Abstract
Immediate restriction of iron initiated by the host is a critical process to protect against bacterial infections and has been described in the liver and spleen, but it remains unclear whether this response also entails a humoral mechanism that would enable systemic sequestering of iron upon infection. Here we show that upon bacterial invasion, host macrophages immediately release extracellular vesicles (EVs) that capture circulating iron-containing proteins. Mechanistically, in a sepsis model in female mice, Salmonella enterica subsp. enterica serovar Typhimurium induces endoplasmic reticulum stress in macrophages and activates inositol-requiring enzyme 1α signaling, triggering lysosomal dysfunction and thereby promoting the release of EVs, which bear multiple receptors required for iron uptake. By binding to circulating iron-containing proteins, these EVs prevent bacteria from iron acquisition, which inhibits their growth and ultimately protects against infection and related tissue damage. Our findings reveal a humoral mechanism that can promptly regulate systemic iron metabolism during bacterial infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All the data supporting the findings of this study are available within this article and the Supplementary Information files. Other information of this study is available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Thornton, F. J., Schäffer, M. R. & Barbul, A. Wound healing in sepsis and trauma. Shock 8, 391–401 (1997).
Cassat, J. E. & Skaar, E. P. Iron in infection and immunity. Cell Host Microbe 13, 509–519 (2013).
Wang, J. & Pantopoulos, K. Regulation of cellular iron metabolism. Biochem. J. 434, 365–381 (2011).
Soares, M. P. & Hamza, I. Macrophages and iron metabolism. Immunity 44, 492–504 (2016).
Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Invest. 126, 1208–1215 (2016).
Brahmer, A. et al. Platelets, endothelial cells and leukocytes contribute to the exercise-triggered release of extracellular vesicles into the circulation. J. Extracell. Vesicles 8, 1615820 (2019).
Bhatnagar, S. & Schorey, J. S. Exosomes released from infected macrophages contain Mycobacterium avium glycopeptidolipids and are proinflammatory. J. Biol. Chem. 282, 25779–25789 (2007).
Bhatnagar, S., Shinagawa, K., Castellino, F. J. & Schorey, J. S. Exosomes released from macrophages infected with intracellular pathogens stimulate a proinflammatory response in vitro and in vivo. Blood 110, 3234–3244 (2007).
Verweij, F. J. et al. Live tracking of inter-organ communication by endogenous exosomes in vivo. Dev. Cell 48, 573–589 (2019).
Villarroya-Beltri, C. et al. ISGylation controls exosome secretion by promoting lysosomal degradation of MVB proteins. Nat. Commun. 7, 13588 (2016).
Jeppesen, D. K. et al. Reassessment of exosome composition. Cell 177, 428–445 (2019).
Kastelowitz, N. & Yin, H. Exosomes and microvesicles: identification and targeting by particle size and lipid chemical probes. ChemBioChem 15, 923–928 (2014).
Keller, M. D. et al. Decoy exosomes provide protection against bacterial toxins. Nature 579, 260–264 (2020).
Drakesmith, H. & Prentice, A. M. Hepcidin and the iron-infection axis. Science 338, 768–772 (2012).
Shutinoski, B. et al. Lrrk2 alleles modulate inflammation during microbial infection of mice in a sex-dependent manner. Sci. Transl. Med. 11, eaas9292 (2019).
Wang, L. et al. Selective modulation of TLR4-activated inflammatory responses by altered iron homeostasis in mice. J. Clin. Invest. 119, 3322–3328 (2009).
Kim, D. K. et al. Inverse agonist of estrogen-related receptor γ controls Salmonella Typhimurium infection by modulating host iron homeostasis. Nat. Med. 20, 419–424 (2014).
Essandoh, K. et al. Blockade of exosome generation with GW4869 dampens the sepsis-induced inflammation and cardiac dysfunction. Biochim. Biophys. Acta 1852, 2362–2371 (2015).
Iguchi, Y. et al. Exosome secretion is a key pathway for clearance of pathological TDP-43. Brain 139, 3187–3201 (2016).
Walker, J. et al. Lipoxin a4 increases survival by decreasing systemic inflammation and bacterial load in sepsis. Shock 36, 410–416 (2011).
Gan, Z. et al. Regulation of macrophage iron homeostasis is associated with the localization of bacteria. Metallomics 11, 454–461 (2019).
Kapetanovic, R. et al. Pig bone marrow-derived macrophages resemble human macrophages in their response to bacterial lipopolysaccharide. J. Immunol. 188, 3382–3394 (2012).
Winn, N. C., Volk, K. M. & Hasty, A. H. Regulation of tissue iron homeostasis: the macrophage “ferrostat”. JCI Insight https://doi.org/10.1172/jci.insight.132964 (2020).
Mayhew, T. M. & Lucocq, J. M. Developments in cell biology for quantitative immunoelectron microscopy based on thin sections: a review. Histochem Cell Biol. 130, 299–313 (2008).
Ratledge, C. & Dover, L. G. Iron metabolism in pathogenic bacteria. Annu. Rev. Microbiol. 54, 881–941 (2000).
Nairz, M., Schroll, A., Sonnweber, T. & Weiss, G. The struggle for iron: a metal at the host–pathogen interface. Cell Microbiol. 12, 1691–1702 (2010).
Otto, B. R., Verweij-van Vught, A. M. & MacLaren, D. M. Transferrins and heme-compounds as iron sources for pathogenic bacteria. Crit. Rev. Microbiol. 18, 217–233 (1992).
Dichtl, S. et al. Dopamine is a siderophore-like iron chelator that promotes Salmonella enterica serovar Typhimurium virulence in mice. mBio https://doi.org/10.1128/mbio.02624-18 (2019).
Haley, K. P. & Skaar, E. P. A battle for iron: host sequestration and Staphylococcus aureus acquisition. Microbes Infect. 14, 217–227 (2012).
Lokken, K. L., Tsolis, R. M. & Bäumler, A. J. Hypoferremia of infection: a double-edged sword? Nat. Med. 20, 335–337 (2014).
Arezes, J. et al. Hepcidin-induced hypoferremia is a critical host defense mechanism against the siderophilic bacterium Vibrio vulnificus. Cell Host Microbe 17, 47–57 (2015).
Ganz, T. Iron in innate immunity: starve the invaders. Curr. Opin. Immunol. 21, 63–67 (2009).
Liao, Z. et al. Heat-killed Salmonella Typhimurium protects mice against carbon ion radiation. J. Int Med Res 48, 300060520924256 (2020).
Abels, E. R. & Breakefield, X. O. Introduction to extracellular vesicles: biogenesis, RNA cargo selection, content, release, and uptake. Cell Mol. Neurobiol. 36, 301–312 (2016).
Rashid, H. O., Yadav, R. K., Kim, H. R. & Chae, H. J. ER stress: autophagy induction, inhibition and selection. Autophagy 11, 1956–1977 (2015).
Moretti, J. et al. STING senses microbial viability to orchestrate stress-mediated autophagy of the endoplasmic reticulum. Cell 171, 809–823 (2017).
Zhang, Y. et al. Hepatotoxicity induced by isoniazid-lipopolysaccharide through endoplasmic reticulum stress, autophagy, and apoptosis pathways in zebrafish. Antimicrob. Agents Chemother. https://doi.org/10.1128/aac.01639-18 (2019).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 332, 1429–1433 (2011).
Nguyên, D. T. et al. Nck-dependent activation of extracellular signal-regulated kinase-1 and regulation of cell survival during endoplasmic reticulum stress. Mol. Biol. Cell 15, 4248–4260 (2004).
Martina, J. A., Chen, Y., Gucek, M. & Puertollano, R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8, 903–914 (2012).
Roczniak-Ferguson, A. et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci. Signal 5, ra42 (2012).
Hua, S. & Wu, S. Y. The use of lipid-based nanocarriers for targeted pain therapies. Front. Pharm. 4, 14 (2013).
Caza, M. & Kronstad, J. W. Shared and distinct mechanisms of iron acquisition by bacterial and fungal pathogens of humans. Front. Cell Infect. Microbiol. 3, 80 (2013).
Patruta, S. I. & Hörl, W. H. Iron and infection. Kidney Int. Suppl. 69, S125–S130 (1999).
Sarwar, H. S. et al. Redox biology of Leishmania and macrophage targeted nanoparticles for therapy. Nanomedicine 12, 1713–1725 (2017).
Wandersman, C. & Delepelaire, P. Bacterial iron sources: from siderophores to hemophores. Annu. Rev. Microbiol. 58, 611–647 (2004).
Hood, M. I. & Skaar, E. P. Nutritional immunity: transition metals at the pathogen-host interface. Nat. Rev. Microbiol. 10, 525–537 (2012).
Xiao, X., Yeoh, B. S. & Vijay-Kumar, M. Lipocalin 2: an emerging player in iron homeostasis and inflammation. Annu. Rev. Nutr. 37, 103–130 (2017).
Lin, J. et al. A Pseudomonas T6SS effector recruits PQS-containing outer membrane vesicles for iron acquisition. Nat. Commun. 8, 14888 (2017).
Carrière, J., Bretin, A., Darfeuille-Michaud, A., Barnich, N. & Nguyen, H. T. T. Exosomes released from cells infected with Crohn’s disease–associated adherent-invasive Escherichia coli activate host innate immune responses and enhance bacterial intracellular replication. Inflamm. Bowel Dis. 22, 516–528 (2016).
Schorey, J. S., Cheng, Y., Singh, P. P. & Smith, V. L. Exosomes and other extracellular vesicles in host–pathogen interactions. EMBO Rep. 16, 24–43 (2015).
Catalano, M. & O’Driscoll, L. Inhibiting extracellular vesicles formation and release: a review of EV inhibitors. J. Extracell. Vesicles 9, 1703244 (2020).
Zhao, Y. et al. Liver governs adipose remodelling via extracellular vesicles in response to lipid overload. Nat. Commun. 11, 1–17 (2020).
Huang, Y. et al. Zika virus propagation and release in human fetal astrocytes can be suppressed by neutral sphingomyelinase-2 inhibitor GW4869. Cell Discov. 4, 1–16 (2018).
Chiou, N. T., Kageyama, R. & Ansel, K. M. Selective export into extracellular vesicles and function of tRNA fragments during T cell activation. Cell Rep. 25, 3356–3370 (2018).
Jeong, M. H., Kim, H. R., Park, Y. J., Chung, K. H. & Kim, H. S. Reprogrammed lung epithelial cells by decrease of miR-451a in extracellular vesicles contribute to aggravation of pulmonary fibrosis. Cell Biol. Toxicol. 38, 725–740 (2022).
Kalluri, R. & LeBleu, V. S. The biology, function, and biomedical applications of exosomes. Science https://doi.org/10.1126/science.aau6977 (2020).
Wortzel, I., Dror, S., Kenific, C. M. & Lyden, D. Exosome-mediated metastasis: communication from a distance. Dev. Cell 49, 347–360 (2019).
Buzás, E. I., Tóth, E., Sódar, B. W. & Szabó-Taylor, K. Molecular interactions at the surface of extracellular vesicles. Semin. Immunopathol. 40, 453–464 (2018).
Luo, L. et al. Epidemiological and clinical differences between sexes and pathogens in a three-year surveillance of acute infectious gastroenteritis in Shanghai. Sci. Rep. 9, 1–9 (2019).
Stanaway, J. D. et al. The global burden of non-typhoidal Salmonella invasive disease: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Infect. Dis. 19, 1312–1324 (2019).
Chen, Y. Q. et al. Delivery of rapamycin by liposomes synergistically enhances the chemotherapy effect of 5-fluorouracil on colorectal cancer. Int J. Nanomed. 16, 269–281 (2021).
Nguyen, T., Du, J. & Li, Y. C. A protocol for macrophage depletion and reconstitution in a mouse model of sepsis. STAR Protoc. 2, 101004–101017 (2021).
Flo, T. H. et al. Lipocalin 2 mediates an innate immune response to bacterial infection by sequestrating iron. Nature 432, 917–921 (2004).
Liu, S. et al. Treatment of infarcted heart tissue via the capture and local delivery of circulating exosomes through antibody-conjugated magnetic nanoparticles. Nat. Biomed. Eng. 4, 1063–1075 (2020).
Dou, G. et al. Chimeric apoptotic bodies functionalized with natural membrane and modular delivery system for inflammation modulation. Sci. Adv. 6, eaba2987 (2020).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant no. 82170925 to Shiyu Liu, no. 81991504 to Y.J. and no. 31800817 to Siying Liu) and National Key Research and Development Program of China (grant no. 2016YFC1101400 to Y.J.).
Author information
Authors and Affiliations
Contributions
H.K. and G.D. designed, performed and interpreted the experiments and wrote the manuscript. L.C. and X.W. performed bacterial experiments and characterized properties of liposome. H.X., X.L., F.D. and Y.L. assisted with animal experiments. X.Y. and Siying Liu performed the histopathological studies and collected data. L.B., H.L. and B.L. contributed to data analysis and interpretation. Shiyu Liu developed the original concept. Shiyu Liu and Y.J. conceived the study and supervised the experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Frederik Verweij, Günter Weiss and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
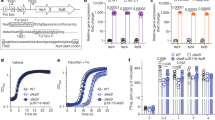
Extended Data Fig. 1 The characterization of S.Tm-infected mice and serum EVs biodistribution in vivo.
a-c, The C57BL/6 J mice were intraperitoneally administrated with S.Tm and blood and tissues samples were collected for further analysis. a, The viable counts of S.Tm in the blood at 24 hours after infection. n = 6 mice. b, Serum (left), hepatic (middle), and splenic (right) iron levels in mice at 24 hours after infection. n = 6 mice. c, The concentration of EVs in serum within 24 hours after infection. n = 3 mice per time point. d, Ex vivo fluorescence images of various organs in mice systemically injected with DiR-labeled serum EVs. n = 3 mice. e, f, Confocal microscopy images showing the uptake of PKH26-labeled EVs (red) by F4/80+ cells (green) (e) or Macro+ cells (green) (f) in liver or spleen. Scale bar, 10 μm. n = 3 biologically independent samples. For a-c, data are presented as the mean ± s.d. For a-c, statistical significance was assessed by unpaired two-sided Student’s t-test.
Extended Data Fig. 2 The effect of EVs release blockade or EVs supplementation on the host defense response to S.Tm infection.
a-c, To evaluate the effects of EVs release blockade on iron homeostasis, uninfected or S.Tm-infected mice were pretreated with GW4869 to block EVs release. a, Western blot analysis of FPN1 and FTH1 expressions in liver or spleen. Experiments were repeated three times and representative images are shown. b, The iron levels in liver or spleen at 12 hours after S.Tm infection. n = 6 mice. c, The viable count of S.Tm in liver and spleen at 12 hours after S.Tm infection. n = 6 mice. d,e, GW4869-pretreated S.Tm-infected mice were injected with EVs derived from uninfected mouse serum (Serum EVs group) or EVs derived from S.Tm-infected mouse serum [Serum(S.Tm)-EVs group]. d, The viable count of S.Tm in the liver and spleen. n = 5 mice. e, Representative fluorescence images of LPS (red) in the liver (upper) and spleen (bottom) and quantitative analysis of the percentage of LPS + area in liver or spleen cells. Scale bar, 50 μm. n = 3 biologically independent samples. For b-e, data are represented as the mean ± s.d. For b-e, statistical significance was assessed by one-way ANOVA with Tukey’s post-hoc test.
Extended Data Fig. 3 S.Tm infection affects iron homeostasis in BMDM.
a, Flow cytometric analysis of the expressions of macrophage surface marker F4/80 and CD11b on BMDM. b, c, BMDM were treated with S.Tm in the absence or presence of GW4869 for 24 hours to determine the iron homeostasis. b, Western blot analysis of expressions of iron-related proteins TfR, CD91, CD163, FTH1, and FPN1. Experiments were repeated three times and representative images are shown. c, The iron content in BMDM. n = 5 biologically independent samples. For c, data are represented as the mean ± s.d. For c, statistical significance was assessed by one-way ANOVA with Tukey’s post-hoc test.
Extended Data Fig. 4 EVs derived from S.Tm-infected BMDM decreased iron level and bacterial number in blood.
a,b, The infected BMDM-derived EVs [BMDM(S.Tm)-EVs group] and infected BMDM-derived EVs pretreated with excess holo-transferrin [BMDM(S.Tm)-EVs+holo-Tf group] were injected into GW4869-pretreated infected mice respectively. a, The iron level in serum. n = 4 mice. b, Viable count of S.Tm in blood. n = 4 mice. For a,b, data are represented as the mean ± s.d. For a,b, statistical significance was assessed by one-way ANOVA with Tukey’s post-hoc test.
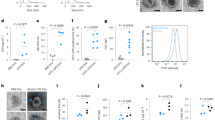
Extended Data Fig. 5 TfR-, CD163-, CD91-bearing EVs derived from S.a-infected mice or BMDM bind iron.
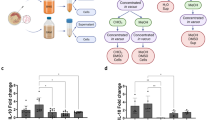
a-c, Wild-type mice was intraperitoneally injected with S.a and the serum were collected after 24 hours. BMDM were infected with S.a for 24 hours at a MOI of 25 and supernatant was collected. EVs were isolated from the serum or supernatant. a, Western blot analysis of TfR, CD163, CD91 and TSG101 expression in EVs derived from uninfected and infected mouse serum (left), and in EVs released from uninfected or infected BMDM (right). Experiments were repeated three times and representative images are shown. b, Immunoelectron microscopy detection for TfR, CD163, and CD91 antibodies in EVs derived from infected mouse serum (upper) or infected BMDM (bottom). Scale bar, 50 nm. Experiments were repeated twice and representative images are shown. c, Schematic diagram of the overall design of the experiments to test the ability of EVs to bind iron in serum. EVs derived from uninfected mouse serum (Serum EVs group) or S.a-infected mouse serum [Serum (S.a)-EVs group] were added to the serum respectively and then removed by ultracentrifugation. The serum supernatant was collected for further analysis. d, e, Total iron levels (d) and transferrin levels (e) in the serum supernatant after incubation with EVs derived from uninfected or infected mouse serum. n = 3 biologically independent samples. f, Growth of S.a in serum supernatant. n = 5 biologically independent samples. For d-f, data are presented as the mean ± s.d. For d and e, statistical significance was assessed by one-way ANOVA with Tukey’s post-hoc test. For f, statistical significance was assessed by one-way ANOVA with Tukey’s post hoc test and Kruskal-Wallis test.
Extended Data Fig. 6 HKS.Tm treatment enhances EVs release and expression of iron-related receptors.
a, b, Wild-type mice were treated with HKS.Tm for 24 hours, and serum EVs were isolated. a, The concentration of EVs in serum. n = 6 mice. b, Western blot analysis of iron-related receptors TfR, CD91, and CD163 expressions on EVs. Experiments were repeated three times and representative images are shown. For a, data are presented as the mean ± s.d. For a, statistical significance was assessed by unpaired two-tailed Student’s t-test.
Extended Data Fig. 7 EVs derived from macrophages protect against bacterial infection.
a, Representative dot plots of mouse spleen macrophages analyses at 48 hours after treatment with liposome-encapsulated clodronate or PBS. The graph showing the quantitative analysis of the percentage of F4/80+/MHCII+ macrophages in the spleen. n = 3 mice. b-e, To explore the role of macrophage EVs release in host EVs level regulation during infection, macrophage-depleted mice were injected intravenously with an equal volume of PBS, BMDM, or Rab27a shRNA-transfected BMDM for 36 hours, followed by S.Tm infection. b, The concentration of EVs in serum at 12 hours after S.Tm infection. n = 5 mice. c, The iron level in serum at 12 hours after S.Tm infection. n = 5 mice. d, Viable count of S.Tm in the blood at 12 hours after S.Tm infection. n = 5 mice. e, Survival rates of mice. n = 6 mice. f, To determine whether macrophage is the major source of serum EVs induced by HKS.Tm, macrophage-depleted mice were treated with an equal volume of PBS or BMDM for 36 hours, followed by HKS.Tm treatment. The graph showed the concentration of EVs in serum. n = 5 mice. For a-d and f, data are presented as the mean ± s.d. For a, statistical significance was assessed by unpaired two-tailed Student’s t-test. For b-d and f, statistical significance was assessed by one-way ANOVA with Tukey’s post-hoc test. For e, statistical significance was performed using the log-rank test.
Extended Data Fig. 8 S.a infection induces EVs release via endoplasmic reticulum stress.
a, Representative fluorescence images of ROS in BMDM infected with S.a at a MOI of 25, and quantitative analysis of the ROS levels in cells. Scale bar, 50 μm. n = 3 biologically independent samples. (b-g), BMDM were infected with S.a in the absence or presence of 4-PBA to clarify the association between ERS and EVs release. b, TEM images showed the ultrastructural morphology of the endoplasmic reticulum in BMDM. Scale bar, 1 μm. High-magnification images of the area marked by yellow boxes are arrayed at the lower panel. White arrow indicated endoplasmic reticulum. c, Western blot analysis of the expression of IRE1α, ATF4, ATF6 and GRP78 in BMDM. In b and c, experiments were repeated three times and representative images are shown. d, Representative fluorescence images of GRP78 in BMDM, and quantitative analysis of the fluorescence intensity Scale bar, 10 μm. n = 3 biologically independent samples. e, Representative fluorescence images of intracellular Ca2+ level in cells, and quantitative analysis of the Ca2+ levels. Scale bar, 4 μm. n = 3 biologically independent samples. f, Intracellular Ca2+ levels measured by fluorescence intensity. n = 3 biologically independent samples. g, The concentration of EVs in supernatant. n = 3 biologically independent samples. For a and d-g, data are presented as the means ± s.d. For a, statistical significance was analyzed by unpaired two-tailed Student’s t-test. For d-g, statistical significance was analyzed by one-way ANOVA with Tukey’s post-hoc test.
Extended Data Fig. 9 The biodistribution of 4-PBA-encapsulated liposome.
a, Ex vivo fluorescent images of various organs in mice injected with RhB-labeled 4-PBA-encapsulated liposome. n = 3 mice. b,c, The uptake of RhB -labeled liposome (red) by macrophages in liver (b) or in vitro cultured BMDM (c) stained with Hoechst (nucleus; blue) and F4/80 (cell membrane; green). Scale bar, 10μm. n = 3 biologically independent sample.
Extended Data Fig. 10 Rab31 regulates the enrichment of iron-related receptors onto EVs.
a, BMDM were treated with S.Tm in the absence or presence of 4-PBA for 24 hours, then EVs were isolated. The TfR, CD91, CD163, and TSG101 expressions on EVs were detected by Western blot. b-d, BMDM were pretreated with Rab31-siRNA and then infected with S.Tm. EVs from BMDM were isolated at 24 hours after S.Tm infection. The Rab31 expressions in BMDM (b) and TfR, CD91, CD163, and TSG101 expressions on EVs (c) were detected by Western blot. d, The concentration of EVs released by BMDM. n = 3 biologically independent samples. e, BMDM were treated with S.Tm in the absence or presence of 4-PBA for 24 hours. The IRE1α and Rab31 expressions in BMDM were detected by Western blot. f-i, BMDM were pretreated with IRE1α-siRNA (f,h) or toyocamycin (g,i) respectively and then were infected with S.Tm. The IRE1α and Rab31 expressions in BMDM (f, g), and TfR, CD91, CD163, and TSG101 expressions on EVs from BMDM (h,i) were analyzed by Western blot. For d, data are presented as the means ± s.d. For d, statistical significance was analyzed by one-way ANOVA with Tukey’s post-hoc test. All western blots were repeated three times and representative images are shown.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and gating strategy.
Supplementary Data 1
Source Data for Supplementary Fig. 1.
Supplementary Data 2
Information on antibodies used in the study.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical Source Data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kuang, H., Dou, G., Cheng, L. et al. Humoral regulation of iron metabolism by extracellular vesicles drives antibacterial response. Nat Metab 5, 111–128 (2023). https://doi.org/10.1038/s42255-022-00723-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-022-00723-5
This article is cited by
-
Bone marrow mesenchymal stem cell-derived exosomal microRNA-382 promotes osteogenesis in osteoblast via regulation of SLIT2
Journal of Orthopaedic Surgery and Research (2023)
-
Macrophage vesicles starve bacteria of iron
Nature Metabolism (2023)
-
Ubiquitination of ASCL1 mediates CD47 transcriptional activation of the AKT signaling pathway, and glycolysis promotes osteogenic differentiation of hBMSCs
In Vitro Cellular & Developmental Biology - Animal (2023)