Abstract
Iron is indispensable for almost all forms of life but toxic at elevated levels1,2,3,4. To survive within their hosts, bacterial pathogens have evolved iron uptake, storage and detoxification strategies to maintain iron homeostasis1,5,6. Recent studies showed that three Gram-negative environmental anaerobes produce iron-containing ferrosome granules7,8. However, it remains unclear whether ferrosomes are generated exclusively by Gram-negative bacteria. The Gram-positive bacterium Clostridioides difficile is the leading cause of nosocomial and antibiotic-associated infections in the USA9. Here we report that C. difficile undergoes an intracellular iron biomineralization process and stores iron in membrane-bound ferrosome organelles containing non-crystalline iron phosphate biominerals. We found that a membrane protein (FezA) and a P1B6-ATPase transporter (FezB), repressed by both iron and the ferric uptake regulator Fur, are required for ferrosome formation and play an important role in iron homeostasis during transition from iron deficiency to excess. Additionally, ferrosomes are often localized adjacent to cellular membranes as shown by cryo-electron tomography. Furthermore, using two mouse models of C. difficile infection, we demonstrated that the ferrosome system is activated in the inflamed gut to combat calprotectin-mediated iron sequestration and is important for bacterial colonization and survival during C. difficile infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All relevant data are within the manuscript and its Supplementary Information. Source data are provided with this paper.
References
Pi, H. & Helmann, J. D. Ferrous iron efflux systems in bacteria. Metallomics 9, 840–851 (2017).
Imlay, J. A. Pathways of oxidative damage. Annu. Rev. Microbiol. 57, 395–418 (2003).
Chandrangsu, P., Rensing, C. & Helmann, J. D. Metal homeostasis and resistance in bacteria. Nat. Rev. Microbiol. 15, 338–350 (2017).
Imlay, J. A. The mismetallation of enzymes during oxidative stress. J. Biol. Chem. 289, 28121–28128 (2014).
Sheldon, J. R. & Heinrichs, D. E. Recent developments in understanding the iron acquisition strategies of Gram positive pathogens. FEMS Microbiol. Rev. 39, 592–630 (2015).
Sheldon, J. R., Laakso, H. A. & Heinrichs, D. E. Iron acquisition strategies of bacterial pathogens. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.VMBF-0010-2015 (2016).
Lessa, F. C. et al. Burden of Clostridium difficile infection in the United States. N. Engl. J. Med. 372, 825–834 (2015).
Kelly, C. P. & LaMont, J. T. Clostridium difficile—more difficult than ever. N. Engl. J. Med. 359, 1932–1940 (2008).
Janka, J. & O’Grady, N. P. Clostridium difficile infection: current perspectives. Curr. Opin. Crit. Care 15, 149–153 (2009).
Rupnik, M., Wilcox, M. H. & Gerding, D. N. Clostridium difficile infection: new developments in epidemiology and pathogenesis. Nat. Rev. Microbiol. 7, 526–536 (2009).
Palmer, L. D. & Skaar, E. P. Transition metals and virulence in bacteria. Annu. Rev. Genet. 50, 67–91 (2016).
Hood, M. I. & Skaar, E. P. Nutritional immunity: transition metals at the pathogen–host interface. Nat. Rev. Microbiol. 10, 525–537 (2012).
Bullen, J. J. The significance of iron in infection. Rev. Infect. Dis. 3, 1127–1138 (1981).
Cassat, J. E. & Skaar, E. P. Iron in infection and immunity. Cell Host Microbe 13, 509–519 (2013).
Parrow, N. L., Fleming, R. E. & Minnick, M. F. Sequestration and scavenging of iron in infection. Infect. Immun. 81, 3503–3514 (2013).
Nakashige, T. G., Zhang, B., Krebs, C. & Nolan, E. M. Human calprotectin is an iron-sequestering host-defense protein. Nat. Chem. Biol. 11, 765–771 (2015).
Corbin, B. D. et al. Metal chelation and inhibition of bacterial growth in tissue abscesses. Science 319, 962–965 (2008).
Moschen, A. R., Adolph, T. E., Gerner, R. R., Wieser, V. & Tilg, H. Lipocalin-2: a master mediator of intestinal and metabolic inflammation. Trends Endocrinol. Metab. 28, 388–397 (2017).
Kell, D. B., Heyden, E. L. & Pretorius, E. The biology of lactoferrin, an iron-binding protein that can help defend against viruses and bacteria. Front. Immunol. 11, 1221 (2020).
McCormick, A. et al. NETs formed by human neutrophils inhibit growth of the pathogenic mold Aspergillus fumigatus. Microbes Infect. 12, 928–936 (2010).
Urban, C. F. et al. Neutrophil extracellular traps contain calprotectin, a cytosolic protein complex involved in host defense against Candida albicans. PLoS Pathog. 5, e1000639 (2009).
Deriu, E. et al. Probiotic bacteria reduce Salmonella typhimurium intestinal colonization by competing for iron. Cell Host Microbe 14, 26–37 (2013).
Bianchi, M., Niemiec, M. J., Siler, U., Urban, C. F. & Reichenbach, J. Restoration of anti-Aspergillus defense by neutrophil extracellular traps in human chronic granulomatous disease after gene therapy is calprotectin-dependent. J. Allergy Clin. Immunol. 127, 1243–1252 e1247 (2011).
Ayling, R. M. & Kok, K. Fecal calprotectin. Adv. Clin. Chem. 87, 161–190 (2018).
Barbut, F. et al. Faecal lactoferrin and calprotectin in patients with Clostridium difficile infection: a case–control study. Eur. J. Clin. Microbiol. Infect. Dis. 36, 2423–2430 (2017).
He, T. et al. Fecal calprotectin concentrations in cancer patients with Clostridium difficile infection. Eur. J. Clin. Microbiol. Infect. Dis. 37, 2341–2346 (2018).
Zackular, J. P. et al. Dietary zinc alters the microbiota and decreases resistance to Clostridium difficile infection. Nat. Med. 22, 1330–1334 (2016).
Ho, T. D. & Ellermeier, C. D. Ferric uptake regulator Fur control of putative iron acquisition systems in Clostridium difficile. J. Bacteriol. 197, 2930–2940 (2015).
Grant, C. R. et al. Distinct gene clusters drive formation of ferrosome organelles in bacteria. Nature 606, 160–164 (2022).
McAllister, K. N., Bouillaut, L., Kahn, J. N., Self, W. T. & Sorg, J. A. Using CRISPR-Cas9-mediated genome editing to generate C. difficile mutants defective in selenoproteins synthesis. Sci. Rep. 7, 14672 (2017).
Byrne, M. E. et al. Desulfovibrio magneticus RS-1 contains an iron- and phosphorus-rich organelle distinct from its bullet-shaped magnetosomes. Proc. Natl Acad. Sci. USA 107, 12263–12268 (2010).
Nitzan, O., Elias, M., Chazan, B., Raz, R. & Saliba, W. Clostridium difficile and inflammatory bowel disease: role in pathogenesis and implications in treatment. World J. Gastroenterol. 19, 7577–7585 (2013).
Sinh, P., Barrett, T. A. & Yun, L. Clostridium difficile infection and inflammatory bowel disease: a review. Gastroenterol. Res. Pract. 2011, 136064 (2011).
Wiskin, A. E., Fleming, B. J., Wootton, S. A. & Beattie, R. M. Anaemia and iron deficiency in children with inflammatory bowel disease. J. Crohns Colitis 6, 687–691 (2012).
Bou-Abdallah, F. The iron redox and hydrolysis chemistry of the ferritins. Biochim. Biophys. Acta Gen. Subj. 1800, 719–731 (2010).
Hintze, K. J. & Theil, E. C. Cellular regulation and molecular interactions of the ferritins. Cell. Mol. Life Sci. 63, 591 (2006).
Glasauer, S., Langley, S. & Beveridge, T. J. Intracellular iron minerals in a dissimilatory iron-reducing bacterium. Science 295, 117–119 (2002).
Glasauer, S. et al. Mixed-valence cytoplasmic iron granules are linked to anaerobic respiration. Appl. Environ. Microbiol. 73, 993–996 (2007).
Välikangas, T., Suomi, T. & Elo, L. L. A systematic evaluation of normalization methods in quantitative label-free proteomics. Brief. Bioinform. 19, 1–11 (2018).
Wagner, F. R. et al. Preparing samples from whole cells using focused-ion-beam milling for cryo-electron tomography. Nat. Protoc. 15, 2041–2070 (2020).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Zheng, S. et al. AreTomo: an integrated software package for automated marker-free, motion-corrected cryo-electron tomographic alignment and reconstruction. J. Struct. Biol. X 6, 100068 (2022).
Yang, J., Norris, J. L. & Caprioli, R. Novel vacuum stable ketone-based matrices for high spatial resolution MALDI imaging mass spectrometry. J. Mass Spectrom. 53, 1005–1012 (2018).
Acknowledgements
We thank members of the Skaar Laboratory for critical comments on the manuscript. This work was supported by the following grants: National Institutes of Health (NIH) grants no. R01 AI73843 (E.P.S.), no. R01 AI164587 (E.P.S.), no. U19 AI174999 (E.P.S.), no. R01 AI138581 (E.P.S. and J.M.S.), no. R00 MH113764 (Q.Z.), no. F32 AI161860 (H.P.), no. K99 AI168483 (H.P.) and no. K23 AI156132 (M.R.N.). Electron tomography was performed through the use of the Vanderbilt Institute of Nanoscale Science and Engineering Core (supported by internal VINSE pilot funding). Cryo-EM was conducted through the use of the Vanderbilt Center for Structural Biology cryo-EM Facility. The Glacios cryo-TEM was acquired by NIH grant no. S10 OD030292. Freeze-substitution was performed in part through the use of the Vanderbilt Cell Imaging Shared Resource (supported by NIH grants no. CA68485, no. DK20593, no. DK58404, no. DK59637 and no. EY08126). We thank M. Dufresne of the Vanderbilt University Mass Spectrometry Research Center for his assistance with MALDI imaging mass spectrometry experiments. We thank W. Chazin for providing purified calprotectin.
Author information
Authors and Affiliations
Contributions
H.P. and E.P.S. conceived and designed the experiments. R.S. and Q.Z. performed the cryo-EM, cryo-ET and cryo-FIB-SEM experiments. J.R.M. performed the electron tomography and cryo-FIB-SEM experiments. A.R.S.K. and J.M.S. performed the IMS experiments. K.N.G.-C. performed the histopathology analysis. E.S.K. prepared the bacterial sections using freeze-substitution. M.R.N. collected the human stool samples and clinical isolates. H.P. performed all other experiments. H.P. and E.P.S. wrote the paper with input from all authors. All authors reviewed the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Robert Britton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 FezB plays a moderate role in protection against oxidative stress.
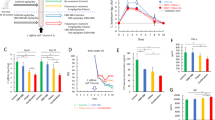
(a) The growth kinetics of C. difficile WT and ΔfezB grown in BHIS with or without treatment of 1 mM FeSO4. (b) Cells were sub-cultured in BHIS and inoculated in BHIS amended with varied concentrations of H2O2. (c) Cells were sub-cultured in BHIS supplemented with 100 µM dipyridyl and inoculated in BHIS amended with 100 µM dipyridyl plus varied concentrations of H2O2. Experiments were conducted three times independently with similar results. Data are presented as mean ± STD (n = 3).
Extended Data Fig. 2 Intracellular metal quantification using ICP-MS.
(a) The intracellular concentrations of Mg, (b) Co, (c) Cu, and (d) Zn were monitored using ICP-MS for four hours after 1 mM FeSO4 was added to cultures of C. difficile WT, ΔfezB, and ΔfezB pJS116-fezB cells grown in BHIS amended with 100 µM dipyridyl. The data are expressed as parts per million (ppm) per gram cell pellet. Experiments were conducted two times independently with similar results. Data are presented as mean ± SEM (n = 6). No significant differences were observed between strains at each timepoint based on two-tailed t tests.
Extended Data Fig. 3 C. difficile produces iron phosphate granules.
(a) EDS spectrum of a C. difficile fur::CT cell shown in Fig. 3a. The Cu signal was from the tomography holder. (b) EDS spectrum of the bacterial section shown in Fig. 3j. The fur::CT cells were stained with uranyl acetate and two distinct uranium (U) peaks were observed. Both bacterial cells and sections were deposited on 200 mesh Ni TEM grids, and strong Ni peaks were observed for both sets of samples. Two distinct Fe peaks (Lα and Kα) were evident for both sets but no other transitional metals were detected above background levels.
Extended Data Fig. 4 Clinical isolates produce ferrosomes.
(a-b) Micrographs of C. difficile isolate 14 grown in BHIS amended with 150 µM EDDHA for 4 h, followed by addition of 500 µM FeSO4 and 3 h incubation. (c-h) Representative micrographs of the same cell in panel b by (c) HAADF-STEM or (d-h) EDS maps of (d) Fe, (e) HAADF merged with Fe signal, (f) O, (g) P, and (h) Ca. Scales bars, (a) 500 nm and (b-h) 200 nm. (i) The integrated EDS spectrum of the micrograph in panel c. Two distinct Fe peaks (Lα and Kα) were detected but no other transitional metals were detected above background levels. The bacterial cells were deposited on 200 mesh Ni TEM grids, and strong Ni peaks were detected. Experiments were conducted using all 12 isolates listed in Table S1 with similar results.
Extended Data Fig. 5 Isolated ferrosomes are amorphous iron- and phosphorus-rich granules.
Images shown are (a) HAADF-STEM, (b) TEM, no crystalline structures were observed in the isolated ferrosomes, (c) HAADF-STEM, (d) Fe EDS map, (e) HAADF merged with Fe signal to show colocalization of Fe signal with electron-dense granules, and EDS maps for (f) O, (g) P, and (h) Ca. (i) The integrated EDS spectrum of the isolated ferrosomes shown in panel c. Three distinct Fe peaks (Lα, Kα, and Kβ) were observed but no other transitional metals were detected above background levels. Some background signal of Ca was detected. The samples were deposited on 200 mesh Ni TEM grids and strong Ni peaks were detected. Scales bars, (a) 200 nm, (b) 10 nm, and (c-h) 100 nm. (j) Elemental quantification based on the EDS spectrum of the isolated ferrosomes. The ratio of P versus Fe is 54:46. Experiments were conducted at least five times with similar results.
Extended Data Fig. 6 Identification of ferrosome-associated proteins using liquid chromatography mass spectrometry (LC-MS).
(a) Schematic of the proteomics experiment. Two comparisons were performed: (b) between the cell lysates of fur::CT before (cell lysate B) and after ferrosome isolation (cell lysate A), and (c) between the isolated ferrosomes from fur::CT cells and the background control (fur::CT ΔfezB). Data shown are both raw intensity and label-free quantification (LFQ) intensity. ND, not detectable. Data are presented as mean ± STD (n = 3). Statistical analyses were done using two-tailed t-test and the p values are denoted in the graphs.
Extended Data Fig. 7 Essentiality of FezB in ferrosome formation.
(a-f) Representative micrographs of a C. difficile fur::CT ∆fezB cell by (a) HAADF-STEM or (b-f) EDS maps of (b) Fe, (c) HAADF merged with Fe signal, (d) O, (e) P, and (f) Ca. (g) The integrated EDS spectrum of the fur::CT ΔfezB cell in panel a. The bacterial cells were deposited on 200 mesh Ni TEM grids, and strong Ni peaks were observed. Two distinct Fe peaks (Lα and Kα) were detected but only disordered Fe precipitates were observed in this double mutant. No other transitional metals were detected above background levels. Scales bars, 1 µm. Experiments were conducted at least six times with similar results.
Extended Data Fig. 8 The entire fez operon is required for ferrosome formation in B. subtilis 168.
(a-b) Micrographs of B. subtilis cells expressing fezAB (WT pDR111-fezAB). No electron-dense ferrosomes were observed. (c-d) Micrographs of B. subtilis cells expressing the entire fez operon (WT pDR111-fezXAB). Electron-dense granules are circled, and disordered iron precipitates are denoted by orange arrows. (a and c) Cells were grown anaerobically in LB; (b and d) Cells were grown anaerobically in LB amended with 100 µM of the iron chelator EDDHA for 4 h, followed by addition of 500 µM FeSO4 and 3 h incubation. IPTG (1 mM) was used to induce expression of either fezAB or the fez operon. (e-j) Representative micrographs of the same cell in panel d by (e) HAADF-STEM or (f-j) EDS maps of (f) Fe, (g) HAADF merged with Fe signal, (h) O, (i) P, and (j) Ca. Scales bars, (a-d) 500 nm and (e-j) 400 nm. (k) The integrated EDS spectrum of the micrograph in panel e. Two distinct Fe peaks (Lα and Kα) were detected but no other transitional metals were detected above background levels. The bacterial cells were deposited on 200 mesh Ni TEM grids, and strong Ni peaks were detected. Experiments were conducted at least three times with similar results.
Extended Data Fig. 9 Cryo-ET reveals that ferrosomes are bound by lipid-like membranes.
(a) A representative micrograph of fur::CT cells visualized using cryo-EM. The cells were processed through rapid vitrification by plunge freezing as described in Methods. A ferrosome cluster evidently present in a thin fur::CT cell was selected for cryo-ET (indicated by purple rectangle). (b) Cryo-ET reveals lipid-like membrane structures within the ferrosome cluster. The full video of the reconstructed electron tomogram is in Supplemental Video 4. (c) A ferrosome cluster was released from lysed fur::CT cells during the vitrification process and selected for cryo-ET for better visualization of the ferrosome membrane structures. (d) Cryo-ET reveals that membrane vesicles are either empty or filled with varied levels of iron phosphate granules. The full video of the reconstructed electron tomogram is in Supplemental Video 5. The membrane vesicles are annotated by purple arrows. (e) A low magnification cryo-EM image shows a FIB-milled lamella containing fur::CT cells. (f) Zoom-in view of the boxed area as indicated by purple rectangle in panel e. The boxed area in panel f was selected for cryo-ET and the reconstructed tomogram is shown in Fig. 4h and Supplemental Video 8. Scale bars, (a, f) 1 µm, (b) 100 nm, (c) 200 nm, (d) 50 nm, and (e) 2 µm. Experiments were conducted at least five times with similar results.
Extended Data Fig. 10 C. difficile resists host-mediated iron sequestration through ferrosome formation.
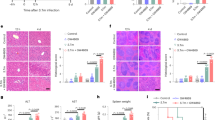
(a) Germination efficiency was examined using C. difficile WT or ΔfezB spores (n = 8). (b) Body weight of C57BL/6 J mice infected with C. difficile WT or ΔfezB spores (n = 20). Mice were treated with cefoperazone (Cfp) prior to infection. (c) Zn and (d) Mn levels were quantified by ICP-MS in fecal samples from four groups of pediatric patients: control (cancer patients), IBD, CDI, and IBD + CDI. The data are expressed as ppm (parts per million) per gram feces, n = 10 for all groups except the group of IBD + CDI (n = 8). (e-k) Infection studies were conducted using an inflammation model where mice were treated with both Cfp and DSS (dextran sulfate sodium) for 5 days (n = 20). (e) Mg, (f) Mn, (g) Zn levels (n = 10), and (h) body weight (n = 20) were monitored. (i) Histology scores were based on the areas of colon that were most severely affected as pathology was often multifocal to segmental. The scoring rubrics are: 0, not present; 1, mild, rare, scattered; 2, moderate, multifocal; 3, marked, locally extensive; and 4, marked, severe, diffuse. (j) Lipocalin (LCN−2) and (k) lactoferrin (LF) levels were quantified during infection. (l) MALDI IMS spectrum showing protein signals, including those associated with calprotectin (S100A8, m/z 10,164, indicated by a purple triangle) and an unidentified protein (m/z 13,807, annotated by a yellow triangle) from murine colon tissue from S100A9−/− (black) or C57BL/6 J (purple) mice infected with C. difficile WT spores. (Inset) Selected window from the protein MALDI IMS spectrum showing protein signal from calprotectin (S100A8, m/z 10,164 indicated with a purple arrow). (m) Body weight of S100A9−/− mice infected with C. difficile WT or ΔfezB spores (n = 25). Mice were treated with Cfp prior to infection. (e-g, j-k, and m) Fecal samples were used for all timepoints except the end timepoint (day 2 or 4), when the mice were euthanized and cecal samples were used. All data are mean ± SEM. Statistical analyses were done using (a-b, h-i, m) two-tailed t tests, (j-k) two-way ANOVA, or (c-g) one-way ANOVA and the significant p values are denoted in the graphs.
Supplementary information
Supplementary Table 4
Mass spectrometry analysis comparing the cell lysates of fur::CT before (cell lysate B) and after ferrosome isolation (cell lysate A).
Supplementary Table 5
Mass spectrometry analysis comparing isolated ferrosomes from fur::CT cells and the background control of fur::CT ΔfezB.
Supplementary Video 1
Reconstructed 3D electron tomograph demonstrating electron-dense granules are produced inside a C. difficile fur::CT cell (Fig. 3g–i). The cell is outlined in green and the granules are highlighted in red.
Supplementary Video 2
Reconstructed tomograms of intact fur::CT cells (Fig. 4a–d) show ferrosome clusters are frequently localized close to the cell membrane. Ferrosome clusters in the centre of the cell might also be close to the top or bottom membrane as they seem to be close to the fiducial markers outside the cell. Scale bar, 100 nm.
Supplementary Video 3
Further reconstructed tomograms of intact fur::CT cells (not included in the figure) show ferrosome clusters are frequently localized close to the cell membrane. Scale bar, 100 nm.
Supplementary Video 4
Cryo-ET unveils lipid-like membrane structures within a ferrosome cluster inside an intact C. difficile fur::CT cell (Extended Data Fig. 9b). Scale bar, 100 nm.
Supplementary Video 5
Cryo-ET shows empty and partially filled lipid vesicles within a cluster of ferrosomes lysed from C. difficile fur::CT cells during vitrification process (Extended Data Fig. 9d). Scale bar, 100 nm.
Supplementary Video 6
Cryo-ET shows that ferrosomes isolated from C. difficile fur::CT cells are membrane bound (Fig. 4e). Scale bar, 50 nm.
Supplementary Video 7
Cryo-ET shows that ferrosomes isolated from C. difficile fur::CT cells are membrane bound (Fig. 4f). Scale bar, 50 nm.
Supplementary Video 8
Reconstructed tomograms of FIB-milled lamella (approximately 200 nm thick) of C. difficile fur::CT cells (Fig. 4h) show ferrosome clusters are frequently localized close to the cell membrane. Empty membrane vesicles and vesicles filled with varied levels of iron phosphate minerals are evident. Scale bar, 100 nm.
Supplementary Video 9
Reconstructed tomograms of FIB-milled lamella (200 nm thick) of C. difficile fur::CT cells (Fig. 4i–l). Scale bar, 100 nm.
Supplementary Video 10
Reconstructed tomograms of FIB-milled lamella (200 nm thick) of C. difficile fur::CT cells (Fig. 4m,n). Scale bar, 100 nm.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pi, H., Sun, R., McBride, J.R. et al. Clostridioides difficile ferrosome organelles combat nutritional immunity. Nature 623, 1009–1016 (2023). https://doi.org/10.1038/s41586-023-06719-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06719-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.