wu, J., Lewis, A. H. & Grandl, J. Touch, tension, and transduction—the function and regulation of piezo ion channels. Trends Biochem. Sci. 42, 57–71 (2017).
Keyes, B. E. et al. Impaired epidermal to dendritic T cell signaling slows wound repair in aged skin. Cell 167, 1323–1338 (2016).
Quan, T. & Fisher, G. J. Role of age-associated alterations of the dermal extracellular matrix microenvironment in human skin aging: a mini-review. Gerontology. 61, 427–434 (2015).
Salzer, M. C. et al. Identity noise and adipogenic traits characterize dermal fibroblast aging. Cell 175, 1575–1590 (2018).
Ichijo, R. et al. Vasculature-driven stem cell population coordinates tissue scaling in dynamic organs. Sci. Adv. 7, eabd2575 (2021).
Gunin, A. G. et al. Age-related changes in angiogenesis in human dermis. Exp. Gerontol. 55, 143–151 (2014).
Detmar, M. et al. Increased microvascular density and enhanced leukocyte rolling and adhesion in the skin of VEGF transgenic mice. J. Invest. Dermatol. 111, 1–6 (1998).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Rusnati, M. et al. Selective recognition of fibroblast growth factor-2 by the long pentraxin PTX3 inhibits angiogenesis. Blood 104, 92–99 (2004).
Doni, A. et al. The long pentraxin PTX3 as a link between innate immunity, tissue remodeling, and cancer. Front. Immunol. 10, 712 (2019).
Valavanidis, A., Vlachogianni, T. & Fiotakis, C. 8-hydroxy-2'-deoxyguanosine (8-OHdG): a critical biomarker of oxidative stress and carcinogenesis. J. Environ. Sci. Health C 27, 120–139 (2009).
Jingpeng, G. et al. Architecture of the mammalian mechanosensitive Piezo1 channel. Nature 527, 64–69 (2015).
Syeda, R. et al. Piezo1 channels are inherently mechanosensitive. Cell Rep. 17, 1739–1746 (2016).
Saotome, K. et al. Structure of the mechanically activated ion channel Piezo1. Nature 554, 481–486 (2018).
Fornai, F. et al. The inflammatory protein pentraxin 3 in cardiovascular disease. Immun. Ageing 13, 25 (2016).
Vasioukhin, V., Degenstein, L., Wise, B. & Fuchs, E. The magical touch: genome targeting in epidermal stem cells induced by tamoxifen application to mouse skin. Proc. Natl Acad. Sci. USA 96, 8551–8556 (1999).
Chhalan, S. M. et al. Piezo1 links mechanical forces to red blood cell volume. eLife 4, e07370 (2015).
Hooper, A. T. et al. Engraftment and reconstitution of hematopoiesis is dependent on VEGFR2-mediated regeneration of sinusoidal endothelial cells. Cell Stem Cell 4, 263–274 (2009).
Ebina, T. et al. 3D clustering of GABAergic neurons enhances inhibitory actions on excitatory neurons in the mouse visual cortex. Cell Rep. 9, 1896–1907 (2014).
Okabe, K. et al. Neurons limit angiogenesis by titrating VEGF in retina. Cell 159, 584–596 (2014).
Butt, H. J. & Jaschke, M. Calculation of thermal noise in atomic force microscopy. Nanotechnology 6, 1 (1995).
Rodríguez-Franco, P. et al. Long-lived force patterns and deformation waves at repulsive epithelial boundaries. Nat. Mater. 16, 1029–1037 (2017).
Hino, N. et al. ERK-mediated mechanochemical waves direct collective cell polarization. Dev. Cell 53, 646–660 (2020).
Lichti, U., Anders, J. & Yuspa, S. H. Isolation and short-term culture of primary keratinocytes, hair follicle populations and dermal cells from newborn mice and keratinocytes from adult mice for in vitro analysis and for grafting to immunodeficient mice. Nat. Protoc. 3, 799–810 (2008).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Kechin, A., Boyarskikh, U., Kel, A. & Filipenko, M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput. Biol. 24, 1138–1143 (2017).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Butler, A. et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
We thank S. Kitano, H. Miyachi, Y. Iizuka, M. Yoshikawa, A. Sada and N. Hino for support and technical advice and A. Enomoto, T. Miyata and K. Kabashima for support on AFM and calcium imaging analyses. We thank staff at the Single-Cell Genome Information Analysis Core in ASHBi for RNA-seq analysis. We thank A. Mantovani and C. Garlanda at IRCCS Humanitas for the Ptx3 KO mice and Y. Kubota at Keio University for the VE-cadherin-creER mice. This work was supported by JSPS KAKENHI 16H06280 (R.I.), 19K17796 (R.I.), 21K16209 (R.I.), 17H05640 (F.T.), 20K21601 (F.T.), 16H06279 (PAGS) (F.T.), AMED under grant no. 20gm5810029 (F.T.), CREST under grant no. JPMJCR2023 (F.T., T.Y. and K.M.), the Takeda Science Foundation (F.T.), KAO Foundation for Arts and Sciences (R.I.), Fujiwara Memorial Foundation (R.I.), ASHBi, supported by the World Premier International Research Center Initiative (WPI) and MEXT Japan. We thank J. Ludovic Croxford and M. Crawford from Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
R.I. and F.T. conceived the project and designed the experiments. R.I. performed most experiments. K.M., A.N. and T.A. analyzed stiffness using AFM. M.K. and T.Y performed sequencing experiments and bioinformatics analyses. T.M. and T.H. contributed to live imaging experiments using two-photon microscopy. S.I. and H.H. contributed to hydrogel-related experiments for keratinocytes. R.I. and F.T. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Salvador Benitah, Richard Cubbon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Aging causes stem cell dysregulation.
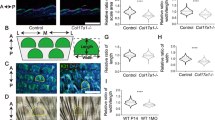
a, Representative immunofluorescence images and fluorescence intensities of COL17 in plantar skin from young (6 months) and aged (24–27 months) mice (n = 5). Data point; 30 cell average/mouse. b, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from young (6 months) and aged (24–27 months) mice (n = 5). Each data point is the average of 30 cells per mouse. c, Representative immunofluorescence images of survivin in IFESCs from control (6 months) and aged (24–27 months) mice. White lines: BM. White-dotted lines: cell boundary. Radial histogram quantification of division angles (n = 5, >10 cells/mouse). d, Gating strategy for flow cytometric analysis of IFESCs derived from the plantar epidermis. Error bars show the s.e.m. Two-tailed t-test (a and b). One-sided Kolmogorov–Smirnov test (c). n = number of mice.
Extended Data Fig. 2 Prolonged calcium flux through Piezo1 causes keratinocyte differentiation and hemidesmosome fragility.
a, Piezo1 fluorescence in situ hybridization (FISH) in plantar skin. Similar results were obtained from three biological independent experiments. White-dotted lines: BM. b, qPCR for Piezo1 in plantar epidermis from aged Piezo1f/f (24 months) and aged Piezo1 cKO (24 months) mice (n = 5). c, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from young Piezo1f/f (2 months), aged Piezo1f/f (24 months) and aged Piezo1 cKO mice (24 months) (n = 5). Data point; 30 cell average/mouse. d, Stiffness values for the dermis of young (6 months) and Piezo1 cKO aged (24 months) plantar skin (n = 5, >30 points/mouse). e, Stiffness values for the dermis of young (6 months), aged (24–27 months) plantar skin (n = 5, >30 points/mouse) and soft and stiff gels (n = 70 points on gels). f, Representative immunofluorescence images and quantification of K10+ cells in primary cultured keratinocytes (n = 3 biological replicates, >130 cells/experiment). g, Representative immunofluorescence images and fluorescence intensities of COL17 in primary cultured keratinocytes (n = 3 biological replicates). Data point; 20 cell average/mouse. h, Representative immunofluorescence images and quantification of K10+ cells in primary cultured Piezo1 cKO keratinocytes (n = 3 biological replicates, >120 cells/experiment). i, Representative immunofluorescence images and fluorescence intensities of COL17 in primary cultured Piezo1 cKO keratinocytes (n = 3 biological replicates). Data point; 20 cell average/experiment. Error bars show the s.e.m. Two-tailed t-test (b,d,f,g,h,i), Tukey’s multiple comparison tests (c). n = number of mice.
Extended Data Fig. 3 Piezo1 agonist Yoda1 induces IFESC dysregulation.
a, Representative immunofluorescence images and quantification of K10+ cells (arrowhead) in the basal layer of young (3 months) plantar skin injected with DMSO or Yoda1 (n = 5, >150 cells/mouse). Aged (24–27 months) plantar skin were injected with DMSO (n = 5, >170 cells/mouse). b, Representative immunofluorescence images and fluorescence intensities of COL17 in plantar skin from young (3 months) plantar skin injected with DMSO or Yoda1 (n = 5). Aged (24–27 months) plantar skin were injected with DMSO (n = 5). Data point; 30 cell average/mouse. c, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from young (3 months) plantar skin injected with DMSO or Yoda1 (n = 5). Aged (24–27 months) plantar skin were injected with DMSO (n = 5). Data point; 30 cell average/mouse. d, Representative immunofluorescence images of survivin in IFESCs from young (3 months) plantar skin injected with DMSO or Yoda1. Aged (24–27 months) plantar skin were injected with DMSO. White lines: BM. White-dotted lines: cell boundary. Radial histogram quantification of division angles (n = 5, >10 cells/mouse). e, Representative immunofluorescence images and quantification of K10+ cells in primary keratinocytes after treatment with DMSO or Yoda1 (n = 3 biological replicates, >120 cells/experiment). f, Representative immunofluorescence images and fluorescence intensities of COL17 in primary keratinocytes after treatment with DMSO or Yoda1 (n = 3 biological replicates). Data point; 20 cell average/experiment. Error bars show the s.e.m. Tukey’s multiple comparison tests (a,b,c), One-sided Kolmogorov–Smirnov test (d), and two-tailed t-test (e,f). n = number of mice.
Extended Data Fig. 4 Regional variability in aging-associated IFESC dysregulation.
a, Representative immunofluorescence images and fluorescence intensities of COL17 in back skin from young (6 months) and aged (24–27 months) mice (n = 5). Data point; 30 cell average/mouse. b, Representative immunofluorescence images and fluorescence intensities of COL17 in tail skin from young (6 months) and aged (24–27 months) mice (n = 5). Data point; 30 cell average/mouse c, Representative immunofluorescence images and quantification of K10+ cells (arrowhead) in the basal layer of tail skin from young (6 months) and aged (24–27 months) mice (n = 5, >150 cells/mouse). d, Representative immunofluorescence images and fluorescence intensities of COL17 in tail skin from young Piezo1f/f (6 months), aged Piezo1f/f (24–27 months) and aged Piezo1 cKO (24–27 months) mice (n = 5). Data point; 30 cell average/mouse e, Representative immunofluorescence images and quantification of K10 + cells (arrowhead) in the basal layer of tail skin from young Piezo1f/f (6 months), aged Piezo1f/f (24–27 months) and aged Piezo1 cKO (24–27 months) (n = 5, >150 cells/mouse). White-dotted lines: BM (c and e). Error bars show the s.e.m. Two-tailed t-test (a,b,c) and Tukey’s multiple comparison tests (d,e). n = number of mice.
Extended Data Fig. 5 Skin vasculatures regulate dermal ECM and epidermal hemidesmosome fragility.
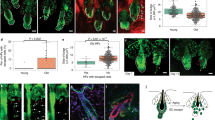
a, Representative immunofluorescence images and quantification of the length of elastin fibers in plantar skin from young (6 months) and aged (24–27 months) mice (n = 5, 10 points/mouse). White-dotted lines: BM. b, qPCR for VEGFA in plantar skin from aged WT (24–27 months) and aged K14VEGF (22 months) mice (n = 5). c, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from young WT (2 months), aged WT (22 months), and K14VEGF mice (22 months). Data point; 30 cell average/mouse. d, qPCR for VEGFR2 in the plantar dermis from VEGFR2f/f (6 months) and VEGFR2 cKO (6 months) mice (n = 5). e, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from VEGFR2f/f (6 months) and VEGFR2 cKO (6 months) mice (n = 5). Data point; 30 cell average/mouse. Error bars show the s.e.m. Two-tailed t-test (a,b,d,e), and Tukey’s multiple comparison tests (c). n = number of mice.
Extended Data Fig. 6 scRNA-seq analysis of mouse plantar dermis from young and aged mice.
a, Whole plantar dermal cell transcriptomes (n = 9,839) from two young (6 months) and two aged (24–27 months) mice plotted on a UMAP plot. The 15 clusters identified by unsupervised clustering are separated by color. Identity and marker genes of each cluster are indicated below the plots. b, Expression of cluster-specific marker genes projected onto UMAP plots.
Extended Data Fig. 7 Cell–cell communication analysis between endothelial cells and fibroblasts during aging.
a, Ligand–receptor interactions inferred by CellPhoneDB were shown. P values calculated in CellPhoneDB were indicated by circle size (scale on top). The means of the average expression level of interacting molecules are indicated by color. b, Violin plots showing the expression of ECM-related genes (Col1a1, Col1a2, Col3a1, Eln and Sparc), angiogenic genes (Sparc and Thy1) and anti-angiogenic genes (Thbs1 and Egr1) in fibroblast clusters (CL-4 and CL-12) from young and aged mice (two mice each). Wilcox test.
Extended Data Fig. 8 Feature plots for each fibroblast cluster.
Expression of cluster-specific marker genes projected onto UMAP plots. Samples were collected from two young and two aged mice.
Extended Data Fig. 9 Aging-induced IFESC dysregulation is ameliorated in Ptx3 KO plantar skin.
a, qPCR for Ptx3 in the plantar dermis from aged WT (24–27 months) and aged Ptx3 KO (24–27 months) mice (n = 5). b, Representative immunofluorescence images and quantification of the length of elastin fibers in plantar skin from young WT (2 months), aged WT (24–27 months), and aged Ptx3 KO (24–27 months) mice (n = 5, 10 points/mouse). White-dotted lines: BM. c, Representative immunofluorescence images and fluorescence intensities of plectin in plantar skin from young WT (2 months), aged WT (24–27 months), and aged Ptx3 KO (24–27 months) mice (n = 5). Data point; 30 cell average/mouse Error bars show the s.e.m. Two-tailed t-test (a), Tukey’s multiple comparison tests (b,c). n = number of mice.
Extended Data Fig. 10 Cellular senescence is partially rescued in Piezo1 cKO, K14VEGF, and Ptx3 KO mice.
a, Representative immunofluorescence images and fluorescence intensities of 8-OHdG in tail skin from young Piezo1f/f (6 months), aged Piezo1f/f (24–27 months) and aged Piezo1 cKO (24–27 months) mice (n = 5). Data point; 30 cell average/mouse. b, Representative immunofluorescence images and fluorescence intensities of 8-OHdG in plantar skin from young WT (2 months), aged WT (22 months) and aged K14VEGF mice (22 months) (n = 5). Data point; 30 cell average/mouse c, Representative immunofluorescence images and fluorescence intensities of 8-OHdG in tail skin from young WT (6 months), aged WT (24–27 months) and aged Ptx3 KO (24–27 months) mice (n = 5). Data point; 30 cell average/mouse. d, Schematic of the mechanism of age-associated IFESC dysregulation. White-dotted lines: BM (a,b,c). Error bars show the s.e.m. Tukey’s multiple comparison tests (a,b,c). n = number of mice.
Supplementary information
Supplementary Table 1
Top 50 DEGs in each dermal cluster.
Supplementary Table 2
Top 50 DEGs in each fibroblast cluster.
Source data
Source Data Fig. 1
Raw data.
Source Data Fig. 2
Raw data.
Source Data Fig. 3
Raw data.
Source Data Fig. 4
Raw data.
Source Data Extended Data Fig. 1
Raw data.
Source Data Extended Data Fig. 2
Raw data.
Source Data Extended Data Fig. 3
Raw data.
Source Data Extended Data Fig. 4
Raw data.
Source Data Extended Data Fig. 5
Raw data.
Source Data Extended Data Fig. 9
Raw data.
Source Data Extended Data Fig. 10
Raw data.
Rights and permissions
About this article
Cite this article
Ichijo, R., Maki, K., Kabata, M. et al. Vasculature atrophy causes a stiffened microenvironment that augments epidermal stem cell differentiation in aged skin. Nat Aging 2, 592–600 (2022). https://doi.org/10.1038/s43587-022-00244-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-022-00244-6
This article is cited by
-
A high-resolution view of the heterogeneous aging endothelium
Angiogenesis (2024)
-
The meaning of adaptation in aging: insights from cellular senescence, epigenetic clocks and stem cell alterations
Nature Aging (2023)
-
Niche stiffness regulates stem cell aging
Nature Aging (2022)