Abstract
The aim to investigate the predictive efficacy of automatic breast volume scanner (ABVS), clinical and serological features alone or in combination at model level for predicting HER2 status. The model weighted combination method was developed to identify HER2 status compared with single data source model method and feature combination method. 271 patients with invasive breast cancer were included in the retrospective study, of which 174 patients in our center were randomized into the training and validation sets, and 97 patients in the external center were as the test set. Radiomics features extracted from the ABVS-based tumor, peritumoral 3 mm region, and peritumoral 5 mm region and clinical features were used to construct the four types of the optimal single data source models, Tumor, R3mm, R5mm, and Clinical model, respectively. Then, the model weighted combination and feature combination methods were performed to optimize the combination models. The proposed weighted combination models in predicting HER2 status achieved better performance both in validation set and test set. For the validation set, the single data source model, the feature combination model, and the weighted combination model achieved the highest area under the curve (AUC) of 0.803 (95% confidence interval [CI] 0.660–947), 0.739 (CI 0.556,0.921), and 0.826 (95% CI 0.689,0.962), respectively; with the sensitivity and specificity were 100%, 62.5%; 81.8%, 66.7%; 90.9%,75.0%; respectively. For the test set, the single data source model, the feature combination model, and the weighted combination model attained the best AUC of 0.695 (95% CI 0.583, 0.807), 0.668 (95% CI 0.555,0.782), and 0.700 (95% CI 0.590,0.811), respectively; with the sensitivity and specificity were 86.1%, 41.9%; 61.1%, 71.0%; 86.1%, 41.9%; respectively. The model weighted combination was a better method to construct a combination model. The optimized weighted combination models composed of ABVS-based intratumoral and peritumoral radiomics features and clinical features may be potential biomarkers for the noninvasive and preoperative prediction of HER2 status in breast cancer.
Similar content being viewed by others
Introduction
Globally, breast cancer is the malignant disease with the highest incidence in women1. Breast cancer is a highly heterogeneous disease, and individualized clinical decision-making is particularly vital. Human epidermal growth factor receptor 2 (Human epidermal growth factor receptor2, HER2) positive is a molecular subtype of breast cancer. Compared with other subtypes, HER2-positive breast cancer has stronger heterogeneity, poorer prognosis, and lower survival rate2. HER2 positivity has been confirmed to promote tumor neovascularization and lymphangiogenesis, thereby affecting tumor growth and metastasis3. Chemotherapy combined with trastuzumab and pertuzumab is currently a first-line targeted therapy program for HER2-positive patients and can significantly improve patient outcomes4. Therefore, HER2 positive can reflect the information on tumor growth and metastasis, and its accurate judgment is crucial for the treatment and prognosis of breast cancer.
Automatic breast volume scanner (ABVS) automatically obtained three-dimensional ultrasound images of the breast using a standardized procedure. ABVS can obtain continuous ultrasound images, which are not influenced by the sonographer's operating technique and clinical experience. ABVS was found to have a higher value than conventional ultrasound in breast cancer screening5, observer consistency6, and preoperative prediction of neoadjuvant chemotherapy response7. Radiomics can analyze the potential relationship between medical imaging and tumor phenotype through high-throughput extraction of a large number of medical imaging features8. Previous studies on molecular subtypes of breast cancer had focused on the internal features of breast tumors, ignoring the peritumoral information9,10. Peritumoral tissues contain higher levels of markers for transcription factor activators, angiogenesis, proliferation, and invasion than tumor tissues, which determine tumor recurrence11,12. Studies had also demonstrated that peritumoral features were associated with the tumor necrosis factor (TNF) signaling pathway, which was involved in tumor angiogenesis, invasion, and metastasis13. Thus, the peritumoral region may embed biological information about tumor growth, invasion, and metastasis and may be a potential biomarker. In addition, studies also confirmed differences in Magnetic resonance imaging (MRI) findings of intratumoral and peritumoral among different molecular subtypes of breast cancer14. Therefore, the utilization of intratumoral and peritumoral features to predict HER2 status should have strong theoretical feasibility.
Some recent studies that predicted HER2 status in breast cancer focused only on radiomics features and did not involve clinically relevant data, which might affect model performance15,16. Model ensemble was an important strategy to optimize models, but the method of model weighted combination belonged to the model ensemble category was less applied in the field of predicting of HER2 status in breast cancer. Therefore, the present study aimed to construct models to predict the HER2 status of breast cancer by utilizing model weighted combination and feature combination methods based on ABVS intratumoral and peritumoral imaging features and clinically relevant features, to obtain the optimal model and provide the best basis for clinical decision-making for breast cancer patients.
Materials and methods
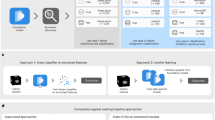
The study was conducted in accordance with the Declaration of Helsinki and approved by the Review Board of the First Hospital of Lanzhou University. Because of the retrospective study, the Ethics Committee of the First Hospital of Lanzhou University exempted written informed consent. Figure 1 showed a flow chart of the research protocol.
Overview of research protocol. Notes: R3mm, model based on peritumoral 3 mm ring of breast tumor; R5mm, model based on peritumoral 5 mm ring of breast tumor; Tumor, model based on radiomics features of the tumor; R5mm+Clinical, model based on radiomics features of the peritumoral 5 mm ring of breast tumor combined with clinical, ABVS, and serology features of breast tumor; R3mm+Clinical, model based on radiomics features of the peritumoral 3 mm ring of breast tumor combined with clinical, ABVS and serology features of breast tumor; Tumor+Clinical, model based on radiomics features of the tumor combined with clinical, ABVS and serology features of breast tumor; Tumor + R5mm, model based on radiomics features of the tumor and those of peritumoral 5 mm ring of breast tumor; Tumor+R3mm, model based on radiomics features of the tumor and those of peritumoral 3 mm ring of breast tumor. ABC, Ada Boosting Classifier; Clinical, Model based on clinical, ABVS, and serology features of breast tumor; ETC, Extra Tree Classifier; GBC, Gradient Boosting Classifier; LGBM, Light Gradient Boosting Machine; RFC, Random Forest Classifier.
Patients
174 patients with invasive breast cancer confirmed in the First Hospital of Lanzhou University from July 1th, 2016 to April 30th, 2022 and 97 patients with invasive breast cancer confirmed in the Ningxia Hui Autonomous Region People's Hospital were collected in this study, which were conducted on May 1–5th, 2022. We had access to the information identifying each patient during or after data collection. Of these, 174 patients in our hospital were randomly divided into the training set and validation set (ratio 8:2), and 97 patients in the external hospital as the test set. Inclusion criteria were: (1) female patients aged between 18 and 80 years; (2) pathologically confirmed invasive breast cancer; (3) ABVS examination before treatment. Exclusion criteria were: (1) radiation therapy, neoadjuvant chemotherapy, or interventional therapy before ABVS examination (n = 48); (2) incomplete clinical, pathological, and serological information (n = 15); and (3) significant artifacts of the tumor area on ABVS images (n = 8). Finally, 174 patients with invasive breast cancer (all female, mean ± standard deviation: 48.8 ± 10.8 years) were included (Fig. 2, Table 1).
Data acquisition
All patients underwent continuous cross-sectional scanning of each breast (interlayer spacing set at 0.5 mm) using a 14L5BV probe (7 MHz, dynamic range 50–55 dB) and a 14L5 linear probe (7–14 MHz frequency range and 10 MHz frequency center) of the ABVS (Siemens, AusonS2000, Munich, Bavaria, Germany), and the acquired axial plane images were transmitted to the workstation to automatically reconstruct sagittal plane and coronal plane images. All examinations were performed by a sonographer with 8 years of experience in ABVS examination. The results of immunohistochemistry (IHC) or fluorescence in situ hybridization (FISH) of breast cancer were considered the reference standard for HER2 status. The following clinical and serological indicators of patients were obtained from the electronic medical record system: age, erythrocytes, hemoglobin, hematocrit, mean erythrocyte volume, mean hemoglobin content, mean hemoglobin concentration, erythrocyte distribution width (standard deviation [SD]), erythrocyte distribution width (coefficient of variation [CV]), leukocytes, percentage of lymphocytes, percentage of monocytes, percentage of neutrophils, percentage of eosinophils, percentage of basophils, the absolute value of lymphocytes, the absolute value of monocytes, the absolute value of neutrophils, the absolute value of eosinophil, the absolute value of basophil, platelets, platelet ratio, mean platelet volume, platelet distribution width, large platelet ratio, cancer antigen 153(CA153), cancer antigen 125(CA125), carcinoembryonic antigen (CEA), total bilirubin, direct bilirubin, and indirect bilirubin.
Region of interest (ROI) marking and ABVS ultrasonic feature extraction
Tumor ROIs were obtained through continuous manual 3D segmentation of breast tumors in the ABVS axial plane by two sonographers (with 8 years and 5 years of ABVS experience) using 3DSlicer version 4.11.2 (BWH, Boston, Massachusetts, USA), and then the ROIs of the 3 mm peritumoral and 5 mm peritumoral were acquired through the 3DSlicer editing function. Two sonographers assessed and recorded ABVS ultrasound features based on the Breast Imaging Reporting and Data System (Breast Imaging Reporting and Data System, BI-RADS), including tumor maximum diameter in the coronal plane, margin, shape, aspect ratio, halo, internal composition, echo, microcalcification, and convergence sign (coronal plane). For undetermined cases, the two sonographers reached a consensus through consultation. Two sonographers were unaware of the HER2 status of breast cancer.
Radiomics feature extraction and model construction
The radiomics features of tumor ROI, 3mm peritumoral ROI, and 5mm peritumoral ROI were extracted using the Radiomics module of IntelliSpace Medicina Scientia (ISMS) version 2.4.0 (Philips Healthcare, Beijing, China) developed based on pyradiomics17. Image types included original, log, and wavelet images. Feature classes contained three-dimensional shape, neighborhood gray-tone difference matrix (Neighboring gray-tone difference matrix, NGTDM), gray dependence matrix (Gray-level dependence matrix, GLDM), gray level co-occurrence matrix (Gray-level co-occurrence matrix, GLCM), first sequence (First order), gray-level run-length matrix (Gray-level run-length matrix, GLRLM) and gray level area matrix (Gray-level size zone matrix, GLSZM). Given the relatively small proportion of HER2-positive cases (45/139) in the training set, the models were trained using the method of oversampling18. Through the Automatic Machine Learning (AML) function of ISMS version 2.4.0, models were constructed based on the radiomics features of the tumor, 3mm peritumoral region, 5mm peritumoral region, and the clinical features (clinical, ABVS ultrasound, and serological features), which were named as Tumor model, R3mm model, R5mm model, and Clinical model, respectively. Among the 13 classifiers of ISMS software, the best classifiers of four types of data sources (the highest sum of the AUC of the training set and the validation set of the classifiers) were selected for constructing four types of data source models.
Radiomics model construction and optimization
The model weighted combinationand feature combination methods were used to construct and optimize the radiomics models. First, based on the four types of data source models, the weighted combination models of Tumor combined with Clinical (Tumor + Clinical), Tumor combined with R3mm (Tumor + R3mm), Tumor combined with R5mm (Tumor + R5mm), R3mm combined with Clinical (R3mm + Clinical), and R5mm combined with Clinical (R5mm + Clinical) were constructed and optimized using the method of a weighted combination of two data source models. In the validation set, alpha-AUC scatter plots of weighted models were plotted depending on the weighting coefficients (alpha), where two model results were combined using alpha*model 1 + (1-alpha) * model 2, to determine the optimal weighting coefficient and AUC of the weighted combination models.
Then, for the above four types of features, Tumor + Clinical, Tumor + R3mm, Tumor + R5mm, R 3 mm, R 5 mm + Clinical models were constructed and optimized using the feature combination method based on a variety of classifiers, and the performance of which was verified in the validation set.
Radiomics model testing
In the test set, the weighted combination models and feature combination models were tested for predictive performance.
Statistical methods
Statistical analyses were performed using SPSS version 24.0 (IBM, Armonk, NY, USA) and R software version 4.0.2 (MathSoft, Seattle, Washington, USA). For non-normally distributed variables, the Mann Whitney U test was utilized to compare statistical differences between the two groups. For normally-distributed variables, t-tests or chi-square tests were conducted. SPSS version 24.0 was performed to draw the Receiver operating characteristic curve (ROC). When the optimization function (0.6 * sensitivity + 0.4 * specificity) was maximum, the cutoff value, sensitivity, and specificity of weighted combination models were taken, and while the Yoden Index was maximum, those of the single data source model and feature combination model was acquired. Delong test was performed to compare the differences in AUCs between two models. R software version 4.0.2 was conducted to draw the scatter plots of the alpha-AUC.
Results
Baseline characteristics of patients
The baseline characteristics of the patients were listed in Table 1, S1 Table, and S2 Table, respectively. There were significant differences (P < 0.05) in shape, aspect ratio, mean erythrocyte volume, and mean hemoglobin content between HER2-positive and HER2-negative groups in the training and validation sets. In the training set, there were significant differences (P < 0.05) in margins, halos, microcalcifications, leukocytes, and the absolute value of neutrophils between the two groups. In the validation set, there were significant differences (P < 0.05) between the two groups in tumor maximum diameter in the coronal plane, hemoglobin, erythrocyte pressure, basophil percentage, platelet ratio, platelet distribution width, and large platelet ratio. In the training and validation sets, 45 cases (45/139) and 11 cases (11/35) patients were HER2-positive breast cancers, respectively.
Comparison of the single data source models
The single-data source models were constructed respectively based on ISMS software. The ROCs of these models in the training set, validation set, and test set are shown in Fig. 3, Table 2. Thus, random forest classifier (RFC), light gradient booster (LGBM), gradient enhancement classifier (GBC), and Extra tree classifier (ETC) were the best classifiers to construct the Tumor model, R3mm model, R5mm model, and Clinical model, respectively. Overall, the AUCs of models decreased sequentially in the training, validation, and test sets. In the validation set, the Clinical model was the highest in terms of AUC 0.803 (95% confidence interval [CI] 0.660–0.947), with a sensitivity of 100% and specificity of 62.5%, followed by R5mm, R3mm, and Tumor models. In the test set, the Clinical model acquired the highest AUC of 0.695 (95% CI 0.583–0.807), the sensitivity of 86.1%, and specificity of 41.9%, followed by the Tumor, R3mm, and R5mm models.
(A) ROCs of the optimal Tumor, the optimal R3mm, the optimal R5mm, and the optimal Clinical model in the training set. (B) ROCs of the optimal Tumor, the optimal R3mm, the optimal R5mm, and the optimal Clinical model in the validation set. (C) ROCs of the optimal Tumor, the optimal R3mm, the optimal R5mm, and the optimal Clinical model in the test set. R3mm, model based on peritumoral 3 mm ring of breast tumor; R5mm, model based on peritumoral 5 mm ring of breast tumor; Tumor, model based on radiomics features of the tumor. ABC, Ada Boosting Classifier; AUC, area under the curve; CI, confidence interval; Clinical, model based on clinical, ABVS, and serology features of breast tumor; ETC, Extra Tree Classifier; GBC, Gradient Boosting Classifier; LGBM, Light Gradient Boosting Machine; RFC, Random Forest Classifier.
Comparison of the weighted combination models
In the validation set, the model weighted combination method was adopted. It could be seen from the alpha-AUC scatter plots that Tumor + Clinical, R3mm + Clinical, and R5mm + Clinical achieved higher AUC when alpha was 0.10, 0.15, and 0.05; Tumor + R3mm owned higher AUC when alpha was 0.05, 0.25, and 0.35; Tumor + R5mm acquired better AUC when alpha was 0.40, 0.45 and 0.50 (Fig. 4, Table 3). In the validation set, the R5mm + Clinical model acquired the highest AUC, 0.826 (95% CI 0.689–0.962), the sensitivity of 100%, and specificity of 62.5%, which was sequentially higher than the R3mm + Clinical model, Tumor + Clinical model, Tumor + R5mm model, and Tumor + R3mm model (Fig. 5, Table 3).
Scatter plots of alpha-AUC of weighted combination models in the validation set. (A) Scatter plots of alpha-AUC of the Tumor model combined with the Clinical model in the validation set. (B) Scatter plots of alpha-AUC of the Tumor model combined with the R5mm model in the validation set. (C) Scatter plots of alpha-AUC of the Tumor model combined with the R3mm model in the validation set. (D) Scatter plots of alpha-AUC of the R5mm model combined with the Clinical model in the validation set. (E) Scatter plots of alpha-AUC of the R3mm model combined with the Clinical model in the validation set. Notes: The two single data source models were combined through the formula of alpha*model 1 + (1-alpha) * model; Alpha, weighted coefficient. AUC, the area under the curve.
ROCs of weighted combination models based on different alphas in the validation set. (A) ROC of the R3mm model combined with the Clinical model in the validation set. (B) ROC of the R5mm model combined with the Clinical model in the validation set. (C) ROC of the Tumor model combined with the Clinical model in the validation set. (D) ROC of the Tumor model combined with R3mm model in the validation set. (E) ROC of the Tumor model combined with R5mm model in the validation set. Notes: Alpha, weighting coefficient. AUC, the area under the curve.
In the test set, the Tumor + Clinical model owned the highest AUC, 0.700 (95% CI 0.590–811), the sensitivity of 86.1%, and specificity of 41.9%, which was higher than those of the R3mm + Clinical model, R5mm + Clinical model, Tumor + R5mm model and Tumor + R3mm model (Fig. 6 and Table 3).
ROCs of weighted combination models based on different alphas in the test set. (A) ROC of the R3mm model combined with the Clinical model in the test set. (B) ROC of the R5mm model combined with the Clinical model in the test set. (C) ROC of the Tumor model combined with the Clinical model in the test set. (D) ROC of the Tumor model combined with R3mm model in the test set. (E) ROC of the Tumor model combined with R5mm model in the test set. Notes: Alpha, weighting coefficient. AUC, the area under the curve.
In the validation and test set, the AUCs of the weighted combination models T + Clinical, T + R5mm, and T + R3mm were better than those of the corresponding single data source model.
Comparison of the feature combination models
The AUC, sensitivity, and specificity of the feature combination models in the validation and test sets were shown in S3 Table. In the validation set, the Tumor + Clinical model had the highest AUC, 0.739 (95% CI 0.556–0.921), with a sensitivity of 81.8% and specificity of 66.7%. In the test set, the Tumor + R3mm model owned the best AUC, 0.668 (95% CI 0.555, 0.782), with a sensitivity of 61.1% and specificity of 71.0%.
Comparison among the single data source model, the feature combination models, and the weighted combination model
Overall, the AUCs of the weighted combination model were higher than most of the corresponding feature combination models and single data source models in both the validation set and the test set. In the validation set, the AUC of the optimal weighted combination model was superior to the optimal feature combination model (0.826 vs. 0.739, P = 0.038), and the optimal single data source model (0.826 vs.0.803, P = 0.446); in the test set, the AUC of the optimal weighted combination model was higher than the optimal feature combination model (0.700 vs. 0.668, P = 0.054), and the optimal single data source model (0.700 vs. 0.695, p = 0.501).
Features analysis of models
R5mm + Clinical and Tumor + Clinical were the optimal radiomics models in the validation and test sets, respectively. Important features for constructing R5mm, Tumor, and Clinical models were shown in S4 Table. For the Clinical model, microcalcifications and aspect ratios were important features in predicting HER2 status, and HER2-positive breast cancers were more likely to show intralesional microcalcifications and growth perpendicular to the skin, as shown in Fig. 7A–C. For the R5mm and Tumor models, Shape_Spherical Disproportion, Shape_Compactness 1, Shape_Compactness 2, and Shape_Elongation features representing tumor shape were key features in predicting HER2 status, and HER2-positive breast cancers tended to be more irregular shapes in the tumor, peritumoral areas (Fig. 7D–F), and ROIs (Fig. 7G–H).
ABVS images of HER2-positive breast tumor and ROIs of the breast tumor region and 5 mm peritumoral region. Notes: Internal microcalcifications in HER2-positive breast cancer in ABVS images (A coronal plane, B axial plane, C sagittal plane; Red arrow identified breast tumor); HER2-positive Breast Cancer Aspect Ratio > 1 in ABVS images (D coronal plane, E axial plane, F sagittal plane; Red arrow identified breast tumor); ROIs of HER2-positive breast tumor region (white area) and 5 mm peritumoral region (red area) were both irregular in shape (G); ROIs of HER2-negavive breast tumor region (white area) and 5 mm peritumoral region(red area) were both relative regular in shape (H).
Discussion
It is an indisputable fact that HER2 is a key therapeutic target for breast cancer. HER2 status is clinically crucial for delaying HER2-positive breast cancer progression, reducing the risk of recurrence19,20, improving treatment outcomes21, and survival19,21. We explored the efficacy of ABVS imaging in predicting HER2 status in breast cancer. Research related to the current study focused on the internal features of breast cancers, ignoring peritumoral information9,10. We concluded that peritumoral tissue could provide as much important information as the tumor itself. In the current research, R5mm + Clinical was the optimal weighted combination model in the validation set, and R3mm + Clinical was the weighted combination model second only to Tumor + Clinical in the test set. This was in part consistent with the previous view that peritumoral information has diagnostic and predictive value for molecular typing of breast cancer22,23,24.
The study confirmed that the model weighted combination method had more advantages than the feature combination method in optimizing the model, and we speculated that the model weighted combination method could preserve and optimize the vital features of the single data source model25. Some studies that predicted HER2 status in breast cancer focused only on image features and did not include relevant clinical data as predictors15, which might lead to the low performance of the model (AUC:0.650). In this study, clinical models, as the optimal single data source model, contributed significantly to the predictive performance of weighted combined models. The reason might be that the ABVS ultrasound features included in the Clinical model could largely reflect tumor heterogeneity26, which was also identical to the findings of Zheng et al27. ABVS, as a three-dimensional breast ultrasound, could provide more comprehensive breast tumor information than conventional ultrasound, which was presumed to be the reason why the AUC of the ABVS-based radiomics model in the current study was higher than that of the conventional ultrasound-based radiomics model (AUC: 0.826 vs. 0.740)16.
We compared the predictive performance of different classifiers when constructing models of intratumoral, peritumoral, and clinical features to select the optimal classifier, optimizing the predictive performance of the single data source model to some extent. Because of the important significance of HER2 positivity for breast cancer diagnosis and treatment, we optimized the cutoff value of ROC to ensure the high sensitivity of the model.
In the current study, we derived important features for constructing the Tumor, R5mm, and Clinical models. In terms of the Tumor model, leukocytes, basic granulocytes, microcalcifications, aspect ratio, and postoperative axillary lymph node metastasis status were crucial features that predict HER2 positivity. This coincided with previous studies demonstrating that basic granulocytes, monocytes, lymphocytes, and microcalcifications were predictors of HER2-positive breast cancer28,29,30. Aspect ratio ≥ 1 and axillary lymph node metastasis were the manifestations of invasive growth and metastasis of breast cancer, which owned a certain predictive value for HER2-positive breast cancer with higher invasiveness4. For R5mm and Clinical models, Shape_SphericalDisproportion, Shape_Compactness 1, Shape_Compactness 2, and Shape_Elongation features representing tumor shape could reflect tumor aggressiveness. In addition, GLSZM, GLCM, and GLRLM, as grayscale features, might reflect the heterogeneity and complexity of tumors31.
Given the performance of the single data source model was unsatisfactory in the validation set, we utilized the method of model weighted to construct a combined model. At the same time, to obtain the optimal weighting coefficient (alpha), we drew an alpha-AUC scatter plot to accurately and intuitively acquire the tendency of the AUC of the weighted combined model to change with alpha. The above method fully optimized the prediction performance of the weighted combination model.
In the present research, the performance of the weighted combination model was significantly superior to that of the feature combination model, but it was not much different from the single data source model. We will explore a more effective combination mode to optimize the combination model in the future.
Our study had the following strengths: Firstly, the multicenter test set was acquired to assess the clinical generalizability of the model. Secondly, combining ABVS radiomics features with relevant clinical and serological features greatly enhanced the predictive performance of the model. Thirdly, the optimal combination model was constructed by the model weighted combination method based on the idea of model ensemble. Forth, with 3D ultrasound data, we're able to provide more comprehensive tumor information for model construction.
The performance of the proposed model may be influenced by the clarity of the ABVS images, the precision of the ROIs, the accuracy of the ABVS ultrasound and the serological features. While the present study showed promising results for predicting HER2 status in breast cancer. However, it also had several limitations: First of all, further research on breast cancer molecular subtypes and assessment of response to neoadjuvant chemotherapy are still needed. In addition, the model should be applied to other ultrasound modalities such as contrast-enhanced ultrasound and ultrasound elasticity32,33. Furthermore, future studies should also expand the number of cases to include other types of breast cancer, verifying the generalizability of the modely34,35. Last but not least, how to apply the research results to clinical practice was the direction that we should strive for in the future.
In summary, the weighted combination model integrating ABVS imaging features, and clinical and serological features could better predict HER2 status in breast cancer patients than the feature combination model and had certain clinical generalizations. The current study provided a simple, non-invasive, and preoperative method for HER2 status prediction, guiding the individualized clinical decision-making for breast cancer patients.
Data availability
The datasets generated and/or analysed during the current study are not publicly available due to protect the privacy of the patients, but are available from the corresponding author on reasonable request.
References
Xia, C. et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin. Med. J. 135, 584–590. https://doi.org/10.1097/cm9.0000000000002108 (2022).
Tokunaga, E. et al. Trastuzumab and breast cancer: developments and current status. Int. J. Clin. Oncol. 11, 199–208. https://doi.org/10.1007/s10147-006-0575-4 (2006).
Gutierrez, C. & Schiff, R. HER2 biology, detection, and clinical implications. Arch. Pathol. Lab. Med. 135, 55–62 (2011).
Loibl, S. & Gianni, L. HER2-positive breast cancer. Lancet 389, 2415–2429. https://doi.org/10.1016/s0140-6736(16)32417-5 (2017).
Wang, H. Y. et al. Automated breast volume scanning: identifying 3-D coronal plane imaging features may help categorize complex cysts. Ultrasound Med. Biol. 42, 689–698. https://doi.org/10.1016/j.ultrasmedbio.2015.11.019 (2016).
Golatta, M. et al. Interobserver reliability of automated breast volume scanner (ABVS) interpretation and agreement of ABVS findings with hand held breast ultrasound (HHUS), mammography and pathology results. Eur. J. Radiol. 82, E332–E336. https://doi.org/10.1016/j.ejrad.2013.03.005 (2013).
Xu, C., Wei, S., Xie, Y., Guan, X. & Yang, B. Three-dimensional assessment of automated breast volume scanner compared with handheld ultrasound in pre-operative breast invasive ductal carcinomas: a pilot study of 51 cases. Ultrasound Med. Biol. 42, 2089–2096. https://doi.org/10.1016/j.ultrasmedbio.2016.05.007 (2016).
Acharya, U. R., Hagiwara, Y., Sudarshan, V. K., Chan, W. Y. & Ng, K. H. Towards precision medicine: from quantitative imaging to radiomics. J. Zhejiang Univ. Sci. B 19, 6–24. https://doi.org/10.1631/jzus.B1700260 (2018).
Song, L., Li, C. & Yin, J. Texture analysis using semiquantitative kinetic parameter maps from DCE-MRI: Preoperative prediction of HER2 status in breast cancer. Front. Oncol. 11, 25. https://doi.org/10.3389/fonc.2021.675160 (2021).
Fan, M. et al. Radiomic analysis reveals DCE-MRI features for prediction of molecular subtypes of breast cancer. PLoS ONE https://doi.org/10.1371/journal.pone.0171683 (2017).
Hill, B. S., Sarnella, A., D’Avino, G. & Zannetti, A. Recruitment of stromal cells into tumour microenvironment promote the metastatic spread of breast cancer. Semin. Cancer Biol. 60, 202–213. https://doi.org/10.1016/j.semcancer.2019.07.028 (2020).
D’Alessio, A., Proietti, G., Sica, G. & Scicchitano, B. M. Pathological and molecular features of glioblastoma and its peritumoral tissue. Cancers https://doi.org/10.3390/cancers11040469 (2019).
Wu, J. et al. Heterogeneous enhancement patterns of tumor-adjacent parenchyma at MR imaging are associated with dysregulated signaling pathways and poor survival in breast cancer. Radiology 285, 401–413. https://doi.org/10.1148/radiol.2017162823 (2017).
Kawashima, H., Inokuchi, M., Furukawa, H., Ikeda, H. & Kitamura, S. Magnetic resonance imaging features of breast cancer according to intrinsic subtypes: Correlations with neoadjuvant chemotherapy effects. Springerplus https://doi.org/10.1186/2193-1801-3-240 (2014).
Li, H. et al. Quantitative MRI radiomics in the prediction of molecular classifications of breast cancer subtypes in the TCGA/TCIA data set. NPJ Breast Cancer 2, 16012 (2016).
Wu, L. et al. Preoperative ultrasound radiomics analysis for expression of multiple molecular biomarkers in mass type of breast ductal carcinoma in situ. BMC Med. Imaging https://doi.org/10.1186/s12880-021-00610-7 (2021).
van Griethuysen, J. J. M. et al. Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77, E104–E107. https://doi.org/10.1158/0008-5472.can-17-0339 (2017).
Amirruddin, A. D., Muharam, F. M., Ismail, M. H., Tan, N. P. & Ismail, M. F. Synthetic Minority Over-sampling TEchnique (SMOTE) and Logistic Model Tree (LMT)-Adaptive Boosting algorithms for classifying imbalanced datasets of nutrient and chlorophyll sufficiency levels of oil palm (Elaeis guineensis) using spectroradiometers and unmanned aerial vehicles. Comput. Electron. Agric. https://doi.org/10.1016/j.compag.2021.106646 (2022).
Waks, A. G. & Winer, E. P. Breast cancer treatment: A review. JAMA 321, 288–300. https://doi.org/10.1001/jama.2018.19323 (2019).
Von Minckwitz, G. et al. Adjuvant pertuzumab and trastuzumab in early HER2-positive breast cancer. N. Engl. J. Med. 377, 702–702. https://doi.org/10.1056/NEJMx170011 (2017).
Rugo, H. S. & Chien, A. J. HER2-positive breast cancer: Is more treatment better?. Lancet Oncol. 17, 268–270. https://doi.org/10.1016/s1470-2045(15)00623-3 (2016).
Braman, N. et al. Association of peritumoral radiomics with tumor biology and pathologic response to preoperative targeted therapy for HER2 (ERBB2)-positive breast cancer. JAMA Netw. Open https://doi.org/10.1001/jamanetworkopen.2019.2561 (2019).
Penn, A. I. et al. Discrimination of benign from malignant breast lesions in dense breasts with model-based analysis of regions-of-interest using directional diffusion-weighted images. BMC Med. Imaging https://doi.org/10.1186/s12880-020-00458-3 (2020).
Zhou, J. et al. Diagnosis of benign and malignant breast lesions on DCE-MRI by using radiomics and deep learning with consideration of peritumor tissue. J. Magn. Reson. Imaging 51, 798–809. https://doi.org/10.1002/jmri.26981 (2020).
Chen, Q. & Li, J. in 11th International Conference on Intelligent Computing (ICIC). 354–362.
Wang, H., Yang, X., Ma, S., Zhu, K. & Guo, S. An optimized radiomics model based on automated breast volume scan images to identify breast lesions: comparison of machine learning methods. J. Ultrasound Med. https://doi.org/10.1002/jum.15845 (2021).
Zheng, X. et al. Deep learning radiomics can predict axillary lymph node status in early-stage breast cancer. Nat. Commun. https://doi.org/10.1038/s41467-020-15027-z (2020).
Barczak, W. et al. Telomere length assessment in leukocytes presents potential diagnostic value in patients with breast cancer. Oncol. Lett. 11, 2305–2309. https://doi.org/10.3892/ol.2016.4188 (2016).
Wiguna, I. G. W. W. et al. Tumor infiltrating lymphocyte and leukocyte ratios as metastatic predictor in HER2 positive luminal type breast cancer. Ann. Oncol. 32, S338–S338. https://doi.org/10.1016/j.annonc.2021.05.703 (2021).
Huang, Z., Chen, L., Wang, Y., Fu, L. & Lv, R. Molecular markers, pathology, and ultrasound features of invasive breast cancer. Clin. Imaging 79, 85–93. https://doi.org/10.1016/j.clinimag.2021.03.039 (2021).
Chen, Y. et al. Prediction of HER2 expression in breast cancer by combining PET/CT radiomic analysis and machine learning. Ann. Nucl. Med. 36, 172–182. https://doi.org/10.1007/s12149-021-01688-3 (2022).
Heydarheydari, S., Birgani, M. J. T. & Rezaeijo, S. M. Auto-segmentation of head and neck tumors in positron emission tomography images using non-local means and morphological frameworks. Pol. J. Radiol. 88, e364–e369. https://doi.org/10.5114/pjr.2023.130815 (2023).
Rezaeijo, S. M., Chegeni, N., Naeini, F. B., Makris, D. & Bakas, S. Within-modality synthesis and novel radiomic evaluation of brain MRI scans. Cancers https://doi.org/10.3390/cancers15143565 (2023).
Hosseinzadeh, M. et al. Prediction of cognitive decline in Parkinson’s disease using clinical and DAT SPECT imaging features, and hybrid machine learning systems. Diagnostics https://doi.org/10.3390/diagnostics13101691 (2023).
Salmanpour, M. R., Hosseinzadeh, M., Rezaeijo, S. M. & Rahmim, A. Fusion-based tensor radiomics using reproducible features: Application to survival prediction in head and neck cancer. Comput. Methods Programs Biomed. https://doi.org/10.1016/j.cmpb.2023.107714 (2023).
Acknowledgements
This work was supported by Gansu Province Science and Technology Program (Grant No. 23JRRA1589), Gansu Health Industry Scientific Research Project (Grant No. GSWSKY2022-54) and the intra-hospital Fund of the First Hospital of Lanzhou University (Grant No. ldyyyn2019-65).
Author information
Authors and Affiliations
Contributions
H.W., W.C., T.L., F.C., and R.L. collected the data. T.L. and F.C. made the formal analysis. H.W. provided the Funding. S.J., L.X., and H.W. were responsible for methodology. H.W. and S.G. conducted the project administration. S.J. provided the software. H.W., S.J., and W.C. wrote the article. All authors reviewed and edited the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, H., Chen, W., Jiang, S. et al. Intra- and peritumoral radiomics features based on multicenter automatic breast volume scanner for noninvasive and preoperative prediction of HER2 status in breast cancer: a model ensemble research. Sci Rep 14, 5020 (2024). https://doi.org/10.1038/s41598-024-55838-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-55838-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.