Abstract
Philadelphia chromosome (Ph)-like acute lymphoblastic leukemia (ALL) is a subset of ALL that demonstrated a high treatment failure rate. One of the hallmarks of Ph-like ALL is PDGFRB gene fusion, with fusion partner proteins often harboring dimerization domains and enhancing the kinase activity of PDGFRB. We determined a novel oncogenic PDGFRB fusion gene, NRIP1::PDGFRB, from a pediatric patient with ALL, encoding a protein with the carboxy-terminal kinase domain of PDGFRB, without the partner peptide. We confirmed the oncogenic potential of NRIP1::PDGFRB in vitro and the efficacy of all ABL1-specific inhibitor generations, including imatinib, dasatinib, nilotinib, and ponatinib, in suppressing this potential. PDGFRB activation mechanism may include juxtamembrane domain truncation in the predicted peptide. In conclusion, we determined a novel fusion gene pattern in Ph-like ALL.
Similar content being viewed by others
Introduction
The development of a risk classification strategy based on molecular subtyping has significantly improved the prognosis of childhood acute lymphoblastic leukemia (ALL) in recent decades1. Risk classification contributed to adapting appropriate treatment options, such as intensified treatment and molecular targeting agents for patients at adverse risk, or treatment with reduced intensity for patients at favorable risk. Philadelphia chromosome (Ph)-like ALL, demonstrates a gene expression profile similar to BCR::ABL1-positive ALL2 and accounts for 15%–30% of B-cell lineage ALL (B-ALL) in children and adults3. Ph-like ALL is associated with high rates of treatment resistance and relapse4. The 5-year event-free survival rates are ~60% and 80% in Ph-like ALL and other childhood ALL subtypes, respectively5. Ph-like ALL often carries oncogenic fusions of tyrosine kinases, including PDGFRB fusions. Most PDGFRB fusions include amino (N)-terminal partner protein with a dimerization motif, such as EBF1, and carboxy (C)-terminal kinase domain of PDGFRB3. The dimerization motif facilitates homodimer formation of the kinase domain, causing autophosphorylation6. Herein, we report a novel truncated form of PDGFRB without a partner protein in B-ALL and confirm its oncogenicity and sensitivity to tyrosine kinase inhibitors (TKIs).
Results
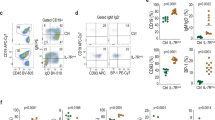
A 4-year-old patient visited the National Cancer Center Hospital in Japan with complaints of fever, malaise, and purpura. Peripheral blood examination revealed 8.4 × 1010/L white blood cells with 81% blasts (Fig. 1a), 5.6 g/dL hemoglobin, and 1.6 × 1010/L platelets. Bone marrow aspiration revealed 90% of myeloperoxidase-negative blasts. Cell surface marker profiling with flow cytometry demonstrated that the blasts were positive for CD19, CD10, CD22, cyCD79a, cy-μ chain, CD27, CD44, and CD66c and negative for T-cell and myeloid markers. The patient was diagnosed with pre-B-ALL based on these results. Figure 1b shows her clinical course. The patient was treated according to the Japanese Pediatric Leukemia/Lymphoma Study Group B-19 protocol and refractory to the early phase of induction chemotherapy, including prednisolone. Complete remission was eventually achieved after the entire induction phase, but minimal residual disease was detected after the early consolidation phase. The patient was refractory to the salvage chemotherapy with blinatumomab, a bispecific antibody against CD3 and CD19 that induces anti-tumor T-cell responses. The patient was scheduled for chimeric antigen receptor T-cell therapy followed by allogeneic hematopoietic stem cell transplantation. The patient’s parents provided informed consent for genetic analyses during induction chemotherapy.
a Representative Giemsa staining of leukemic cells. b Clinical timeline of patient’s treatment history from diagnosis; treatments at different time points are shown along the top. The blue and red lines indicate the ratio of blasts and tumor cells detected by flow cytometry (FCM). The asterisk indicates the ratio of PDGFRB FISH-positive cells. PSL prednisolone, VCR vincristine, DNR daunorubicin, L-ASP L-asparaginase, 6-MP mercaptopurine, CPM cyclophosphamide, Ara-C cytarabine, DEX dexamethasone, VP-16 etoposide. Intrathecal chemotherapy was administered throughout each treatment phase. Minimal residual disease (MRD) positivity was detected on Day 112. c Reads of NRIP1::PDGFRB fusion in each genomic locus; dashed line indicates the genomic breakpoint. d PDGFRB break-apart FISH analysis is depicted; green and red dots indicate 5' and 3' ends of PDGFRB DNA probe. e Chimeric reads of NRIP1::PDGFRB fusion in exon12 of PDGFRB; colored portions of the reads indicate mismatched bases. M methionine, W tryptophan. f Schematic representation of NRIP1::PDGFRB fusion. g Reads in PDGFRB locus; dashed line indicates the genomic breakpoint, and bottom panel shows the ratio of reads between exons before and after the breakpoint. h Expression levels of PDGFRB and the representative genes downstream of activated tyrosine kinases in ALL in Ph-like and Ph ALL groups (left, n = 7) and ALLs of other subtypes (right, n = 7); red and blue dots indicate cases with NRIP1::PDGFRB and EBF1::PDGFRB fusions. The box plots show medians (lines), interquartile ranges (IQRs; boxes), and ± 1.5 × IQRs (whiskers).
RNA sequencing (RNA-seq) of leukemia cells revealed the presence of a novel NRIP1::PDGFRB fusion gene, where an untranslated region of NRIP1 intron 3 was connected to exons 12–23 of PDGFRB (Fig. 1c). Cytogenetic analysis with fluorescence in situ hybridization (FISH) revealed a split signal of PDGFRB in 88/100 leukemic cells analyzed (Fig. 1d). Whole exome sequencing of leukemic blasts revealed no other genetic abnormalities. Interestingly, the fusion gene breakpoint resides in the middle of exon 12 of PDGFRB, followed by methionine for translation initiation (Fig. 1e). Thus, the protein resulting from NRIP1::PDGFRB has an amino acid sequence for C-terminus of PDGFRB, with a conserved tyrosine kinase domain, while it lacks N-terminal extracellular and juxtamembrane (JM) domains (Fig. 1f). The coding sequence of NRIP1 is excluded from the fusion transcript; thus, promoter swapping was considered a potential mechanism for the increased expression of truncated PDGFRB caused by this translocation. The increase in the expression levels of PDGFRB was detected between exons before and after the breakpoint, indicating the result of promoter swapping in the translocated allele (Fig. 1g). Considering the expression levels of representative genes downstream of activated tyrosine kinases in ALL7 from the data of our previous study8, our case belonged to Ph-like/Ph-positive group (Fig. 1h). Additionally, PDGFRB expression levels increased in PDGFRB rearranged cases, including this one.
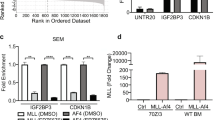
We stably transduced a murine pro-B-cell line, Ba/F3, with NRIP1::PDGFRB to validate the oncogenic potential of NRIP1::PDGFRB. Wild-type (WT) PDGFRB cDNA was also transduced as a positive control. Ba/F3 cells expressing NRIP1::PDGFRB and WT PDGFRB cDNA survived upon IL-3 withdrawal for 1 week (Fig. 2a). We detected NRIP1::PDGFRB-generated truncated form of PDGFRB and its phosphorylation (Fig. 2b), as well as the excessive phosphorylation of downstream targets of NRIP1::PDGFRB (Fig. 2d). Next, we incubated these cells with different concentrations of known ABL1 TKIs and one BRAF kinase inhibitor (vemurafenib). As demonstrated in Fig. 2c, Ba/F3 cells that express NRIP1::PDGFRB or WT PDGFRB were sensitive to all ABL1 TKI generations (imatinib, dasatinib, nilotinib, and ponatinib), with a trend toward a lower IC50 in cells expressing NRIP1::PDGFRB than WT PDGFRB. Reduced phosphorylation of downstream targets of NRIP1::PDGFRB was achieved by ABL1 TKI administration, but not by other TKIs, supporting the results of the drug sensitivity assay (Fig. 2d).
a Left panel shows Ba/F3 outgrowths transduced with NRIP1::PDGFRB compared to Ba/F3 cells transduced with a mock vector upon IL-3 withdrawal; right panel shows flow cytometry images of Ba/F3 cells transduced with a mock vector, including NRIP1::PDGFRB and PDGFRB WT; transformed clones were detected only in Ba/F3 cells transduced with NRIP1::PDGFRB and PDGFRB WT vectors; FSC forward scatter, SSC side scatter. b Western blot analyses of Ba/F3 cells transduced with NRIP1::PDGFRB or PDGFRB WT vectors; truncated form of PDGFRβ and phosphorylated PDGFRβ were shown on the left. c Ba/F3 cell sensitivities transduced with NRIP1::PDGFRB or PDGFRB WT vectors to imatinib, dasatinib, nilotinib, ponatinib, and vemurafenib; data are presented as the average of three independent experiments; vertical axis indicates viability as calculated by cell number. d Western blot analysis of Ba/F3 cells, transduced with NRIP1::PDGFRB, treated with ponatinib (1 nM) and vemurafenib (1 nM); after 24 h of tyrosine kinase inhibitor exposure, lysates were prepared and immunoblotted. e Structures of WT PDGFRB and NRIP1::PDGFRB predicted with AlphaFold2. TM transmembrane domain, JM juxtamembrane domain. f Schematic representation of PDGFRB fusion pattern; TM transmembrane domain, JM juxtamembrane domain.
Discussion
Patients with Ph-like ALL frequently show tyrosine kinase fusions, and EBF1 is a major fusion partner of PDGFRB, found in 73% of fusions3. To date, almost all fusion partners carry dimerization motifs, such as the coiled-coil domain. Fusion to the protein with dimerization motifs results in PDGFRB kinase domain homodimerization, causing kinase autophosphorylation and activation. Such a response potentiates RAS/MAPK and PI3K pathways and promotes cell proliferation6.
In contrast, the encoded protein by NRIP1::PDGFRB lacks a partner protein with a dimerization domain, although we confirmed its growth-inducing ability through excessive autophosphorylation. Although rare, hematological malignancies demonstrated PDGFRB fusions without a dimerization protein9. The partner proteins of PDGFRB in the fusion protein encoded by DTD1::PDGFRB, MRC1::PDGFRB fusions10,11, and G3BP1::PDGFRB demonstrated no dimerization domains while the oncogenic ability has been experimentally confirmed12.
The characteristics of these fusion protein types are truncated JM domain, detected in our case. JM domain in tyrosine kinase receptors has been reported as an autoinhibitory domain that suppresses kinase activity through conformational proximity13. Additionally, some tyrosine kinase families, other than PDGFRB, are activated by JM dysfunction. FLT3 and KIT have crystal structures similar to PDGFRB. An internal tandem duplication (ITD) of JM domain in FLT3 causes a structural alteration of JM domain, resulting in constitutive activation of its enzymatic function and cell proliferation14. FLT3-ITD alterations occur in acute myeloid leukemia (AML), accounting for ~30% of AML cases15. Mutations in JM domain of KIT in gastrointestinal stromal tumors (GIST) demonstrated a similar activation mechanism16. TKIs are effective and clinically used for FLT3-ITD-positive AML and KIT rearranged GISTs17,18. FIP1L1::PDGFRA fusion, one of the major oncogenic fusion genes of myeloproliferative disorders, is another example of JM dysfunction. The translated protein contains truncated JM and kinase domains of PDGFRA19. Similarly, proliferative ability20 and response to TKI21 have been demonstrated.
Importantly, Stover et al. revealed an increase in enzymatic ability with the absence of tryptophan-566 (W566) in JM region of PDGFRB20, and Chen et al. reported the crucial role of W566 in maintaining JM domain assembly22. The protein encoded by NRIP1::PDGFRB, in our case, lacks JM domain (Fig. 2e); the fusion transcript excluded the sequence encoding W566 (Fig. 1e). A recent study reported a novel PDGFRB fusion gene, CD74::PDGFRB, in Ph-like ALL in addition to the known patterns of PDGFRB fusions (Fig. 2f)23. The sequence encoding W566 was conserved in the transcript, but PDGFRB translation starts from the same translation start site as NRIP1::PDGFRB, causing the same form of JM truncated PDGFRB protein. Specifically, they experimentally revealed that the truncated PDGFRB without W566 harbors a stronger kinase activity than truncated PDGFRB retaining W566. Additionally, they revealed that CD74::PDGFRB did not dimerize as strongly as EBF1::PDGFRB, a representative PDGFRB fusion gene with partner protein harboring dimerization domain. PDGFRB protein with truncated JM results in excessive downstream phosphorylation as shown in our case, but dimerization may not be necessary for PDGFRB autophosphorylation in JM dysregulated cases. Altogether, truncated JM is a novel oncogenic form of PDGFRB aberration in Ph-like ALL.
The 5-year event-free survival of Ph-like ALL with PDGFRB rearrangement was 50%24. Accumulating reports indicated the efficacy of TKIs against Ph-like ALL, including those with PDGFRB rearrangement25,26,27, although they remained prospectively not validated. Currently, an ongoing prospective trial aims to confirm the efficacy of dasatinib in patients with Ph-like ALL with specific fusions (Children’s Oncology Group’s AALL1131, NCT02883049). We and others23 confirmed the proliferative capacity and response to TKIs in JM dysregulated PDGFRB; thus, our data will be beneficial for future patient selection. In conclusion, our study identified a novel truncated PDGFRB fusion in Ph-like ALL without fusion partner peptides which can be targeted by TKIs.
Methods
Sample
We used a bone marrow aspiration specimen for RNA sequencing (RNA-seq). The patient’s parents signed a written informed consent for genetic analyses and publication of the case report. The National Cancer Center Research Ethics Review Board approved this study (2015-059). We followed the ethical principles of the Declaration of Helsinki.
RNA sequencing
We extracted total RNA from the bone marrow sample and prepared and subjected RNA-seq libraries to next-generation sequencing as previously described28. We used Arriba to detect gene fusion29.
Primers for NRIP1::PDGFRB fusion
We identified NRIP1::PDGFRB fusion transcript by cDNA PCR from the patient sample using the following primer sets:
NRIP1 forward: TTGGATTGTGAGCTATTTCAGAAC
PDGFRB reverse: AGGGTTTGGGGCACAACACGTCAG
Cell culture
The wild-type PDGFRB cDNA and NRIP1::PDGFRB cDNA coding regions were inserted into the pMXS plasmid. Ba/F3 cells were infected with the generated retroviruses from each plasmid.
Drug sensitivity assay
Cells were seeded into 96-well plates at a 100 μL volume. After overnight incubation, cells were treated with each drug, including imatinib (Selleck), dasatinib (Selleck), nilotinib (Selleck), ponatinib (Selleck), and vemurafenib (Selleck), at doses ranging from 0.1 nM to 1 μM, incubated for 72 h. Subsequently, 10 μL of PrestoBlue (Thermo Fisher Scientific) was added to the plates, and the fluorescence was measured after 3 h of incubation.
Clinical sequence data
Sequencing data of Japan Adult Leukemia Study Group (JALSG) B-ALL clinical samples were obtained from the Japanese Genotype–Phenotype Archive (accession JGAS00000000047)8, which is hosted by the DNA Databank of Japan.
Western blot
Standard protocols were used for protein detection by immunoblot analysis, using primary antibodies PDGFRβ (#3169, 1:1000 dilution), phospho-PDGFRβ (Tyr751) (#3161, 1:1000 dilution), Akt (#4691, 1:1000 dilution), phospho-Akt (Ser473) (#4060, 1:1000 dilution), Erk1/2 (#4695, 1:1000 dilution), phospho-Erk1/2 (#4370, 1:1000 dilution), and β-Actin (#4970, 1:1000 dilution) purchased from Cell Signaling Technology. Uncropped immunoblots blots of each Figure are included in Supplementary Fig. 1.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Sequencing data is deposited at Gene Expression Omnibus (GEO) under the accession number GSE242858.
References
Hunger, S. P. & Mullighan, C. G. Acute lymphoblastic leukemia in children. N. Engl. J. Med. 373, 1541–1552 (2015).
Mullighan, C. G. et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N. Engl. J. Med. 360, 470–480 (2009).
Harvey, R. C. & Tasian, S. K. Clinical diagnostics and treatment strategies for Philadelphia chromosome-like acute lymphoblastic leukemia. Blood Adv. 4, 218–228 (2020).
Roberts, K. G. et al. High frequency and poor outcome of philadelphia chromosome-like acute lymphoblastic leukemia in adults. J. Clin. Oncol. 35, 394–401 (2017).
Loh, M. L. et al. Tyrosine kinome sequencing of pediatric acute lymphoblastic leukemia: a report from the children’s oncology group TARGET project. Blood 121, 485–488 (2013).
Guérit, E., Arts, F., Dachy, G., Boulouadnine, B. & Demoulin, J. B. PDGF receptor mutations in human diseases. Cell Mol. Life Sci. 78, 3867–3881 (2021).
Harvey, R. C. et al. Development and validation of a highly sensitive and specific gene expression classifier to prospectively screen and identify B-Precursor Acute Lymphoblastic Leukemia (ALL) patients with a Philadelphia Chromosome-Like (“Ph-like” or “BCR-ABL1-Like”) signature for therapeutic targeting and clinical intervention. Blood 122, 826–826 (2013).
Yasuda, T. et al. Recurrent DUX4 fusions in B cell acute lymphoblastic leukemia of adolescents and young adults. Nat. Genet. 48, 569–574 (2016).
Walz, C. et al. Characterization of three new imatinib-responsive fusion genes in chronic myeloproliferative disorders generated by disruption of the platelet-derived growth factor receptor β gene. Haematologica 92, 163–169 (2007).
Gosenca, D. et al. Identification and functional characterization of imatinib-sensitive DTD1-PDGFRB and CCDC88C-PDGFRB fusion genes in eosinophilia-associated myeloid/lymphoid neoplasms. Genes Chrom. Cancer 53, 411–421 (2014).
Eissa, S. S. et al. Dasatinib induces a dramatic response in a child with refractory juvenile xanthogranuloma with a novel MRC1-PDGFRB fusion. Blood Adv. 4, 2991–2995 (2020).
Jan, M. et al. A cryptic imatinib-sensitive G3BP1-PDGFRB rearrangement in a myeloid neoplasm with eosinophilia. Blood Adv. 4, 445–448 (2020).
Hubbard, S. R. Juxtamembrane autoinhibition in receptor tyrosine kinases. Nat. Rev. Mol. Cell Biol. 5, 464–471 (2004).
Kiyoi, H., Ohno, R., Ueda, R., Saito, H. & Naoe, T. Mechanism of constitutive activation of FLT3 with internal tandem duplication in the juxtamembrane domain. Oncogene 21, 2555–2563 (2002).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Corless, C. L., Fletcher, J. A. & Heinrich, M. C. Biology of gastrointestinal stromal tumors. J. Clin. Oncol. 22, 3813–3825 (2004).
Daver, N., Schlenk, R. F., Russell, N. H. & Levis, M. J. Targeting FLT3 mutations in AML: review of current knowledge and evidence. Leukemia 33, 299–312 (2019).
Casali, P. G. et al. Time to definitive failure to the first tyrosine kinase inhibitor in localized GI stromal tumors treated with imatinib as an adjuvant: a european organisation for research and treatment of cancer soft tissue and bone sarcoma group intergroup randomized trial in collaboration with the australasian gastro-intestinal trials group, UNICANCER, French Sarcoma Group, Italian Sarcoma Group, and Spanish Group for research on sarcomas. J. Clin. Oncol. 33, 4276–4283 (2015).
Cools, J. et al. A tyrosine kinase created by fusion of the PDGFRA and FIP1L1 genes as a therapeutic target of imatinib in idiopathic hypereosinophilic syndrome. N. Engl. J. Med. 348, 1201–1214 (2003).
Stover, E. H. et al. Activation of FIP1L1-PDGFRα requires disruption of the juxtamembrane domain of PDGFRα and is FIP1L1-independent. Proc. Natl Acad. Sci. USA 103, 8078–8083 (2006).
Khoury, J. D. et al. The 5th edition of the World Health Organization classification of haematolymphoid tumours: myeloid and histiocytic/dendritic neoplasms. Leukemia 36, 1703–1719 (2022).
Chen, J. et al. Positive and negative regulatory roles of the WW-like domain in TEL-PDGFbetaR transformation. Blood 104, 535–542 (2004).
Sadras, T. et al. Unusual PDGFRB fusion reveals novel mechanism of kinase activation in Ph-like B-ALL. Leukemia 37, 905–909 (2023).
den Boer, M. L. et al. Outcomes of paediatric patients with B-cell acute lymphocytic leukaemia with ABL-class fusion in the pre-tyrosine-kinase inhibitor era: a multicentre, retrospective, cohort study. Lancet Haematol. 8, e55–e66 (2021).
Roberts, K. G. et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N. Engl. J. Med. 371, 1005–1015 (2014).
Tanasi, I. et al. Efficacy of tyrosine kinase inhibitors in Ph-like acute lymphoblastic leukemia harboring ABL-class rearrangements. Blood 134, 1351–1355 (2019).
Zhang, X. et al. Pediatric acute lymphoblastic leukemia with Pdgfrb fusions: a multicentre retrospective study. Blood 140, 6137–6139 (2022).
Tanaka, Y. et al. Transcriptional activities of DUX4 fusions in B-cell acute lymphoblastic leukemia. Haematologica 103, e522–e526 (2018).
Uhrig, S. et al. Accurate and efficient detection of gene fusions from RNA sequencing data. Genome Res. 31, 448–460 (2021).
Acknowledgements
We are grateful to Hitoshi Ichikawa, Sachiyo Mitani, Maiko Matsuda, Erika Arakawa, Reina Takeyama, and the medical staff of the National Cancer Center Hospital. We thank the patient and the patient’s family who contributed to this study. This study was supported in part by grants from the Grant-in-Aid for Scientific Research under Grant Number 21K12117 (to T.U.) and National Cancer Center Research and Development Fund 2020-J-2 for NCC Biobank and NCC Core Facility (to K.S.).
Author information
Authors and Affiliations
Contributions
H.M., C.O., and Y.T. designed the study. Y.T., T.U., and S.Koj. performed sequencing data analyses. B.M. performed functional assays. B.M., M.S., A.A., K.Tao., and K.Tan. engaged in patient care. K.S., S.Y., S.Koh., M.K., N.K., Y.G., Y.Y., and A.H. performed library preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Miyazaki, B., Ueno, T., Sugiyama, M. et al. Promoter swapping of truncated PDGFRB drives Ph-like acute lymphoblastic leukemia. npj Precis. Onc. 7, 132 (2023). https://doi.org/10.1038/s41698-023-00485-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-023-00485-7