Abstract
The reciprocal connections between the cerebellum and the cerebrum have been suggested to simultaneously play a role in brain size increase and to support a broad array of brain functions in primates. The cerebello-cerebral system has undergone marked functionally relevant reorganization. In particular, the lateral cerebellar lobules crura I-II (the ansiform) have been suggested to be expanded in hominoids. Here, we manually segmented 63 cerebella (34 primate species; 9 infraorders) and 30 ansiforms (13 species; 8 infraorders) to understand how their volumes have evolved over the primate lineage. Together, our analyses support proportional cerebellar-cerebral scaling, whereas ansiforms have expanded faster than the cerebellum and cerebrum. We did not find different scaling between strepsirrhines and haplorhines, nor between apes and non-apes. In sum, our study shows primate-general structural reorganization of the ansiform, relative to the cerebello-cerebral system, which is relevant for specialized brain functions in an evolutionary context.
Similar content being viewed by others
Introduction
The human cerebellum plays a role in various brain functions including executive, social, and emotional processing1,2,3,4, spanning across motor and cognitive systems. Behavioral deficits across these wide-ranging behavioral domains have also been shown in lesion studies5,6, suggesting the cerebellum does more than its canonical motor functions7. Functional magnetic resonance imaging (fMRI) studies reveal a complex cerebellar functional map with distinct activations8,9,10 and resting-state connectivity profiles11,12,13,14, distributed across the cortex. The cerebellum may encode errors, with the highly repetitive and evolutionarily conserved15,16,17,18 cerebellar circuit supporting the same computational process across behavioral domains (the universal cerebellar transform theory)19,20,21,22,23. This has prompted the idea that the location of cerebellar functions may be chiefly related to its inputs and outputs23. Nonetheless, functional diversity may exist at the algorithmic level22, with cerebellar areas integrating information from distributed brain areas24,25,26, instead of simply mapping one-to-one to inputs. In either scenario, cerebellar function depends heavily on connectivity with the rest of the brain. However, how cerebellar volume has changed over the course of primate evolution relative to one of its primary connections, the cerebrum, remains incompletely understood.
The question of how cerebellar structure and its (evolutionary) reorganization may reveal its function27,28,29 was already posed in 1906 by Louis Bolk and still motivates evolutionary studies today. So far, comparative studies of the primate cerebello-cerebral system (CCS) have indicated that CCS hyperscaling likely drives primate brain size30. Moreover, volumes31,32 and neuron numbers33,34 of primate CCS components evolve predictably. Furthermore, the rate of cerebellar volumetric35 and surface area36 evolution may have increased relative to the neocortex, specifically towards humans. Last, contemporary humans and apes have larger cerebella relative to recent human ancestors37. A reciprocal CCS connection, supportive of cerebellar learning in motor and non-motor function38,39,40,41, was first established in non-human primates (NHPs). Within the highly connected CCS, motor and prefrontal cortex (PFC) streams were largely separated into distinct projection zones23,42,43,44,45,46. Such separated information streams can be described as distinct functional modules, supporting relatively distinct brain functions. Note that this macroscale description of functional modules contrasts descriptions of the smallest operational unit that performs the cerebellar computation47.
The cerebellum is not only connected to many areas of the cerebral cortex but also to the subcortex and other areas of the body. Its macroscale evolution could thus reflect selection on a combination of behaviors including sensorimotor, vestibular, and socio-cognitive abilities, as well as changes in social niche, diet, and mode of locomotion. However, macroscale evolutionary analyses of specific functional modules48,49,50 may reflect somewhat more specific selective forces relevant to primate evolution. Although it is impossible to summarize a brain area’s function in a single term—especially in an evolutionary context – human crura I-II appear to primarily be involved in coordinating more distributed processes associated with social cognition, language, and emotion11,12,13,14,24,51. This idea is furthered by functional imaging, which reveals a strong involvement in the multi-demand and socio-linguistic networks52, together suggesting the area’s general involvement in complex integrative computation. For example, crura I-II shows fMRI activations in tasks involving language and theory of mind, but also executive and emotional processing9,10. Resting-state profiles and meta-analytic connectivity studies show that these functions may mainly come from cerebral connectivity, as the area is connected with cerebral areas also involved in such transmodal, integrative functions12,13,14,53. A cross-species homolog of human crura I-II has been referred to as the ansiform area54,55. This term is accredited to Louis Bolk’s work27 on the mammal cerebellum, where he first defined cerebellar lobular terminology29. We adopt the term ‘ansiform’ here to refer to this area. It is not guaranteed that ansiform functions established in humans will be organized in the same way in other primates. However, strong conservation of CCS structural connectivity across primates, alongside high correspondence between structural and functional connectivity, makes it reasonable to assume globally similar functions are supported by this area across primates.
The ansiform is a remarkable cerebellar area. It has a uniquely prolonged developmental trajectory56. Moreover, its disruption is often connected to brain disorders with cognitive alterations57,58,59 while developmental disruption strongly alters normal socio-cognitive development60. In the capuchin (Cebus apella), it is interconnected with frontal rather than motor cortical areas. As in all cerebello-cerebral functional modules, a reciprocal closed-loop system connects the ansiform with transmodal cerebral areas through distinct pontine, thalamic, and dentate nucleus zones23,42,43,44,45. Recent tractography work also illustrates high similarity between the human and chimpanzee CCS26. Most generally, primate ansiforms connect reciprocally to transmodal cerebral cortical areas11,12,13,14,26,51,61,62. Accordingly, evolutionary changes to the ansiform may reflect the selection of abilities supported by an ansiform-transmodal cerebral cortical module.
Reorganization63,64,65,66 of the CCS and its associated connectivity have thus been hypothesized to be important for global functional divisions within the cerebellum23,46,50. Indeed, reorganization within primate brains, rather than brain size alone, may best characterize anthropoids50,63. We focus our current study on both the whole cerebellar volume and, more specifically, the volume of the ansiform. Relatively large lateral cerebellar hemispheres (mostly consisting of the ansiform) can be noted in four vertebrate clades including primates67 and particularly hominoids68. Specifically, ansiform volume fractions of the cerebellum are larger in humans than in macaques69, and appear larger in primates than in mice and rats54. These gross anatomical expansions in primates, alongside cerebral transmodal expansions50,51,70,71,72, could reflect selection of large-scale CCS networks31,67,69,73,74 that are generally expected to support complex functional processes in cognitive, socio-linguistic, multi-demand networks52. Indeed, expansion of the ansiform from macaques to chimps to humans69 occurs alongside expansion of the PFC72, as well as the termination zone that connects them in the cerebral peduncles46. Currently, it remains unclear how the ansiform scales relative to the CCS, when taking allometry (scaling relationships) and phylogeny (evolutionary relationships) among species and traits into account.
Here we assessed primate CCS evolution through phylogenetic comparative analysis of cerebellar volumes. We used a large (34 species, 63 specimens) open MRI dataset alongside modern phylogenetic comparative methods to consolidate previous evidence of isometric (volume of region y and region x expand at the same rate) scaling between cerebellum and cerebrum31,32,75. We evaluated potential differences in slopes or intercepts of cerebellar scaling in apes and non-apes35,37, as well as strepsirrhines and haplorhines. Additionally, we aimed to assess ansiform hyper-allometry54,68,69 (hyperscaling; volume of y expands faster than x) in an allometric, phylogenetic context. Where possible, we explored intraspecific variability, substantial among brain traits68,76,77,78,79,80 and an inherent challenge for comparative primatology81. Our analyses confirmed chiefly isometric scaling between cerebellum and cerebrum. Re-analysis of the Stephan collection82 data indicated that this relation may be hypo-allometric (hyposcaling; volume of y expands slower than x), a finding supported by several robustness analyses of our data. Conversely, accounting for allometry and phylogeny, we find that primate-general ansiform hyper-allometry directly explains previous observations of this area’s large relative size in large-brained primates68,69,83.
Results
Neuroanatomical traits
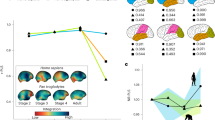
First, we report volumetric variations across the 34 primate species included in our study (Fig. 1; Table 1). Intraspecific data spread for species with enough (N ≥ 4) observations (asterisks in Fig. 1 mark species with multiple observations) is given as median absolute deviations (MADs) (Table 1). Absolute volumes were largest in apes, consistent with body size scaling.
We obtained the consensus tree for the 34 species in our dataset from 10kTrees Arnold et al.142. It represents the best-supported evolutionary relationships between the primates in our sample. Archeological epochs are superimposed on the tree to provide a temporal perspective of predicted species bifurcations. Internal node numbers are plotted and can be used to identify ancestral characters (Supplementary Table 1). Extant species included in the current study are provided on the right and are colored by clade membership. Additionally, the sample size per species is given in parentheses. Species with multiple specimens are marked with asterisks.
Intraspecific variability is substantial
For several species (N ≥ 4) we analyzed intraspecific variability. Variability, quantified as MADs as percentage of the median, ranged from 4.8 to 15.8% for cerebellar volume, and from 6.2 to 20.7% for cerebral volume. Intraspecific ansiform observations were only available for humans (median: 43.73 ± 5.00 cm3; MAD = 11.4%) and chimpanzees (median: 15.54 ± 1.93 cm3; 12.4%) (Table 1, Supplementary Figs. 1, 2). Human volumes were systematically higher in males. The chimpanzee data were generally similar between sexes (Supplementary Fig. 3).
Evolutionary model testing
Next, we evaluated fit of evolutionary models, with the best-supported model used in subsequent analyses. Akaike Information Criterion (AIC) testing84 revealed the following support for evolutionary models: Brownian Motion (BM) (λ = 1.0; AIC = 6209.62) > Early Burst (AIC = 6216.25) » BM (λ = 0.0 (star-phylogeny); AIC = 6267.87) » Ornstein-Uhlenbeck (OU) (single α; AIC = 6609.29) » OU (α per trait; AIC = 7143.13) » OU (full multivariate α; AIC = 8063.37). BM was best supported, significantly more so than Early Burst. Chi-squared testing revealed further support for the λ = 1.0 versus the λ = 0.0 model (χ2 = 53.68). Thus, we adopted BM (λ = 1.0) for subsequent analyses.
Ancestral character estimations
Next, we aimed to estimate ancestral traits based on BM evolution. We provide ancestral volumes for cerebellar, cerebral, and ansiform volumes in Supplementary Table 1 and full ancestral character estimations (ACEs) can be found on GitHub.
Ratios are confounded by allometry
To visualize the evolutionary dynamics of the CCS we mapped ancestral character estimations (ACEs) to the phylogenetic tree. We plotted 95% confidence intervals for ACEs (Fig. 2), with uncertainty intuitively increasing with time to present (Supplementary Fig. 4). Cerebellar and cerebral volumes showed virtually identical evolutionary dynamics (Fig. 2a, b), with the largest volumes in apes. Cerebellar volume at the ancestral node of the 34-species tree was estimated at 1856 mm3, resembling the ring-tailed lemur in our data. Ansiforms (Fig. 2d) were also largest in apes. We observed that ratios of cerebellum-to-cerebrum (Fig. 2c) and ansiform-to-cerebellum (Fig. 2e) may vary greatly across species (see also Table 1). However, these ratios result directly from the allometries described in (Fig. 3a, c; next section), and should thus not be interpreted functionally. We highlight how traits developed over evolutionary time more directly in phenograms (Supplementary Fig. 5).
Ancestral character estimations (ACEs) based on the Brownian Motion (BM) model of trait evolution are provided for absolute (a, b, d) and relative (c, e) volumes, alongside node-wise 95% confidence intervals (colored bars on tree nodes). ACEs for cerebellar (a), cerebral (b), and relative cerebellar-to-cerebral (c) volumes were calculated from, and mapped to, the full 34-species tree, whereas ansiform (d) and relative ansiform-to-cerebellar volumes (e) were calculated from, and mapped to, the 13-species tree. Note that the ratios in (c, e) serve merely to illustrate how ratios may result from the allometries described in Fig. 3a, c. Gradient colors on the tree represent log10-transformed brain area volumes (in mm3) (a, b, d), and volumetric fractions (in percentages) (c, e), respectively. The ancestral node was estimated at 73 million years to present. N = 34 species in (a–c) and N = 13 in (d, e).
Phylogenetic generalized least squares (PGLS) regressions for cerebellar volume regressed on cerebral volume (a, b), and ansiform regressed on rest of cerebellar (c) and cerebral (d) volumes. Median volumetric data were taken from 34-species (a) and 13-species (b–d). Log10-transformed neuroanatomical measures were plotted and overlaid with regression lines obtained in the respective PGLS models. 95% (black dotted line) and 99% (red dotted line) confidence intervals are provided alongside a fictive isometric scaling relationship with the same intercept (blue line). Exclusion of the isometric scaling relationship from the confidence intervals was taken to indicate significant allometry, as indicated by the asterisks. a, b Cerebellar volumes regressed on cerebral volumes for full data (a) and for the 13 species with complete data (b) both illustrate isometric scaling trending towards hypo-allometry. Lemuriformes and Hominidea were the two clades with the most impressive cerebellar-to-cerebral volume ratios (Fig. 2c) and were thus specifically colored here (Lemuriformes colored in a bold gray; (a–d) to show how ratios were confounded by the primate-general allometry between areas of the cerebello-cerebral system. Although species belonging to these infraorders displayed slightly higher cerebellum-to-cerebrum scaling than the primate sample as a whole (a), most of the differences in ratio resulted directly from allometry. Zooming in, (a) illustrates that while most Hominoidea, including Homo sapiens, Pongo pygmaeus, and Hylobates lar fall on the regression line of the sample, all but one of the Lemuriformes (Lemur catta; on the line) exceed the general trend. Additionally, in both clades, several species approach isometry, with one member of each falling on the isometric line (Gorilla gorilla gorilla and Mirza coquereli). (b) Illustrates reducing of the PGLS slope, connected to the smaller sample of species. c, d Ansiform volume regressed on the rest of cerebellar volume (c) and cerebral volume (d) both illustrate hyper-allometric scaling relationships. Because of strong positive allometry, species with larger cerebella and cerebra are expected to have larger relative ansiforms, directly accounting for the high ratios reported in (Fig. 2e). c, d Both illustrate that Hominoidea do not have uniquely large ansiforms. Although some Hominoidea lie above this steep regression line, so do several other—smaller—species (as colored in green; Mirza coquereli, Aotus trivirgatus, Cebus apella, and Papio hamadryas). NS = non-significant. N = 34 in (a), and N = 13 in (b–d).
Allometric scaling relationships
Accelerated ansiform scaling in primates
Next, we aimed to formally quantify scaling relationships within the CCS. More specifically, we assessed scaling between cerebellum and cerebrum, and ansiform and cerebellum or cerebrum. We employed phylogenetic generalized least squares (PGLS) regression in the context of BM evolution (Fig. 3). Median cerebellar volumes were regressed on median cerebral volume for the full 34-species data (Fig. 3a) and for the 13 species with complete data (Fig. 3b). The scaling relationships were slightly below isometry in both cases (slope34 = 0.955 and slope13 = 0.940), but statistically evolved isometrically. Ansiform volume scaled hyper-allometrically to the rest of cerebellar (ROC; Fig. 3c) and cerebral (Fig. 3d) volumes (slopeROC = 1.297; slopecerebrum = 1.245), with a lower intercept for the cerebrum (interceptcerebrum = −2.784) than for the ROC (interceptROC = −1.833). Together, the ansiform scaled hyper-allometrically to both the ROC and cerebral volumes.
No distinct allometries across major primate bifurcations
To explore divergent patterns across primate groups, we split the data into strepsirrhine (9 species) and haplorhine (25) subsets (Fig. 4a–e). We also split the data into ape and non-ape subsets for cerebellum regressions (7 apes (N = 24); 27 non-apes (N = 39)) and ansiform (5 apes (N = 22); 8 non-apes (N = 8)) (Fig. 4f–i). Main cerebellar ~ cerebral regressions are replotted in (Fig. 4a, f).
Data were split into strepsirrhine and haplorhine (a–e) and ape and non-ape (f–i) subsets. Separate PGLS regressions were performed in these groups. Importantly, none of the grade shifts were significant as assessed by phylogenetic ANCOVA (Pr > F are given for the varying slopes, intercept, or both). For strepsirrhines, a slightly higher intercept was observed for cerebellar scaling relative to the cerebrum (a; main analysis). When regressed against body mass, cerebella scaled similarly between groups (b), whereas the cerebral PGLS had a slightly lower intercept in strepsirrhines relative to haplorhines (c). This showed the opposite pattern relative to the main PGLS (a). Within the brain however, cerebellar volumes appeared to be more influential on the regression in (a). The jump in intercept for strepsirrhines in cerebellar PGLS mirrored the main PGLS (d), while cerebra scaled virtually identically between groups (e). For apes/non-apes, it appeared that ape cerebella were relatively large (f; main analysis). This (merely visual) shift was slightly accentuated by restricting the analyses to haplorhines as in Barton and Venditti35. (g). Ansiform regressions on the rest of cerebellar (h) and cerebral (i) volumes revealed no differences. Together, the lack of significant differences in any pANCOVA argue for primate-general scaling. Nstrepsirrhine = 9; Nhaplorhine = 25 (a–e); Nape = 7; and Nnon-ape = 27 (f); Nape = 7; and Nnon-ape = 18 (g); and Nape = 5 and Nnon-ape = 8 (h, i).
First, in the strepsirrhine-haplorhine analysis, we observed that although cerebellar volume scaled virtually identically across groups (Fig. 4b), cerebral volume appeared somewhat smaller relative to body mass in strepsirrhines (Fig. 4c). However, we show that relative to the rest of the brain, cerebella appeared larger in strepsirrhines (Fig. 4d). Conversely, cerebral scaling was virtually identical across groups (Fig. 4e). Differences were minor, with phylogenetic ANCOVA (pANCOVA) revealing no significant differences in scaling in any comparison. Supplementary Fig. 6 illustrates how cerebellar and cerebral volumes differ relative to brain volume across these groups due to allometry. To further evaluate how brain-body scaling may contribute to our findings, we visualized encephalization in our data (Fig. 5). We observed a characteristically high encephalization quotient (EQ) for humans, in line with previous work (6.51; Jerison’s EQ = 7.5985). We found that brain-body scaling differed across clades and was widely distributed between even closely related primates, e.g., between apes.
Here, we plotted the number of standard deviations from the brain volume–body mass regression versus body mass to visualize how they covary among clades and major primate bifurcations (a). Gray shading indicates the 95% confidence interval for the linear regression. We also replicated Jerison’s encephalization quotient (EQ) in our data, describing brain-to-body scaling relative to expectation based on allometric scaling (b). We used the allometry described by Jerison100 for mammals: \({EQ}=\frac{0.12}{{P}^{2/3}}\), with P = body mass. EQ varied greatly between clades, and somewhat less within them. Deviations from our primate brain-body regression show little within-clade consistency or relation to grouping. *Indicates suspected outliers; **indicates missing values. N = 34 species (63 observations).
Second, for apes and non-apes, we observed no significant differences between cerebellar-cerebrum PGLS regressions (Fig. 4f), even when restricting the analysis to haplorhines (Fig. 4g). No scaling differences were apparent for the ansiform (Fig. 4h, i). Fisher’s R-to-Z transformation showed that fit was always worse when fitting separate regressions for apes and non-apes (Supplementary Note 1).
Replication in the Stephan dataset
To further validate the stability of our results, we replicated PGLS analysis in an alternative set of primates, the Stephan dataset82 (Fig. 5). This dataset contains postmortem primate brains, with volumes corrected for shrinkage. BM was again the best-supported model (AIC = −19.13), with BM ≈ Early Burst (AIC = −17.51) ≈ OU (diagonal α; AIC = −16.03). It significantly outperformed the star-model (χ2 = 84.60). We adopted BM to map the ancestral character estimations (ACEs) to the tree. Cerebellar and cerebral volume ACEs displayed only slightly larger spread and clade-wise distribution was similar. PGLS for cerebellar volume regressed on cerebral volume indicated significant hypo-allometry, with its slope of approximately 0.92 being slightly reduced relative to the main analysis (Fig. 5b). This slope was again able to explain observed ratios (Fig. 5a) well.
Robustness analyses and exploration of data
Lastly, we aimed to assess robustness of our results. First, we showed that cerebellar ~ cerebral scaling differed between data from the museum collection (MNHN) and rest of the data (Supplementary Fig. 7a). Notwithstanding, the museum PGLS scaling resembled the full data PGLS: \(y=-0.274+0.977x\) (Supplementary Fig. 7b). We went on to simulate shrinkage across areas and brains (including other intraspecific variation). We showed that the cerebellar–cerebral regression supported a range of allometries: from relatively strong hypo-allometries (~0.8) to smaller hyper-allometries (~1.1) (Supplementary Fig. 7c, d). Conversely, ansiform regressions were resilient to this uncertainty and argued firmly for hyper-allometry (Supplementary Fig. 7g–j). Introducing artificial 2x intraspecific variability revealed that most PGLS analyses supported cerebello-cerebral hypo-allometry, despite intraspecific variation (Supplementary Fig. 7e, f).
Rerunning PGLS in a more conservative subset that excluded potential cerebellar volume outliers showed stronger cerebello-cerebral hypo-allometry (Supplementary Fig. 8a), tending toward significance while better matching the Stephan PGLS (Supplementary Fig. 8b). Ansiform regression on the rest of the cerebellar volume increased in hyperscaling (Supplementary Fig. 8c), while that on cerebral volume remained virtually identical (Supplementary Fig. 8d).
Lastly, considering our finding of primate-general scaling, we explored if ratio differences across clades (Fig. 2) could be directly related to body size or brain volume. Cerebellum-to-cerebrum ratios were neither related to body mass86 (R2adj. = −0.03; p = 0.88) (Supplementary Fig. 9a) nor brain volume (R2adj. = −0.01; p = 0.45) (Supplementary Fig. 9b). Conversely, ansiform-to-cerebellum ratios were related to both body mass (R2adj. = 0.50; p < 0.01**) (Supplementary Fig. 9c) and brain volume (R2adj. = 0.60; p < 0.0001****) (Supplementary Fig. 9d). This relationship between ansiform fraction and brain or body size results directly from strong positive allometric scaling.
Discussion
In the current work, we studied cerebellar volumes for 34 primate species (N = 63) and ansiform volumes for 13 species (N = 30). The cerebellum, and the ansiform specifically, are part of important cerebello-cerebral loops and networks contributing to transmodal, integrative functions12,13,14,31,39,45,46,50,61,62,69,87,88. Together, our analyses showed support for isometric-to-hypo-allometric scaling between the cerebellum and cerebrum. Conversely, the ansiform displayed strong hyper-allometry versus the rest of the cerebello-cerebral system (CCS). We did not find statistical support for different scaling between strepsirrhines and haplorhines, nor between apes and non-apes. Given this primate-general scaling, species with larger brains are expected to have larger relative ansiforms, explaining previous reports of their high ratios in chimpanzees and humans69.
Our analyses of CCS evolution showed the strongest support for Brownian Motion (BM) (λ = 1.0). This fits previous findings for the evolution of neocortical traits89 and is further supported by additional analyses in data from the Stephan collection82. The BM model, describing random incremental changes predictable by time since divergence, fits with constrained scaling of the CCS in our study. Although departures from BM have been interpreted as indicating adaptive variation90,91,92, BM does not rule out adaptation93. Recently, generalization of the BM model has been shown to represent both neutral drift and rapid adaptive evolutionary change. BM as implemented here has more difficulty describing the latter mode of evolution, because it assumes neutral and gradual change94. It remains open whether homogeneous models (a single regimen per tree, with useful statistical properties93) or heterogeneous models (different regimes on subtrees) will best elucidate primate evolutionary complexity94.
Our main PGLS supports isometric cerebello-cerebral31,32,95,96,97 evolution, tending towards hypo-allometry (Fig. 3a; ~0.95). To assess the effects of potential confounding factors, we ran several robustness analyses. These indicated that our main results generally hold when PGLS is rerun in data i) without potential outliers identified from the literature, ii) from only museum specimens, and iii) accounting for shrinkage differences between brain areas. Introducing shrinkage differences within-brain showed that cerebello-cerebral scaling may range from rather strong hypo-allometries (~0.8) to slightly weaker hyper-allometries (~1.1) (Supplementary Fig. 7c, d). Introducing severe artificial shrinkage differences across-brains showed mostly reduced allometries relative to our main PGLS (Supplementary Fig. 7e, f), as did reanalysis without literature outliers (Supplementary Fig. 7a; ~0.93) and the Stephan replication (Supplementary Fig. 7b; ~0.92). A recent study reported similar and significant hypo-allometric cerebello-cerebral surface area scaling across mammals (~0.92)97. The museum (MNHN)-only PGLS was the exception (Supplementary Fig. 7a; ~0.98), with an increased slope relative to our main analysis. Although our results might hint at a slight cerebello-cerebral hypo-allometry, it becomes clear from these data how sparsely sampled, low-n, comparative data may lead to contradictory conclusions98, especially when considering near-isometric scaling. Our robustness analyses showed that providing several complementary views on comparative data may reveal more than small sample p-values.
Previous studies have reported tight primate cerebello-cerebral scaling31,32, as well as indications that cerebella may have become relatively large95 and undergo accelerated growth in hominoidea35. Although we found no statistical evidence for relatively large cerebella in apes, restricting analyses to haplorhines (as in Barton and Venditti35) somewhat accentuates a small non-significant grade shift in apes (Fig. 4g). Additionally, apes’ positive deviations in PGLS (Figs. 3, 6) hinted slightly towards accelerated cerebellar growth in apes35. A combination of small samples, near-isometric scaling, and relatively small differences between apes and non-apes may cause conflicting conclusions on cerebello-cerebral scaling. However, our study did not consider branch-wise evolutionary rates, which appeared more strongly accelerated in apes35. In line with the notion of accelerated cerebellar growth, relatively large cerebella are noted in contemporary humans and apes, but not in recent human ancestors37.
The Stephan et al.82 collection reported cerebellar and cerebral volumes for 34 species available in 10kTrees Arnold et al.142. Ancestral character estimations (ACEs) (a) and phylogenetic generalized squares regression (PGLS) (b) were repeated in this complementary dataset. Despite only partial overlap in species, ACEs for cerebellar-to-cerebral volume ratios (a) largely mirrored the main analysis (Fig. 2c). PGLS regression between cerebellar and cerebral volumes in the Stephan collection (b) illustrated generally similar scaling as the primary analysis (Fig. 3a), albeit reduced at approximately 0.92 (black line), and with its 95% confidence intervals (black dotted line), but not 99% confidence intervals (red dotted line), excluding isometry (blue line). All apes and lemurs fell on or above the regression line, approaching isometry and mirroring the main analysis (Fig. 3a). Altogether, the analyses again show how allometry causes diverse ratios between brain areas to arise across species. Legend: apes are colored as in Fig. 3a, with legend again shown here in (b). Lemuriformes are colored here as well, with solid grays representing species overlapping with the main analysis, and solid blacks indicating lemur species unique to the Stephan dataset (a, b). Lastly, other species overlapping between both datasets, that were not specifically considered, are colored a shaded gray (a, b). N = 34 species.
Relatively large cerebella were also noted in lemurs, although this may result from smaller cerebra across strepsirrhines. Although strepsirrhines are smaller-brained and have smaller cerebra, their cerebella might be slightly larger than those of haplorhines (although not significantly), whereas cerebra scaled virtually identically relative to brain volume between groups (Fig. 4). Although brain-body scaling (Fig. 5) may be related to cognitive ability, as per the encephalization quotient hypothesis85,99,100, we argue that reorganization of behavior-supporting systems within brains may be more relevant63,64,65. Brain reorganization63,64,65,66,101 may paint a nuanced and complementary picture regarding links to comparative ability, alongside (relative) brain size85,99,100,102,103. Within the brain, both apes and lemurs appear to deviate positively from primate-general cerebello-cerebral scaling in PGLS. Future studies may seek to explore how relatively large cerebella may result from cerebral scaling, as well as investigate behavioral correlates of large cerebella. Both are likely quite different between strepsirrhines and haplorhines.
Relative to the CCS, the ansiform scaled significantly hyper-allometrically with a high slope (~1.3) (Fig. 3c, d). In contrast to cerebello-cerebral scaling, relative ansiform scaling appeared to remain hyper-allometric despite simulating unequal shrinkage across areas (~1.1 to ~1.6) (Supplementary Fig. 7g–j). Such hyperscaling likely drives much of the lateral cerebellar expansion observed in large-brained hominoidea68, and primates more generally67, and perhaps occurs alongside cerebral transmodal expansions (together part of a CCS functional module)50,73. It also explains larger ansiform fractions in larger-brained primates69,83. We found that the ansiform fraction, but not the cerebello-cerebral fraction, scales with brain volume (Supplementary Fig. 9), an expected outcome of a strong hyper-allometry. Together, these findings paint a coherent picture of primate-general hyperscaling of the ansiform, that exceeds the hyperscaling previously reported for the prefrontal cortex (1.18x relative to neocortex)72.
The implications of such hyperscaling are manifold. First, it supports the notion of modular evolution49 amidst anatomical constraints104,105 of the CCS. Specifically, a combination of developmental and functional48 constraints likely leads to tight scaling of cerebello-cerebral structures31,32. Amidst this isometric scaling, the ansiform-prefrontal link starts to scale at a steeper slope50,67,73. Small timing differences in cell division cycles have been found to explain hyperscaling of (neo)cortical surface area in primates106. Common neurodevelopmental mechanisms may thus underlie both tight cerebello-cerebral scaling across vertebrates31,32,34,71,87,88 and modular ansiform expansions107, suggesting two prominent evolutionary theories49,104 are not mutually exclusive50.
Connected to this, primate-general hyperscaling can reframe brain scaling as a matter of size. Humans may have specialized cognitive abilities and even exceptionally large relative brain size85, but evolution has not acted to make humans exempt from its laws (the scala naturæ). Although specific anatomical specializations108,109,110 may in part cause human abilities, large human brains85 may also lead to some level of behavioral emergence, through an interaction of allometries that affect whole-brain scaling. Specifically, we speculate that the combination of many primate-general allometries might lead to increasingly abundant neuroanatomical systems that support the emergence of abilities that are more than the sum of their parts111. The ansiform-transmodal cortex module clearly contributes to integrative primate behaviors10,12,13,14,24,50,67,69,112, but its evolution is primate-general. Anatomically, humans may merely represent an extreme of many allometries: their large CCSs30 having enlarged modules supporting complex and abstract functional processing51,71,72, increased white-matter connectivity46,51,113, and a large neuronal pool33,34. The same architecture at a different scale may support relatively unique behaviors.
The anything-you-can-get-basis of comparative studies warrants some consideration. For example, most species in the current work are represented by one specimen, so that much of natural variation remains unaccounted for. First, it is important to consider that intraspecific variability in any brain volume is substantial77,79,80. Small intraspecific samples can thus not be representative of the range of natural variability. Although we show robustness of our general conclusions at the primate-level, this means that deviations for individual species cannot be used to infer adaptation, since precise volumes may differ. We show variability of volumes across our data to be substantial78,79,80, as they include specimens across sexes and adult ages.
We took care to provide the most accurate volumetric measurements possible by manually segmenting cerebella. Relative to automated segmentations in CERES114, volumes were systematically higher for the cerebellum. This resulted from under-inclusion at cerebellar edges by CERES and over-inclusion at the pontine-white matter border in manual segmentation, intending to isolate the cerebellum similarly across species. Universal guidelines for comparative segmentation might help decrease differences, crucial for relatively small-sized comparative studies.
Rerunning PGLS in data excluding potential outlying cerebellar volumes from the literature31,68,82,95,115,116 (Supplementary Fig. 8) revealed that the main PGLS was robust, but had potentially somewhat overestimated cerebello-cerebral scaling. Discrepancies between these results and volumes reported across the literature show the impact of intraspecific variation.
Reassuringly, the ansiform was observed to consistently take up a large fraction of the cerebellum across humans (N = 10) and chimpanzees (N = 9), predicted by the main allometry (~1.3). Our data generally resemble prior observations for mean ansiform volumes in humans (current study: 43.94 cm3; Balsters et al.: 53.65 cm3; Makris et al.: 43.01 cm3), intraspecific variability (5.00 cm3; 8.01 cm3; 6.38cm3), and relative ansiform-to-cerebellum fraction (30.76%; 36.51%; 29.58%)69,83. Reanalysis of ansiform scaling in larger datasets will be necessary to confirm our observations as they may be partially driven by an underpowered sample and influenced by specimen-specific idiosyncrasies81,98,117.
Our study compared brain areas within specimens, helping to minimize the influence of variability across brains. Nonetheless, shrinkage could alter exact scaling formulae and would ideally be accounted for. We assumed that intraspecific variability, compounded by measurement error due to unequal shrinkage, may lead to a twofold difference in volumetric estimates77,79,80. Simulating the effect of introducing this volumetric variability in cerebello-cerebral PGLS (Supplementary Fig. 7e, f), we found generally reduced allometries relative to our main PGLS. This result was consistent with analysis of the Stephan dataset, which used shrinkage-corrected volumes82.
Although sometimes assumed to be of minor importance101, within-brain shrinkage was expected to play a larger role. Cerebellar, cerebral, and ansiform shrinkage may differ118, perhaps based on white-matter content119. Simulating this shrinkage (Supplementary Fig. 7c, d, g–j) revealed that conclusions on cerebello-cerebral scaling may differ depending on how areas have shrunk with respect to one another. This may also explain contradictory conclusions on near-isometric cerebellar scaling across datasets35. Ansiform hyperscaling was robust to this shrinkage. In the future, standard recording of fresh brain weights, and calculation of time-related shrinkage across brain areas may help combat uncertainty in volumetric estimates and, potentially, contradictory results81,98.
Systematic analysis of larger datasets containing both many species and sizable intraspecific samples might resolve previous controversies78,81,98,120,121. Creating community-wide guidelines for data collection and statistical handling can facilitate the integration of data from primate studies50,122,123,124. Statistical methods to incorporate intraspecific samples already exist125,126,127. To further data integration, databases can be used to earmark primate brains with unique identifiers, linked to relevant metadata including sex, age, body weight, captivity status, data availability, and inclusion in specific studies. Lastly, museum collections can greatly expand comparative samples128.
Summarizing, we report cerebellar volumetric data for 34 species (N = 63) and ansiforms (crura I-II) for 13 species (N = 30). Our phylogenetic analyses provide corroborating evidence for isometry-to-hypo-allometry between cerebellar and cerebral volumes31,32,97 and provide evidence for primate-general ansiform hyperscaling69,83. A combination of constrained104 and modular49 evolution may explain changes in the volume of the primate cerebello-cerebral system. The consistent finding of cerebello-cerebral coevolution argues for their structure and function being intimately linked. The strong positive allometry of the ansiform in primates, potentially alongside that of structurally connected cerebral transmodal cortices51,71,72,129, illustrates modular evolution within the primate cerebello-cerebral system. However, larger datasets are needed to elucidate exact primate scaling and determine if species or groups of species deviate from allometry more than expected.
In conclusion, within the tightly coevolving cerebello-cerebral system, the ansiform-transmodal cerebrum functional module may be a primary area of interest in the study of human and comparative cognitive correlates in the future. Large datasets, in combination with relevant metadata and sizable intraspecific samples, will increase understanding of the behavioral implications of primate hyperscaling of this module within the cerebello-cerebral system.
Methods
Data provenance and acquisition
Primate comparative data are rare and sizable datasets are difficult to obtain. We used the collection of primate species collated from different sources in the Brain Catalogue Primates project89, available on BrainBox83. It included 35 species (N = 66), which covered the primate phylogenetic tree relatively well, with 9 infraorders from strepsirrhine and haplorhine suborders. Cebidae (7 species), Hominoidea (7), Lemuriformes (6), and Papionini (5) were the most extensively sampled (Fig. 1). The data combines MRI images obtained from diverse sources, including donations, open data, and acquisitions from Heuer et al. (2019)89 (Supplementary Data 1). The current study followed the institutional review board guidelines of the corresponding institutions. Newer acquisitions included MRI scans from the Vertebrate Brain Collection of the National Museum of Natural History in Paris, France89. Scans included both fully abstracted brains and in situ brains, either full body scans or only including the skull, and were acquired on three scanners: two types of 3 T Siemens scanners (Tim Trio and Prisma) and a 11.4 T Bruker Bioscpec. Scanner. Beyond the new scans obtained by Heuer et al. (2019)89, the Brain Catalogue Primates project89 includes nine chimpanzee T1-weighted (T1w; 3 T) MRI scans from the National Chimpanzee Brain Resource (NCBR; chimpanzeebrain.org)96, two crab-eating an one rhesus macaque (T1w; 7 T) from the Pruszinsky Lab130, four crab-eating and four rhesus macaques (T1w; 3 T) from the Primate Data-Exchange (PRIME-DE)131, a crab-eating macaque and eastern gorilla downloaded from braincantalogue.org, a bonobo, gibbon, and a western lowland gorilla (T1w; 1.5 T) from the NCBR96, and finally 10 human brains (T1w; 3 T) from the ABIDE-I dataset132. ABIDE-I gained approval from the local Institutional Review Boards and had the 18 Health Insurance Portability and Accountability-protected health information identifiers removed132. No new primate brains were obtained in his study. Together, T1- or T2-weighted (T1w or T2w) MRI anatomical scans were obtained at either 1.5 T, 3 T, 7 T or 11.4 T for up to 12 h. For all brains, 3D gradient-echo sequences (FLASH) images were obtained89. MRI quality descriptors including scanning resolutions, and signal-to-noise (SNR) can be found in Supplementary Data 1. For full provenance and acquisition parameters, see Heuer et al.89.
Data preprocessing and quality control
Human132,133 and non-human primate (NHP)89 data were already preprocessed. SNRs of the primate scans be found in Supplementary Data 1. Comparative datasets often vary in data quality, due to differences in brain size, tissue conservation (in vivo, in situ, or extracted brains), MRI resolution, and SNR, as well as geometrical brain differences, and more idiotypic differences related to individual specimens. During manual segmentation (see the following) extensive visual inspection of the data was performed by NM (as well as previously by KH and RT)89. The red howler monkey was excluded due to irreparable damage. Extensive damage was also noted in the black-and-white ruffed lemur, eastern gorilla, and gibbon. Additionally, two cerebellar were not noticeably missing any tissue, but were deformed. Volumetric estimates for specimens with damaged brains need to be treated with due diligence.
Lastly, postmortem brains shrink. Comparing postmortem to fresh brain weight82 is the general practice for estimating shrinkage. Unfortunately, we could not measure fresh brain volume, nor could we specifically determine brain tissue gravity in our data, necessary to determine unequal shrinkage across areas. We did thus not correct for shrinkage. Although one would ideally correct volumes for individualized specimen- and brain area-specific shrinkage factors, determined within a controlled experimental environment, this is usually not practically feasible and to our knowledge has not been done to this point (see Stephan et al.82 for the closest approximation of this ideal). Previous comparisons between volumes obtained from in vivo MRI (Yerkes sample) and postmortem brains (Stephan sample) indicated no difference in reliability as it relates to the study of allometry68. Finally, our specific within-brain study design, comparing volume within specimens, should appreciably lessen the effect of shrinkage across brains.
Manual segmentation procedure
To provide the most accurate volumetric measurements in the highly variable primate data, we manually delineated the areas of interest directly on the MRI data134. Before this manual segmentation, we semi-automatically obtained initial cerebellar masks (serial 2D-outlines in MRI sections used to reconstruct the cerebellum in 3D). We did this by subtracting existing cerebral masks from whole brain masks with StereotaxicRAMON (github.com/neuroanatomy/StereotaxicRAMON) or using Thresholdmann (github.com/neuroanatomy/thresholdmann) to interpolate between thresholds at manually selected locations. Initial cerebellar masks were uploaded to BrainBox135 for collaborative segmentation. Lastly, we used StereotaxicRAMON for mathematical morphology operations that preserve original mask topology (Fig. 7a).
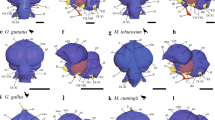
a A schematic representation of the manual segmentation pipeline. T1- and T2-weighed (T1w and T2w) MRI scans previously described in Heuer et al.89 were used to make initial cerebellar masks by subtracting cerebral masks from whole brain masks with StereotaxicRAMON. Some initial cerebellar masks were made through local thresholding and interpolation with Thresholdmann. Cerebellar masks were then uploaded to BrainBox for manual segmentation. Manual segmentation was performed iteratively with interpolation between slices through mathematical morphology operations, until segmentations reached satisfactory quality. Volumes were then downloaded with a custom python script for subsequent analysis in R. b–c Example segmentations in stereotaxic planes for the cerebellum and ansiform in the hamadryas baboon. b Cerebellar segmentations primarily involved removal of the brain stem, removing erroneously marked tissue and fissures, and reconstructing damaged tissue. c Ansiform segmentations additionally involved identification of superior posterior and ansoparamedian fissures, and segmentation between these. Segmentations were made so they did not enter vermal portions of lobule VII. Example masks are color-coded (see legend), but the meaning of the colors themselves is arbitrary. Scale bars are provided for each image.
Cerebellum
Manual segmentations (Fig. 7b) consisted of brainstem removal, outer cerebellar boundary determination, erroneously marked sulci erasure, and damaged tissue reconstruction. Segmentations (N = 65 specimens) were performed by NM, with SLV providing preliminary segmentations for a subset of specimens. Damaged tissue was reconstructed by methodical manual interpolation from adjacent, non-damaged slices. Segmentations were refined until satisfactory, using the three stereotaxic planes and alternating manual segmentation and mathematical morphology operations. All segmentations were approved by KH and SLV.
Ansiform
Segmentation (Fig. 7c) relied on identification of the superior posterior (SPF) and ansoparamedian (APMF) fissures17,69,136. Since data quality was variable, segmentation was performed in a subset of representative specimens where fissures were identifiable: 5 apes (N = 22; including 10 humans and 9 chimpanzees) and 8 non-apes (N = 8). Segmentations were again iteratively performed until satisfaction. An interobserver strategy validated segmentations made by NM. For six species, a secondary observer (MA) provided blinded segmentations alongside NM. Additionally, a tertiary observer (VS) segmented human and chimpanzee ansiforms in consultation with NM. All segmentations were checked by KH and SLV.
Reliability of segmentations
Reliability of ansiform segmentations was assessed by ANOVA:
Human cerebellar and ansiform manual segmentations were compared with automated segmentations from CERES114 at volbrain.upv.es:
and
Intraclass correlation (ICC) was additionally calculated137. ICC of 0.75–0.90 indicates good reliability and 0.90–1.00 excellent reliability138.
Cerebellar volume was significantly related to method (Pr(>F) < 0.01). ICC was .26 (95% confidence interval (CI): −0.38 < ICC < 0.74). Inspection revealed that manual segmentations were on average 20.4 cm3 (16.2%) larger than CERES segmentations. Ansiform volume was not related to method (Pr(>F) = 0.286), with ICC of 0.84 (95% CI: .51 < ICC < 0.96). Manual volumes were only 2.7 cm3 (6.6%) larger. One specimen differed by 6.9 cm3 (18.2%). It represented an average-sized cerebellum, unlikely to alter the results substantialy. Full results can be found on GitHub.
Manual ansiform volumes were significantly related to species (Pr(>F) < 0.001). No significant observer effect was detected (Pr(>F) = 0.468), with ICC of .99 (95% CI: 0.92 < ICC < 1.00).
Neuroanatomical traits
Traits of interest included cerebellar, cerebral89, and ansiform volumes (Supplementary Fig. 1). Cerebellar-to-cerebral and ansiform-to-cerebellum ratios were provided merely to show that volume ratios observed within extant primate’s CCS69,101 result directly from allometric scaling (Supplementary Fig. 1). Ratios are strongly confounded by allometry, with hyper-allometries leading to high ratios with increasing size, and vice versa. Volumes could not be corrected for shrinkage. Absolute traits were log10-transformed. Normality was examined for traits with multiple observations (≥ 4) per species (Fig. 1) by Shapiro–Wilk tests139 and outliers were visualized on boxplots (± 1.5 IQR from Q1 and Q3). This led to the exclusion from subsequent analyses of one rhesus and one crab-eating macaque (Supplementary Fig. 2).
Uncertainties in comparative data
Primate datasets are rare and generally work on an all-you-can-get-basis. Consequently, covariates known to have substantial influence on brain volumes, such as sex and age, are generally unavailable. In our study, we could not account for sex or age across the sample. Provenances of the data were diverse, potentially leading to differences in the extent of shrinkage of the included brains82,140. These sources of measurement variability may compound intraspecific variability in primate brain traits80 and together lead to large uncertainties in volumetric estimates, a problem well-known to comparative primatologists81. These challenges may lead to spurious results when not appropriately addressed98,117,141.
In our study, the comparison of traits within brains helps partially account for differences between them. Although minor relative to the cross-species scale of investigation, we investigated intraspecific variation where possible. We made sure to exclude any suspected juvenile specimens and examined the distribution of traits among the sexes in chimpanzees and humans (Supplementary Fig. 3). Lastly, we ran several robustness analyses to examine the potential effect of i) different provenances of the data, ii) shrinkage across tissues and brains, and iii) potentially outlying values based on previous literature.
Statistics and reproducibility
Consensus phylogenetic trees (the primate family structure) for the full (Fig. 1) and ansiform analyses were obtained from the 10kTrees142 primate database (10ktrees.nunn-lab.org; version 3). They were constructed from 17 genes and 7 different loci142 (10ktrees.nunn-lab.org/downloads/10kTrees_Documentation.pdf). We used R version R 4.1.0 for statistical analyses143.
Extant primate traits are not statistically independent, sharing variable amounts of evolutionary history144. Brain traits of species that diverged more recently are more likely to be similar. We detected substantial phylogenetic signal, and correlations between cerebello-cerebral traits illustrated severe collinearity (Fig. 8), demonstrating the importance of accounting for phylogeny and allometry. Therefore, before performing evolutionary analyses, we assessed what evolutionary model best described trait evolution among our species. Fit of extant traits were tested with three common evolutionary models that describe modes of evolution, using Rphylopars125. Importantly, the best-supported evolutionary model would inform all subsequent analyses. Models included Brownian Motion (BM)145, Ornstein-Uhlenbeck (OU)146 (with single alpha, alpha per trait, or full multivariate alpha matrix), and Early Burst92. First, BM describes traits varying randomly over time in direction and extent, leading to differences between species predictable by time since divergence144. Second, OU processes behave like BM with a specific trait optimum147,148. Lastly, Early Burst processes92 – which reflect rapid change after adaptive radiations (e.g., adaptation to a nocturnal niche) followed by comparative trait stability – may explain trait evolution. To assess the influence of phylogeny within the data, we chi-squared tested BM with full (Pagel’s λ = 1.0) versus a model without (λ = 0.0; star-model) phylogenetic signal. Fit was determined by Akaike information criterion (AIC; lower is better)125,149. A difference of four to seven points represents significantly less support, while a difference exceeding ten indicates no support for the worse-performing model.
Volumes recorded in the current study were correlated with the measurements from Heuer et al.89. The lower diagonal displays the correlations of log10-transformed variables, which were not corrected for phylogeny and included all specimens. These correlations serve to illustrate the strong collinearity between the traits, which are an expected outcome of allometric scaling. Corresponding R2 coefficients for the correlations can be found on the upper diagonal. Cerebellar, cerebral, and ansiform volumes correlated strongly and positively with all other variables, except fold wavelength, for which a negative correlation was observed. N = 34 species.
We next mapped extant primate traits back to their ancestral nodes along the phylogenetic tree. This ancestral character estimation (ACE) combines information from extant traits and the tree, finding values for ancestor traits consistent with the best-supported evolutionary model. In other words, if traits evolved following this model, ACEs provide estimations of traits of ancestor species. Visualizing ACEs on the consensus phylogenetic tree facilitates qualitative examination of primate-general and clade-specific evolutionary trends that result from allometric scaling.
To account for trait non-independence and reveal allometric scaling between traits, we performed phylogenetic generalized least squares (PGLS)144,150 regressions implemented in nlme151. Regressions were performed for cerebellar volume, regressed on cerebral volume for all species (1) and those with complete (including ansiform) data (2); and ansiform volume regressed on rest of cerebellar volume (3) and cerebral volume (4), all taking species medians.
To better understand potentially divergent patterns, we tested for differences in cerebello-cerebral scaling across two primate bifurcations: strepsirrhines and haplorhines, and apes and non-apes. Using a phylogenetic version of ANCOVA (pANCOVA; github.com/JeroenSmaers/evomap), separate regressions assessed the effects of allowing different intercepts (grade shifts), slopes (evolutionary rate of y relative to x), or both. High cerebellar-to-cerebral scaling may be driven by i) large cerebella or ii) small cerebra. It is known that strepsirrhines have smaller brains than haplorhines86. Therefore, we performed PGLS for cerebellar and cerebral volumes against body mass data (obtained from a recent compilation86) and brain volume data. Strepsirrhine and haplorhine scaling differences were further illustrated by ancestral reconstructions of volumes normalized against brain volume.
Since we had the a priori hypothesis that apes and non-apes would have different cerebellar and ansiform scaling, we additionally calculated R2likelihood152,153 for the full regressions, and for ape or non-ape regressions separately. Fisher’s R-to-Z transformation (cran.r-project.org/web/packages/psych/index.html) determined the fit. To improve comparability with a previous study that reported a grade shift of cerebellar-to-neocortical scaling in apes35, in a haplorrhine-only dataset, we repeated our PGLS analyses within haplorhines (N = 25 species).
Replication and robustness analyses
We repeated our main cerebellar–cerebral PGLS and ACE analyses in the Stephan collection82. Cerebellar and cerebral volumes were matched to primates in 10kTrees142. This led to a partially overlapping (species-wise) dataset of 34 species. This dataset corrected brain area volumes for shrinkage.
To examine the robustness of our main results, we ran a trio of additional analyses. First, we examined data provenance. Most data came from the MNHN, containing brains conserved for several decades. To assess whether unequal shrinkage between these and other data may have driven our results, we ran PGLS on MNHN and non-MNHN datasets separately. Secondly, we specifically examined shrinkage across areas, rerunning cerebellar–cerebral PGLS on 10.000 simulated datasets, in which we introduced random shrinkage factors between cerebellum and cerebrum of 0.91 to 1.1 (assuming 10% different shrinkage difference). In a connected step, we introduced uncertainties in volumetric estimates, consistent with potentially unaccounted-for shrinkage82, compounding intraspecific variability79,80. We again simulated 10.000 datasets, this time introducing random factors of 0.5 to 2.0 in PGLS. Thirdly, in lieu of intraspecific sample sizes for most species, we excluded potential outlier cerebellar volumes based on the literature31,68,82,95,115,116, rerunning PGLS in a more conservative subsample. This analysis excluded observations differing more than a factor two from previous observations. Differences between PGLSs were quantified through pANCOVA.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data obtained in the current project, including volumes and ratios, but also intermediate text files and figures, are uploaded to the GitHub repository accompanying this project: github.com/NevMagi/34primates_cerebellum. Input files necessary to replicate our results can also be found here, alongside full-size figures to aid legibility. The source data behind the graphs can be found in Supplementary Data 2. MRI data have been collated from different sources, and published Open Access89. The Stephan collection (https://doi.org/10.1159/000155963) was not published Open Access. Therefore, access to the data was requested by the authors through the Copy Clearance Center Marketplace and facilitated through the Central Library of Forschungszentrum Jülich GmbH. Reuse permission for these data may be requested from the publisher.
Code availability
Custom computer code was used at all stages of analysis. Some of the scripts used are adapted scripts89. Full custom code can be obtained from GitHub (github.com/NevMagi/34primates_cerebellum). Code is immediately available for usage under an Apache 2.0 License, which can be found on GitHub. A formal release of the data, as per 01-11-2023 was published to Zenodo (https://doi.org/10.5281/zenodo.10054902)154. Full accreditation of R software used in the project can be found in Supplementary Table 2 and their references155,156,157,158,159,160,161 are added at the end of the reference list, unless the software was explicitly mentioned and cited earlier in the manuscript.
References
Van Overwalle, F. et al. Consensus paper: cerebellum and social cognition. Cerebellum 19, 833–868 (2020).
Koziol, L. F. et al. Consensus paper: the cerebellum’s role in movement and cognition. Cerebellum 13, 151–177 (2014).
Mariën, P. et al. Consensus paper: language and the cerebellum: an ongoing enigma. Cerebellum 13, 386–410 (2014).
Adamaszek, M. et al. Consensus paper: cerebellum and emotion. Cerebellum 16, 552–576 (2017).
Botez, M. I., Gravel, J., Attig, E. & Vézina, J. L. Reversible chronic cerebellar ataxia after phenytoin intoxication: possible role of cerebellum in cognitive thought. Neurology 35, 1152–1157 (1985).
Schmahmann, J. D. & Sherman, J. C. The cerebellar cognitive affective syndrome. Brain 121, 561–579 (1998).
Luciani, L. Das Kleinhirn: neue Studien zur normalen und pathologischen Physiologie. (E. Besold, 1893).
Kruithof, E. S., Klaus, J. & Schutter, D. J. L. G. The human cerebellum in reward anticipation and outcome processing: an activation likelihood estimation meta-analysis. Neurosci. Biobehav. Rev. 149, 105171 (2023).
Stoodley, C. J. The cerebellum and cognition: evidence from functional imaging studies. Cerebellum 11, 352–365 (2012).
Stoodley, C. J. & Schmahmann, J. D. Functional topography in the human cerebellum: a meta-analysis of neuroimaging studies. NeuroImage 44, 489–501 (2009).
Buckner, R. L., Krienen, F. M., Castellanos, A., Diaz, J. C. & Thomas Yeo, B. T. The organization of the human cerebellum estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 2322–2345 (2011).
Xue, A. et al. The detailed organization of the human cerebellum estimated by intrinsic functional connectivity within the individual. J. Neurophysiol. 125, 358–384 (2020).
Marek, S. et al. Spatial and temporal organization of the individual human cerebellum. Neuron 100, 977–993.e7 (2018).
Guell, X., Schmahmann, J. D., Gabrieli, J. D. E. & Ghosh, S. S. Functional gradients of the cerebellum. eLife 7, e36652 (2018).
Eccles, J. C., Ito, M. & Szentágothai, J. The Cerebellum as a Neuronal Machine. The Cerebellum as a Neuronal Machine (Springer, 1967). https://doi.org/10.1007/978-3-662-13147-3.
Nieuwenhuys, R. Comparative anatomy of the cerebellum. Prog. Brain Res. 25, 1–93 (1967).
Larsell, O. & Jansen, J. The Comparative Anatomy and Histology of the Cerebellum: Vol. 2. From Monotremes through Apes. vol. 2 (University of Minnesota Press, 1970).
Larsell, O. & Jansen, J. The Comparative Anatomy and Histology of the Cerebellum: Vol. 1. From Myxinoids through Birds. vol. 1 (University of Minnesota Press, 1967).
Marr, D. & Poggio, T. From understanding computation to understanding neural circuitry. A. I. memo. 357, 1–22 (1976).
Marr, D. Vision: A Computational Investigation of Visual Representation in Man Vol. 8 (Freeman and Company, 1982).
Schmahmann, J. D. Disorders of the cerebellum: Ataxia, dysmetria of thought, and the cerebellar cognitive affective syndrome. J. Neuropsychiatry Clin. Neurosci. 16, 367–378 (2004).
Diedrichsen, J., King, M., Hernandez-Castillo, C., Sereno, M. & Ivry, R. B. Universal transform or multiple functionality? understanding the contribution of the human cerebellum across task domains. Neuron 102, 918–928 (2019).
Ramnani, N. The primate cortico-cerebellar system: anatomy and function. Nat. Rev. Neurosci. 7, 511–523 (2006).
King, M., Shahshahani, L., Ivry, R. B. & Diedrichsen, J. A task-general connectivity model reveals variation in convergence of cortical inputs to functional regions of the cerebellum. eLife 12, e81511 (2023).
Shahshahani, L., King, M., Nettekoven, C., Ivry, R. & Diedrichsen, J. Selective recruitment: Evidence for task-dependent gating of inputs to the cerebellum. https://doi.org/10.1101/2023.01.25.525395 (2023).
Chauvel, M. Singularity of the White Matter Structural Connectivity of the Human Brain Compared to the Chimpanzee Brain (Université Paris-Saclay, 2023).
Bolk, L. Das Cerebellum der Säugetiere: eine vergleichend anatomische Untersuchung (Fischer, 1906).
Glickstein, M. & Voogd, J. Lodewijk Bolk and the comparative anatomy of the cerebellum. Trends Neurosci. 18, 206–210 (1995).
Larsell, O. & Von Berthelsdorf, S. The ansoparamedian lobule of the cerebellum and its correlation with the limb-muscle masses. J. Comp. Neurol. 75, 315–340 (1941).
Smaers, J. B. & Vanier, D. R. Brain size expansion in primates and humans is explained by a selective modular expansion of the cortico-cerebellar system. Cortex 118, 292–305 (2019).
Smaers, J. B., Steele, J. & Zilles, K. Modeling the evolution of cortico-cerebellar systems in primates. Ann. N. Y. Acad. Sci. 1225, 176–190 (2011).
Whiting, B. A. & Barton, R. A. The evolution of the cortico-cerebellar complex in primates: anatomical connections predict patterns of correlated evolution. J. Hum. Evol. 44, 3–10 (2003).
Herculano-Houzel, S. The remarkable, yet not extraordinary, human brain as a scaled-up primate brain and its associated cost. Proc. Natl Acad. Sci. 109, 10661–10668 (2012).
Azevedo, F. A. C. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Barton, R. A. & Venditti, C. Rapid evolution of the cerebellum in humans and other great apes. Curr. Biol. 24, 2440–2444 (2014).
Sereno, M. I. et al. The human cerebellum has almost 80% of the surface area of the neocortex. Proc. Natl Acad. Sci. USA 117, 19538–19543 (2020).
Weaver, A. H. Reciprocal evolution of the cerebellum and neocortex in fossil humans. Proc. Natl Acad. Sci. USA 102, 3576–3580 (2005).
Doya, K. What are the computations of the cerebellum, the basal ganglia and the cerebral cortex? Neural Netw. 12, 961–974 (1999).
Shine, J. M. & Shine, R. Delegation to automaticity: The driving force for cognitive evolution? Front. Neurosci. 8, 90 (2014).
Leiner, H., Leiner, A. & Dow, R. Does the cerebellum contribute to mental skills? Behav. Neurosci. 100, 443–454 (1986).
Leiner, H., Leiner, A. & Dow, R. Cerebro-cerebellar learning loops in apes and humans. Ital. J. Neurol. Sci. 8, 423–436 (1987).
Kelly, R. M. & Strick, P. L. Cerebellar loops with motor cortex and prefrontal cortex of a nonhuman primate. J. Neurosci. 23, 8432–8444 (2003).
Dum, R. P. & Strick, P. L. An unfolded map of the cerebellar dentate nucleus and its projections to the cerebral cortex. J. Neurophysiol. 89, 634–639 (2003).
Middleton, F. A. & Strick, P. L. Cerebellar projections to the prefrontal cortex of the primate. J. Neurosci. 21, 700–712 (2001).
Strick, P. L., Dum, R. P. & Fiez, J. A. Cerebellum and nonmotor function. Annu. Rev. Neurosci. 32, 413–434 (2009).
Ramnani, N. et al. The evolution of prefrontal inputs to the cortico-pontine system: Diffusion imaging evidence from macaque monkeys and humans. Cereb. Cortex 16, 811–818 (2006).
Apps, R. et al. Cerebellar modules and their role as operational cerebellar processing units. Cerebellum 17, 654–682 (2018).
Montgomery, S. H., Mundy, N. I. & Barton, R. A. Brain evolution and development: adaptation, allometry and constraint. Proc. R. Soc. B: Biol. Sci. 283, 20160433 (2016).
Barton, R. A. & Harvey, P. H. Mosaic evolution of brain structure in mammals. Nature 405, 1055–1058 (2000).
Magielse, N., Heuer, K., Toro, R., Schutter, D. J. L. G. & Valk, S. L. A Comparative perspective on the cerebello-cerebral system and its link to cognition Cerebellum https://doi.org/10.1007/s12311-022-01495-0 (2022).
Buckner, R. L. & Krienen, F. M. The evolution of distributed association networks in the human brain. Trends Cogn. Sci. 17, 648–665 (2013).
Nettekoven, C. et al. A hierarchical atlas of the human cerebellum for functional precision mapping. https://doi.org/10.1101/2023.09.14.557689 (2023).
Riedel, M. C. et al. Meta-analytic connectivity and behavioral parcellation of the human cerebellum. NeuroImage 117, 327–342 (2015).
Luo, Y. et al. Lobular homology in cerebellar hemispheres of humans, non-human primates and rodents: a structural, axonal tracing and molecular expression analysis. Brain Struct. Funct. 222, 2449–2472 (2017).
Sugihara, I. Crus I in the rodent cerebellum: its homology to crus i and ii in the primate cerebellum and its anatomical uniqueness among neighboring lobules. Cerebellum 17, 49–55 (2018).
Liu, X. et al. A multifaceted gradient in human cerebellum of structural and functional development. Nat. Neurosci. 25, 1129–1133 (2022).
Moberget, T. et al. Cerebellar gray matter volume is associated with cognitive function and psychopathology in adolescence. Biol. Psychiatry 86, 65–75 (2019).
Dong, D. et al. Compression of cerebellar functional gradients in schizophrenia. Schizophrenia Bull. 46, 1282–1295 (2020).
Morimoto, C. et al. Volumetric differences in gray and white matter of cerebellar Crus I/II across the different clinical stages of schizophrenia. Psychiatry Clin. Neurosci. 75, 256–264 (2021).
Badura, A. et al. Normal cognitive and social development require posterior cerebellar activity. eLife 7, e36401 (2018).
Palesi, F. et al. Contralateral cerebello-thalamo-cortical pathways with prominent involvement of associative areas in humans in vivo. Brain Struct. Funct. 220, 3369–3384 (2015).
Palesi, F. et al. Contralateral cortico-ponto-cerebellar pathways reconstruction in humans in vivo: Implications for reciprocal cerebro-cerebellar structural connectivity in motor and non-motor areas. Sci. Rep. 7, 1–13 (2017).
Smaers, J. B. & Soligo, C. Brain reorganization, not relative brain size, primarily characterizes anthropoid brain evolution. Proc. R. Soc. B: Biol. Sci. 280, 20130269 (2013).
Passingham, R. E. & Smaers, J. B. Is the prefrontal cortex especially enlarged in the human brain? allometric relations and remapping factors. Brain, Behav. Evol. 84, 156–166 (2014).
Halley, A. C. & Krubitzer, L. Not all cortical expansions are the same: the coevolution of the neocortex and the dorsal thalamus in mammals. Curr. Opin. Neurobiol. 56, 78–86 (2019).
Neubert, F.-X., Mars, R. B., Thomas, A. G., Sallet, J. & Rushworth, M. F. S. Comparison of human ventral frontal cortex areas for cognitive control and language with areas in monkey frontal cortex. Neuron 81, 700–713 (2014).
Smaers, J. B., Turner, A. H., Gómez-Robles, A. & Sherwood, C. C. A cerebellar substrate for cognition evolved multiple times independently in mammals. eLife 7, e35696 (2018).
MacLeod, C. E., Zilles, K., Schleicher, A., Rilling, J. K. & Gibson, K. R. Expansion of the neocerebellum in Hominoidea. J. Hum. Evol. 44, 401–429 (2003).
Balsters, J. H. et al. Evolution of the cerebellar cortex: the selective expansion of prefrontal-projecting cerebellar lobules. NeuroImage 49, 2045–2052 (2010).
Sherwood, C. C., Bauernfeind, A. L., Bianchi, S., Raghanti, M. A. & Hof, P. R. Human brain evolution writ large and small. Prog. Brain Res. 195, 237–254 (2012).
Hill, J. et al. Similar patterns of cortical expansion during human development and evolution. Proc. Natl Acad. Sci. USA 107, 13135–13140 (2010).
Bush, E. C. & Allman, J. M. The scaling of frontal cortex in primates and carnivores. Proc. Natl Acad. Sci. USA 101, 3962–3966 (2004).
Smaers, J. B. Modeling the evolution of the cerebellum. from macroevolution to function. Prog. Brain Res. 210, 193–216 (2014).
Smaers, J. B., Steele, J., Case, C. R. & Amunts, K. Laterality and the evolution of the prefronto-cerebellar system in anthropoids. Ann. N. Y. Acad. Sci. 1288, 59–69 (2013).
Rilling, J. K. Human and nonhuman primate brains: are they allometrically scaled versions of the same design? Evol. Anthropol. 15, 65–77 (2006).
Schoenemann, P. T., Budinger, T. F., Sarich, V. M. & Wang, W. S. Y. Brain size does not predict general cognitive ability within families. Proc. Natl Acad. Sci. USA 97, 4932–4937 (2000).
Giedd, J. N. et al. Child psychiatry branch of the national institute of mental health longitudinal structural magnetic resonance imaging study of human brain development. Neuropsychopharmacology 40, 43–49 (2015).
Charvet, C. J., Darlington, R. B. & Finlay, B. L. Variation in human brains may facilitate evolutionary change toward a limited range of phenotypes. Brain Behav. Evol. 81, 74–85 (2013).
Reardon, P. K. et al. Normative brain size variation and brain shape diversity in humans. Science 360, 1222–1227 (2018).
Croxson, P. L., Forkel, S. J., Cerliani, L. & Thiebaut de Schotten, M. Structural variability across the primate brain: a cross-species comparison. Cereb. Cortex 28, 3829–3841 (2018).
Harmon, L. J. & Losos, J. B. The effect of intraspecific sample size on type I and type II error rates in comparative studies. Evolutionn 59, 2705–2710 (2005).
Stephan, H., Frahm, H. & Baron, G. New and revised data on volumes of brain structures in insectivores and primates. Folia Primatologica 35, 1–29 (1981).
Makris, N. et al. MRI-based surface-assisted parcellation of human cerebellar cortex: an anatomically specified method with estimate of reliability. Neuroimage 25, 1146–1160 (2005).
Burnham, K. P. & Anderson, D. R. Multimodel inference: Understanding AIC and BIC in model selection. Sociol. Methods Res. 33, 261–304 (2004).
Jerison, H. J. Evolution of the Brain and Intelligence (Academic Press, 1973).
Isler, K. et al. Endocranial volumes of primate species: scaling analyses using a comprehensive and reliable data set. J. Hum. Evol. 55, 967–978 (2008).
Gutiérrez-Ibáñez, C., Iwaniuk, A. N. & Wylie, D. R. Parrots have evolved a primate-like telencephalic-midbrain-cerebellar circuit. Sci. Rep. 8, 9960 (2018).
Muller, A. S. & Montgomery, S. H. Co‐evolution of cerebral and cerebellar expansion in cetaceans. J. Evol. Biol. 32, 1418–1431 (2019).
Heuer, K. et al. Evolution of neocortical folding: a phylogenetic comparative analysis of MRI from 34 primate species. Cortex 118, 275–291 (2019).
O’Meara, B. C., Ané, C., Sanderson, M. J. & Wainwright, P. C. Testing for different rates of continuous trait evolution using likelihood. Evolution 60, 922–933 (2006).
Harmon, L. J., Schulte, J. A., Larson, A. & Losos, J. B. Tempo and mode of evolutionary radiation in Iguanian lizards. Science 301, 961–964 (2003).
Harmon, L. J. et al. Early bursts of body size and shape evolution are rare in comparative data. Evolution 64, 2385–2396 (2010).
Harmon, L. Phylogenetic comparative methods: learning from trees. https://doi.org/10.32942/OSF.IO/E3XNR (2019).
Elliot, M. G. & Mooers, A. Ø. Inferring ancestral states without assuming neutrality or gradualism using a stable model of continuous character evolution. BMC Evol. Biol. 14, 226 (2014).
Rilling, J. K. & Insel, T. R. Evolution of the cerebellum in primates: differences in relative volume among monkeys, apes and humans. Brain Behav. Evol. 52, 308–314 (1998).
Rilling, J. K. & Insel, T. R. The primate neocortex in comparative perspective using magnetic resonance imaging. J. Hum. Evol. 37, 191–223 (1999).
Heuer, K. et al. Diversity and evolution of cerebellar folding in mammals. eLife 12, e85907 (2023).
Wartel, A., Lindenfors, P. & Lind, J. Whatever you want: Inconsistent results are the rule, not the exception, in the study of primate brain evolution. PLoS ONE 14, e0218655 (2019).
Jerison, H. J. The theory of encephalization. Ann. N. Y. Acad. Sci. 299, 146–160 (1977).
Jerison, H. J. Brain, body and encephalization in early primates. J. Hum. Evol. 8, 615–635 (1979).
Hanson, A., Grisham, W., Sheh, C., Annese, J. & Ridgway, S. Quantitative examination of the bottlenose dolphin cerebellum. Anat. Rec. 296, 1215–1228 (2013).
Van Schaik, C. P., Triki, Z., Bshary, R. & Heldstab, S. A. A farewell to the encephalization quotient: a new brain size measure for comparative primate cognition. Brain Behav. Evol. 96, 1–12 (2021).
Deaner, R. O., Isler, K., Burkart, J. & Van Schaik, C. Overall brain size, and not encephalization quotient, best predicts cognitive ability across non-human primates. Brain Behav. Evol. 70, 115–124 (2007).
Finlay, B. L. & Darlington, R. B. Linked regularities in the development and evolution of mammalian brains. Science 268, 1578–1584 (1995).
Yopak, K. E. et al. A conserved pattern of brain scaling from sharks to primates. Proc. Natl Acad. Sci. USA 107, 12946–12951 (2010).
Rakic, P. A small step for the cell, a giant leap for mankind: a hypothesis of neocortical expansion during evolution. Trends Neurosci. 18, 383–388 (1995).
Charvet, C. J., Striedter, G. F. & Finlay, B. L. Evo-devo and brain scaling: candidate developmental mechanisms for variation and constancy in vertebrate brain evolution. Brain Behav. Evol. 78, 248–257 (2011).
Semendeferi, K., Armstrong, E., Schleicher, A., Zilles, K. & Van Hoesen, G. W. Prefrontal cortex in humans and apes: a comparative study of area 10. Am. J. Phys. Anthropol. 114, 224–241 (2001).
Eichert, N. et al. Cross-species cortical alignment identifies different types of anatomical reorganization in the primate temporal lobe. eLife 9, e53232 (2020).
Amiez, C. et al. The relevance of the unique anatomy of the human prefrontal operculum to the emergence of speech. Commun. Biol. 6, 1–12 (2023).
Herculano-Houzel, S. The evolution of human capabilities and abilities. Cerebrum 2018, 05–18 (2018).
Wong, C. H. Y. et al. Fronto-cerebellar connectivity mediating cognitive processing speed. NeuroImage 226, 117556 (2021).
Bush, E. C. & Allman, J. M. The scaling of white matter to gray matter in cerebellum and neocortex. Brain Behav. Evol. 61, 1–5 (2003).
Romero, J. E. et al. CERES: a new cerebellum lobule segmentation method. NeuroImage 147, 916–924 (2017).
Maseko, B. C., Spocter, M. A. & Haagensen, M. & Manger, P. R. Elephants have relatively the largest cerebellum size of mammals. Anat. Rec. 295, 661–672 (2012).
Navarrete, A. F. et al. Primate brain anatomy: new volumetric MRI measurements for neuroanatomical studies. Brain Behav. Evol. 91, 109–117 (2018).
Hooper, R., Brett, B. & Thornton, A. Problems with using comparative analyses of avian brain size to test hypotheses of cognitive evolution. PLoS ONE 17, e0270771 (2022).
Quester, R. & Schröder, R. The shrinkage of the human brain stem during formalin fixation and embedding in paraffin. J. Neurosci. Methods 75, 81–89 (1997).
Kretschmann, H. J., Tafesse, U. & Herrmann, A. Different volume changes of cerebral cortex and white matter during histological preparation. Microsc Acta 86, 13–24 (1982).
Sherwood, C. C. & Smaers, J. B. What’s the fuss over human frontal lobe evolution? Trends Cogn. Sci. 17, 432–433 (2013).
Barton, R. A. & Venditti, C. Reply to Smaers: Getting human frontal lobes in proportion. Proc. Natl Acad. Sci. 110, E3683–E3684 (2013).
Bakker, R., Wachtler, T. & Diesmann, M. CoCoMac 2.0 and the future of tract-tracing databases. Front. Neuroinformatics 6, 30 (2012).
Mars, R. B. et al. Primate comparative neuroscience using magnetic resonance imaging: promises and challenges. Front. Neurosci. 8, 298 (2014).
Friedrich, P. et al. Imaging evolution of the primate brain: the next frontier? NeuroImage 228, 117685 (2021).
Goolsby, E. W., Bruggeman, J. & Ané, C. Rphylopars: fast multivariate phylogenetic comparative methods for missing data and within-species variation. Methods Ecol. Evolut. 8, 22–27 (2017).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Powell, L. E., Isler, K. & Barton, R. A. Re-evaluating the link between brain size and behavioural ecology in primates. Proc. R. Soc. B: Biol. Sci. 284, 20171765 (2017).
Herbin, M. et al. Do not dispose of historic fluid collections: evaluating research potential and range of use. Collect. Forum 34, 157–169 (2021).
Donahue, C. J., Glasser, M. F., Preuss, T. M., Rilling, J. K. & Van Essen, D. C. Quantitative assessment of prefrontal cortex in humans relative to nonhuman primates. Proc. Natl Acad. Sci. USA 115, E5183–E5192 (2018).
Arbuckle, S. A., Diedrichsen, J. & Pruszynski, J. A. Non-human primate anatomicals. https://doi.org/10.5281/ZENODO.1319671 (2018).
Milham, M. P. et al. An open resource for non-human primate imaging. Neuron 100, 61–74.e2 (2018).
Di Martino, A. et al. The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol. Psychiatry 19, 659–667 (2014).
Craddock, C. et al. The Neuro Bureau Preprocessing Initiative: open sharing of preprocessed neuroimaging data and derivatives. Front. Neuroinformatics 7 (2013).
Morey, R. A. et al. A comparison of automated segmentation and manual tracing for quantifying hippocampal and amygdala volumes. Neuroimage 45, 855–866 (2009).
Heuer, K., Ghosh, S., Robinson Sterling, A. & Toro, R. Open Neuroimaging Laboratory. Res. Ideas Outcomes 2, e9113 (2016).
Faber, J. et al. Manual sub-segmentation of the cererbellum. https://doi.org/10.1101/2022.05.09.22274814 (2022).
Gamer, M., Lemon, J., Fellows, I. & Singh, P. Irr: Various coefficients of interraterreliability and agreement. R Package Version 0.84, https://CRAN.R-project.org/package=irr (2012).
Koo, T. K. & Li, M. Y. A guideline of selecting and reporting intraclass correlation coefficients for reliability research. J. Chiropr. Med. 15, 155–163 (2016).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples) on JSTOR. Biometrika 52, 591–611 (1965).
Hedrick, B. P. et al. Assessing soft-tissue shrinkage estimates in museum specimens imaged with diffusible iodine-based contrast-enhanced computed tomography (diceCT). Microsc Microanal. 24, 284–291 (2018).
Freckleton, R. P. The seven deadly sins of comparative analysis. J. Evol. Biol. 22, 1367–1375 (2009).
Arnold, C., Matthews, L. J. & Nunn, C. L. The 10kTrees website: a new online resource for primate phylogeny. Evolut. Anthropol. 19, 114–118 (2010).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 125, 1–15 (1985).
Freckleton, R. P., Harvey, P. H. & Pagel, M. Phylogenetic analysis and comparative data: a test and review of evidence. Am. Nat. 160, 712–726 (2002).
Uhlenbeck, G. E. & Ornstein, L. S. On the theory of the Brownian motion. Phys. Rev. 36, 823–841 (1930).
Lande, R. Natural selection and random genetic drift in phenotypic evolution. Evolution 30, 314–334 (1976).
Hansen, T. F. Stabilizing selection and the comparative analysis of adaptation. Evolution 51, 1341–1351 (1997).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Control 19, 716–723 (1974).
Symonds, M. R. E. & Blomberg, S. P. in Modern Phylogenetic Comparative Methods and their Application in Evolutionary Biology 105–130 (Springer Berlin Heidelberg, 2014). https://doi.org/10.1007/978-3-662-43550-2_5.
Pinheiro, J., Bates, D. & R Core Team. nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1-163, https://CRAN.R-project.org/package=nlme (2023).
Ives, A. R. R2s for Correlated Data: Phylogenetic Models, LMMs, and GLMMs. Syst. Biol. 68, 234–251 (2019).
Ives, A. & Li, D. rr2: An R package to calculate R2s for regression models. J. Open Source Softw. 3, 1028 (2018).
Magielse, N. NevMagi/34primates_cerebellum: 01-11-2023-V1.0.1 (2023). https://doi.org/10.5281/zenodo.10054901.
Wei, T. & Simko, V. R package ‘corrplot’: Visualization of a Correlation Matrix. Version 0.92, https://github.com/taiyun/corrplot (2021).
Guillerme, T. dispRity: A modular R package for measuring disparity. Methods Ecol. Evol. 9, 1755–1763 (2018).
Wickham, H. ggplot2. (Springer International Publishing, 2016). https://doi.org/10.1007/978-3-319-24277-4.
Lenth, R. V. Least-Squares Means: The R Package lsmeans. J. Stat. Softw. 69, 1–33 (2016).
Revell, L. J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Evol. 3, 217–223 (2012).
Revelle, W. psych: Procedures for Psychological, Psychometric, and Personality Research. R Package Version 1.0–95. Evanst. Ill (2013).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
Acknowledgements
The authors would like to thank Dr. Carol MacLeod and Dr. Alexandra de Sousa for enthralling discussions about the cerebellum. We would furthermore like to thank Dr. Joshua Grant for their proofreading of the final version of the manuscript before acceptance. RT and KH are supported by the French Agence Nationale de la Recherche, projects NeuroWebLab (ANR-19-DATA-0025) and DMOBE (ANR-21-CE45-0016). KH received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement No101033485 (Individual Fellowship). Last, this work was funded in part by Helmholtz Association’s Initiative and Networking Fund under the Helmholtz International Lab grant agreement InterLabs-0015, and the Canada First Research Excellence Fund (CFREF Competition 2, 2015–2016), awarded to the Healthy Brains, Healthy Lives initiative at McGill University, through the Helmholtz International BigBrain Analytics and Learning Laboratory (HIBALL), including NM, SBE, and SLV.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
N.M.: Conceptualized the manuscript; wrote and adapted custom computer code; performed manual segmentations; performed analyses; wrote the manuscript; incorporated coauthor revisions manuscript; incorporated reviewers’ suggestions manuscript and analysis. R.T.: Provided MRI data; wrote original custom code; wrote used software; provided revision comments manuscript. V.S. and M.A. provided manual segmentations of ansiform areas for 12 and 6 specimens, respectively. S.B.E.: Provided revision comments manuscript; provided funding for the project. K.H.: Provided MRI data; provided semi-automated segmentation and software support; wrote original custom code; wrote used software; provided revision comments manuscript; provided supervision. S.L.V.: Conceptualized the manuscript; provided a subset of primary segmentations; provided revision comments on manual segmentations, the manuscript, and computer code (multiple occasions); provided supervision; provided funding for the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests
Peer review
Peer review information
Communications Biology thanks Robert Barton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Magielse, N., Toro, R., Steigauf, V. et al. Phylogenetic comparative analysis of the cerebello-cerebral system in 34 species highlights primate-general expansion of cerebellar crura I-II. Commun Biol 6, 1188 (2023). https://doi.org/10.1038/s42003-023-05553-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05553-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.