Abstract
Nonhost resistance of Arabidopsis thaliana against the hemibiotrophic fungus Colletotrichum tropicale requires PEN2-dependent preinvasive resistance and CYP71A12 and CYP71A13-dependent postinvasive resistance, which both rely on tryptophan (Trp) metabolism. We here revealed that CYP71A12, CYP71A13 and PAD3 are critical for Arabidopsis’ postinvasive basal resistance toward the necrotrophic Alternaria brassicicola. Consistent with this, gene expression and metabolite analyses suggested that the invasion by A. brassicicola triggered the CYP71A12-dependent production of indole-3-carboxylic acid derivatives and the PAD3 and CYP71A13-dependent production of camalexin. We next addressed the activation of the CYP71A12 and PAD3-dependent postinvasive resistance. We found that bak1-5 mutation significantly reduced postinvasive resistance against A. brassicicola, indicating that pattern recognition contributes to activation of this second defense-layer. However, the bak1-5 mutation had no detectable effects on the Trp-metabolism triggered by the fungal penetration. Together with this, further comparative gene expression analyses suggested that pathogen invasion in Arabidopsis activates (1) CYP71A12 and PAD3-related antifungal metabolism that is not hampered by bak1-5, and (2) a bak1-5 sensitive immune pathway that activates the expression of antimicrobial proteins.
Similar content being viewed by others
Introduction
The resistance of an entire plant species against all isolates of particular pathogens is called nonhost resistance1. Colletotrichum tropicale (hereafter Ctro), formally called C. gleosporioides, is a hemibiotrophic fungal pathogen that causes anthracnose on its host mulberry; however, it is not able to enter the nonhost Arabidopsis because Arabidopsis blocks pathogen entry via activation of preinvasive resistance. Preinvasive resistance of Arabidopsis against Ctro involves PENETRATION2 (PEN2) and PEN32,3,4,5,6,7,8. Nonhost preinvasive resistance toward Ctro also needs EDR1 (ENHANCED DISEASE RESISTANCE 1)9.
Importantly, even the pen2 edr1 mutant is still not fully susceptible to Ctro10, because strong postinvasive resistance is newly activated once Ctro enters epidermal cells of the mutants defective in the preinvasive resistance. We reported previously that Arabidopsis cyp79B2 cyp79B3 double mutant is fully susceptible to the nonadapted pathogen Ctro, i.e., the mutant is defective in both preinvasive and postinvasve resistance10. CYP79B2 and CYP79B3 are key enzymes for the biosynthesis of tryptophan (Trp)-derived antimicrobial metabolites. CYP79B2/CYP79B3 convert Trp into indole-3-acetaldoxime (IAOx)11, and this precursor is then converted into several compounds for antimicrobial immunity, such as PEN2 substrates indole-glucosinolates (IGs), PAD3 (PHYTOALEXIN-DEFICIENT3)-dependent camalexin, and 4-hydroxy-ICN (4-OH-ICN) whose biosynthesis requires CYP82C212,13,14. We have shown that the pen2 pad3 mutant is partially defective in postinvasive resistance to Ctro, indicating that the Arabidopsis phytoalexin, camalexin, is a critical factor for this.
In addition to serving as a precursor of IGs and camalexin, IAOx can be also converted to indole-3-carboxylic acid and its derivatives (ICAs). We have shown recently that CYP71A12 but not CYP71A13 has an important contribution to the accumulation of ICAs in leaves upon both Ctro and P. cucumerina inoculation15. On the other hand, loss of CYP71A13 reduced camalexin accumulation in leaves upon infection by multiple pathogens15,16, whereas a single loss of CYP71A12 did not reduce camalexin accumulation upon Ctro and P. cucumerina infection15. These findings suggest distinct roles of these two homologous P450 monooxygenases in the responses toward pathogen infection. Importantly, the pen2 cyp71A12 double mutant exhibits a partial reduction in postinvasive resistance to Ctro15, which was similar to that of the pen2 pad3 plants. This indicates that CYP71A12-dependent synthesis of ICAs as well as camalexin synthesis is critical for postinvasive resistance to Ctro, whereas CYP71A12 and PAD3 are dispensable for the preinvasive resistance: Ctro cannot invade the pad3 single mutant or the cyp71A12 single mutant15.
It has been reported that in addition to the above mentioned pests, camalexin is critical for the immunity of Arabidopsis to additional filamentous pathogens including a necrotrophic fungus Alternaria brassicicola (hereafter called Ab)17,18. However, it remains unclear whether CYP71A12 and ICAs are involved in Arabidopsis immunity to other fungi including Ab.
Immune responses in plants, including biosynthesis of specialized metabolites, can be triggered by pattern recognition-receptor (PRRs). BAK1 (BRASSINOSTEROID INSENSITIVE 1-ASSOCIATED RECEPTOR KINASE 1) is known to act as a coreceptor with multiple PRRs, including FLS2 and EFR, via ligand-induced heteromerization19,20,21,22. BAK1 was initially identified as a positive regulator of the brassinosteroid response (BR)23,24. Correspondingly, the bak1 null mutants such as bak1-4 mutant have a defect not only in FLS2 and EFR-dependent immune responses, but are also hyposensitive to BR. In contrast to the null alleles, the bak1–5 allele is impaired in pathogen-associated molecular pattern (PAMP)-triggered immunity, but not in BR signaling25. Importantly, bak1-5 is more severely impaired in defense responses than the null mutant bak1-425.
Here, we investigated whether CYP71A12-dependent synthesis of ICAs might be involved in Arabidopsis immunity, especially postinvasive resistance, to Ab that infects Brassicaceae plants. As a result, we found that the invasion by Ab triggers the CYP71A12-dependent accumulation of ICAs as well as camalexin. Furthermore, evaluation of lesion development and microscopic observation of the pathogen invasion suggested the involvement of CYP71A12, together with PAD3, in postinvasive resistance to Ab.
We also asked how Arabidopsis recognizes Ab invasion to activate the CYP71A12 and PAD3-dependent postinvasive resistance. To this end we investigated function of BAK1 in immunity and ICAs formation and found that the two bak1 mutations, especially bak1–5, reduce postinvasive resistance to Ab, revealing the involvement of a PRR system in the recognition of Ab invasion for the activation of defense. Unexpectedly, we found that the bak1–5 mutation has no negative impact on the invasion-triggered activation of biosynthesis of camalexin or ICAs derived from Trp.
Results
CYP71A12 contributes to the immunity of Arabidopsis against the necrotrophic pathogen Alternaria brassicicola independently of CYP71A13 and PAD3
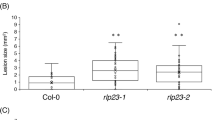
Ab is a necrotrophic fungal pathogen that infects several Brassicaceae spp., including cabbage and canola but is restricted within limited lesions when inoculated on leaves of Arabidopsis accession Col-09,17,26. To investigate the roles of CYP71A12 in the postinvasive immunity of Arabidopsis against Ab, we used the Ryo-1 strain of Ab for the inoculation assay (Fig. 1). During our former study on postinvasive resistance against Ctro we investigated Trp metabolism-related mutants including cyp71A12 in the pen2 background, because Ctro was not able to invade these mutants in the absence of the pen2 mutation (15). In this study, to focus on postinvasive resistance against Ab, we used these pen2 background mutants for the Ab inoculation assay to exclude a potential effect of PEN2 on preinvasive resistance against this pathogen. Our evaluation of lesion development at 4 days postinoculation (dpi) revealed that the pen2 cyp71A12 mutant showed enhanced susceptibility to Ab, as compared with the pen2 plants, indicating contribution of CYP71A12 to the immunity towards Ab (Fig. 1). We also found that lesion development in the pen2 mutant was not significantly different from that in the wild-type (WT) Col-0 leaves (Fig. 1), suggesting that opposite with the impact on several filamentous pathogens2,6, PEN2 has likely no detectable contribution to the Arabidopsis immunity against Ab at least in the Col-0 accession.
PAD3 and CYP71A12 are involved in the immunity of Arabidopsis against the necrotrophic pathogen Alternaria brassicicola (Ab). (A) Lesion development caused by Ab on Arabidopsis mutant plants with defects in Trp metabolism pathways. Conidial suspensions (1 × 105 conidia/mL) of Ab were drop-inoculated onto mature leaves of 4–5-week-old plants. The photograph was taken at 4 days postinoculation (dpi). (B) Quantification of lesion development. Conidial suspensions of Ab were drop-inoculated onto tested plants. At 4 dpi, lesion areas were measured and the relative values to Col-0 (WT plants) were calculated. Means and standard deviations (SDs) were calculated from three independent experiments. The statistical significance of differences between means was determined by Tukey’s honestly significant difference (HSD) test. Means not sharing the same letter are significantly different (P < 0.05). N.D., not determined.
We also found that the pen2 pad3 mutants have increased susceptibility to the Ab Ryo-1 (Fig. 1), indicating the importance of PAD3-dependent camalexin synthesis, consistent with previous reports17,26. Also, an additional mutation in CYP71A13 further increased lesion development in pen2 cyp71A12 (Fig. 1). Opposite with PAD3 and CYP71A12/A13, cyp82C2 mutation did not cause significant changes in Ab development suggesting that CYP82C2 together with 4-OH-ICN do not contribute to the immunity toward this pathogen (Fig. 1). We were unable to perform proper quantitative analysis of lesion development in the cyp79B2 cyp79B3 mutant at 4 dpi because the lesions had already merged. However, quantitative analysis at 3 dpi revealed that the cyp79B2 cyp79B3 mutant was the most susceptible to Ab among all tested genotypes (Supplementary Fig. S1). As PEN2 appeared to be unlikely essential for the immunity against Ab, we also investigated the effects of the mutations of PAD3, CYP71A12, CYP71A13 and CYP82C2 on the immunity to Ab in the WT background in addition to the pen2 background by using pad3, cyp82C2, cyp71A12, cyp71A12 cyp71A13 mutants. The obtained result was similar to the finding on the mutations in the pen2 background (Supplementary Fig. S2).
Biosynthesis of ICAs and camalexin occurs during postinvasive resistance toward Ab
Pastorczyk, M. et al.15 reported that (1) the expression of CYP71A12 and PAD3 is strongly induced in the pen2 mutant defective in preinvasive resistance, but not in WT, upon the inoculation of the non-adapted pathogen Ctro and (2) CYP71A12 and PAD3 are involved in postinvasive resistance against Ctro. To assess whether CYP71A12 and PAD3 are expressed during postinvasive resistance against Ab, we investigated the expression pattern of these genes at several time points after Ab inoculation (at 4, 12, 24, and 48 h postinoculation, hpi). Although lesion development by Ab in pen2 was comparable to that in WT (Fig. 1), we used the pen2 mutant for the analysis to exclude a potential contribution of PEN2 to preinvasive resistance to Ab.
We found that the expression of CYP71A12 and PAD3 started to be induced at 12 hpi in pen2, and the corresponding expression levels were stronger elevated at later time points (Fig. 2A). By contrast, we did not detect any induction at 4 hpi (Fig. 2A). In parallel, we also investigated the temporal infection behavior of Ab in Arabidopsis. We found that conidia of Ab had already germinated at 4 hpi, however, we did not detect any host invasion at this time (Fig. 2B). The fungus started to invade the pen2 plants at 12 hpi and the invasion ratio became elevated at later time points (Fig. 2B,D). These findings indicate a link between CYP71A12 and PAD3 induction and the initiation of host invasion in the Ab-Arabidopsis interactions, strongly suggesting that the expressions of CYP71A12 and PAD3 are triggered by Ab invasion. Furthermore, we also revealed that simultaneous loss of both CYP71A12 and CYP71A13 produced no detectable effects on the preinvasive resistance against Ab (Fig. 2C,D). These findings suggest that CYP71A12 and CYP71A13 are involved in the postinvasive resistance against Ab. We also found that the invasion ratio of Ab in pen2 was not significantly different from that in WT (Fig. 2C), in contrast to the case of Ctro6.
The invasion of Ab activates the expression of PAD3 and CYP71A12. (A) Expression of PAD3 and CYP71A12 following Ab inoculation. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto 4–5-week-old pen2 plants and kept at 100% humidity. The samples were collected at 4, 12, 24, and 48 hpi. Each gene transcript was quantified by quantitative polymerase chain reaction (qPCR) using the gene-specific primers listed in Supplementary Table S2. Values were normalized to the expression level of UBC21. The means and SDs were calculated from three independent experiments. Statistical comparisons between mock and Ab treated samples were conducted using two-tailed Student’s t tests (**P < 0.01). (B) Quantitative analysis of the Ab invasion ratio among pen2 plants. Conidial suspensions (1 × 105 conidia/mL) of Ab were drop-inoculated onto pen2 plants and kept at 100% humidity. The inoculated leaves were collected at 4, 12, 24, and 48 hpi, and then subjected to a trypan blue viability staining assay. The presence or absence of invasive hyphae from at least 50 germinating conidia were counted in each experiment. The means and SDs were calculated from three independent experiments. Statistical comparisons of the Ab invasion ratios at 12, 24, and 48 hpi against that at 4 hpi were conducted using two-tailed Student’s t tests (**P < 0.01). (C) PEN2, CYP71A12, and CYP71A13 are dispensable for preinvasive resistance against Ab. Aliquots of 5 μL of Ab conidial suspension were drop-inoculated onto leaves of 4–5-week-old plants. At 12 hpi, the inoculated leaves were collected and stained with trypan blue, and then invasive hyphae were observed by light microscopy. At least 50 germinating conidia were counted in each experiment. The means and SDs were calculated from three independent experiments. Statistical analysis using two-tailed Student’s t tests showed no significant differences among the genotypes. (D) Light microscopy observations. At 12 hpi, a part of the germinating conidia developed invasive hyphae (arrows) inside the inoculated plants. The top images are focused on conidium and the bottom images are focused on invasive hypha. c conidium, g germ tube. Bars = 50 µm.
We then assessed whether our observations made at the gene expression level could be supported with metabolite profiles. We found that the pathogen-induced accumulation of two ICAs, glucoside of 6-hydroxy-indole-3-carboxylic acid (6OGlcICA) and glucose ester of indole-3-carboxylic acid (ICAGlc), was detected at 24 hpi, but not at 4 hpi with Ab in both WT and pen2 (Fig. 3A; Supplementary Fig. S3), which matched strongly with the gene expression data (Fig. 2A). Similar results were also obtained during the analysis of camalexin accumulation (Fig. 3A). Therefore, we conclude that the synthesis of ICAs and camalexin is triggered by Ab invasion.
Invasion by Ab induced CYP71A12-dependent biosynthesis of 6-hydroxy-ICA and PAD3-dependent biosynthesis of camalexin. (A) The accumulation of 6-hydroxy-ICA (6OGlcICA) and camalexin in Arabidopsis plants inoculated with Ab. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto Col-0 (WT) and pen2 plants. As a control, water was sprayed as a mock treatment. The means of metabolites (nmol/g fresh weight, FW) and SDs from four biological independent samples are shown in the graph. The statistical significance of differences between means was determined by Tukey’s HSD test. Means not sharing the same letter are significantly different (P < 0.05). (B) CYP71A12 is required for the Ab-triggered accumulation of 6OGlcICA, whereas PAD3 and CYP71A13 are required for the accumulation of camalexin. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto tested mutant lines. The statistical significance of differences between means in each time point was determined by Tukey’s HSD test. Means not sharing the same letter are significantly different (P < 0.05).
Reduced accumulation of ICAs correlates with the breakdown of the postinvasive resistance toward Ab
We have reported recently that the accumulation of ICAs triggered by inoculation with P. cucumerina is reduced in the cyp71A12, but not in the cyp71A13 mutant15. By contrast, the P. cucumerina-triggered accumulation of camalexin was reduced in the cyp71A13, but not in the single cyp71A12 mutant15. Therefore, we assumed that the CYP71A12-dependent production of ICAs contributes to Arabidopsis postinvasive immunity to Ab independently of CYP71A13-dependent camalexin production. As mentioned above, we also found that the cyp79B2 cyp79B3 mutant exhibited more severe defects in postinvasive resistance against Ab than the pen2 cyp71A12 cyp71A13 mutant (Fig. 1 and Supplementary Fig. S1), although the reason was not clear. To get further insights on these aspects, we investigated the Trp-related metabolite profiles of the aforementioned mutants: pen2, pen2 pad3, pen2 cyp71A12, pen2 cyp71A12 cyp71A13, and cyp79B2 cyp79B3.
Obtained results revealed that the simultaneous loss of CYP79B2 and CYP79B3 completely abolished the Ab-triggered accumulation of ICAs and camalexin, whereas the loss of PAD3 canceled camalexin accumulation, but had no effects on the accumulation of ICAs (Fig. 3B; Supplementary Fig. S4). In the pen2 cyp71A12 mutant, the Ab-triggered accumulation of ICAs was reduced significantly compared with the pen2 mutant, opposite with camalexin accumulation that was rather increased compared with pen2 (Fig. 3B; Supplementary Fig. S4). These results further strengthen the idea that the accumulation of ICAs triggered by the Ab invasion is critical for the postinvasive resistance of Arabidopsis against this necrotrophic fungal pathogen. In the pen2 cyp71A12 cyp71A13 mutant, the ICA levels that accumulated upon Ab invasion were similar to those observed in the pen2 cyp71A12 mutant, but camalexin accumulation in the triple mutant was completely diminished, in contrast to pen2 cyp71A12 (Fig. 3B; Supplementary Fig. S4), further supporting the importance of CYP71A13 for the pathogen-induced accumulation of camalexin, but not ICAs.
It is noteworthy that the leaves of pen2 cyp71A12 cyp71A13 mutant still accumulated clearly detectable amounts of ICAs at both 24 hpi and 48 hpi of Ab, whereas camalexin in this triple mutant was under the detection limit at the same time points (Fig. 3B; Supplementary Fig. S4). As mentioned above, the cyp79B2 cyp79B3 mutant exhibited a more severe phenotype to Ab inoculation compared with the pen2 cyp71A12 cyp71A13 mutant (Fig. 1A,B; Supplementary Fig. S1). The cyp79B2 cyp79B3 mutant is entirely defective in the production of not only camalexin, but also ICAs (Fig. 3B; Supplementary Fig. S4). Also, both cyp79B2 cyp79B3 and pen2 cyp71A12 cyp71A13 mutants are commonly defective in biosynthesis of the PEN2-dependent IG-hydrolysis products (leading to unidentified antifungal compounds), these data suggested that the different levels of susceptibility to Ab between pen2 cyp71A12 cyp71A13 and cyp79B2 cyp79B3 is likely caused by the differential accumulation of ICAs in these two mutants.
In addition to ICAs and camalexin, CYP79B2/CYP89B3 enzymes are essential for biosynthesis of other Trp-derived metabolites including IGs15,27. The pen2 cyp71A12 cyp71A13 mutant lacks the PEN2-dependent IG-hydrolysis products, but retains the ability to produce IGs, which can be activated by another enzyme. It has been reported that IG biosynthesis in Arabidopsis is controlled by the transcription factors MYB34, MYB51, and MYB122, whereas these factors are dispensable for the pathogen triggered biosynthesis of camalexin and ICAs27. Thus, to assess the possibility that the different susceptibility to Ab between pen2 cyp71A12 cyp71A13 and cyp79B2 cyp79B3 mutants might be linked to IG biosynthesis, we compared phenotypes of pen2 cyp71A12 cyp71A13 and cyp71A12 cyp71A13 myb34 myb51 myb122 lines following Ab inoculation. We first performed the metabolite analyses on the cyp71A12 cyp71A13 myb34 myb51 myb122 mutant leaves at 24 hpi of Ab (Supplementary Fig. S5). The obtained results showed that this mutant was defective in IG biosynthesis upon Ab infection. Subsequently, we tested susceptibility of this mutant towards Ab in an inoculation assay. We found that lesion development in the cyp71A12 cyp71A13 myb34 myb51 myb122 mutant leaves was similar to that in the pen2 cyp71A12 cyp71A13 mutant (Supplementary Fig. S6). These results indicate that PEN2-independent IG hydrolysis is likely not involved in the postinvasive resistance against Ab. Thus, the different susceptibility between the pen2 cyp71A12 cyp71A13 mutant and the cyp79B2 cyp79B3 mutant is not linked to IG-deficiency. Together with the involvement of CYP71A12-dependent ICAs synthesis in the postinvasive resistance of Arabidopsis against Ab, we consider that the remaining ICAs in pen2 cyp71A12 cyp71A13 are still effective against Ab. Consistent with this idea, the pen2 cyp71A12 mutant was more susceptible to Ab than the pen2 mutant (Fig. 1).
The bak1–5 mutation reduces the postinvasive resistance of Arabidopsis to Ab, independently of pathogen-triggered ICAs and camalexin biosynthesis
Next, we asked how Arabidopsis recognizes the invasion by Ab to activate the biosynthesis of ICAs and camalexin. The candidate mechanism critical for this process is the PRRs-dependent PAMP recognition machinery28,29. When Arabidopsis recognizes PAMPs, at least some of the cognate PRRs, including FLS2 and EFR receptor-like kinases (RLKs), form complexes with coreceptors such as BAK119,20,21,22. Therefore, we decided to assess the possible involvement of BAK1 in the invasion-triggered accumulation of ICAs and camalexin. For this purpose, we used two mutant alleles of BAK1, bak1–4 and bak1–5; of these, bak1–4 is a BAK1 null allele19, and bak1–5 is a semi-dominant allele with a specific phenotype related to PAMP responsiveness25. Importantly, it was reported that bak1-5 is more severely impaired in elf18 and flg22 responses than the null mutant bak1-425.
Because the main results on gene expression and metabolite accumulation were derived from the pen2-background mutants in this study, we used the pen2 bak1–4 mutant30 and the newly generated pen2 bak1–5 mutant. We first performed Ab inoculation assay on these plant lines and found that both pen2 bak1–4 and pen2 bak1–5 plants have reduced immunity to Ab, although the pen2 bak1–5 plants were more susceptible than the pen2 bak1–4 plants (Fig. 4A,C). Similar results were observed for single bak1–4 and bak1–5 as compared with WT plants (Supplementary Fig. S7). We then evaluated whether the bak1–5 mutation would reduce preinvasive resistance to Ab. To assess this, we compared the invasion behavior of Ab in the pen2 mutant with that in the pen2 bak1–5 mutant. The invasion ratio in the pen2 bak1–5 mutant was similar to that in the pen2 mutant, suggesting that the bak1–5 mutation does not have detectable impact on preinvasive resistance to Ab (Fig. 4B). These results indicate that the bak1–5 mutation reduces postinvasive resistance to Ab, i.e., PRR systems likely function in the recognition of Ab invasion.
The bak1 mutations reduced the immunity of Arabidopsis against Ab. (A) Quantification of lesion development. Conidial suspensions of Ab (1 × 105 conidia/mL) were drop-inoculated onto true leaves of 4–5-week-old plants. At 4 dpi, lesion areas were measured and the relative values to Col-0 (WT plants) were calculated. The means and SDs were calculated from three independent experiments. The statistical significance of differences between means was determined by Tukey’s HSD test. Means not sharing the same letter are significantly different (P < 0.05). (B) The bak1–5 mutation did not reduce preinvasive resistance against Ab in the pen2 mutant. Aliquots of 5 μL of conidial suspension (1 × 105 conidia/mL) of Ab were drop-inoculated onto leaves of 4–5-week-old plants of the tested mutants. At 12 hpi, the inoculated leaves were collected and stained with trypan blue, and then invasive hyphae were observed under light microscopy. At least 50 germinating conidia were counted in each experiment. The means and SDs were calculated from three independent experiments. Statistical analysis using two-tailed Student’s t tests showed no significant differences between pen2 and pen2 bak1–5 mutants. (C) Lesion development caused by Ab on two pen2 bak1 mutants. Conidial suspensions (1 × 105 conidia/mL) of Ab were drop-inoculated onto mature leaves of 4–5-week-old plants. The photograph was taken at 4 dpi.
Because we found that the bak1–5 mutation reduced postinvasive resistance to Ab, we next investigated whether bak1–5 would have negative effects on the Ab invasion-triggered activation of camalexin and ICAs biosynthesis. We checked the gene expression of PAD3 and CYP71A12 in the pen2 and pen2 bak1–5 mutants after Ab inoculation. Surprisingly, the bak1–5 mutation did not cancel the induced expression of PAD3 and CYP71A12 in pen2 (Figs. 2A, 5A). Furthermore, we found that the accumulation of camalexin and ICAs upon Ab invasion was not reduced in the pen2 bak1–5 mutant compared with the pen2 mutant (Fig. 5B and Supplementary Fig. S8). The accumulation level of ICAs was even higher in pen2 bak1–5 than pen2 at 48 hpi, which might be due to enhanced infection in pen2 bak1–5 (Fig. 5B and Supplementary Fig. S8). Collectively, these results indicate that the bak1–5 mutation does not reduce the Ab-triggered activation of camalexin and ICAs biosynthesis, although this mutation reduces postinvasive resistance to Ab.
The bak1–5 mutation did not reduce the Ab-invasion triggered accumulation of 6-hydroxy-ICA and camalexin. (A) The Ab-invasion triggered activation of CYP71A12 and PAD3 expression in the pen2 bak1–5 mutant. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto 4–5-week-old plants. The samples were collected at 4, 12, 24, and 48 hpi. Each gene transcript was quantified by RT–qPCR using the gene-specific primers listed in Supplementary Table S2. Values were normalized to the expression level of UBC21. The statistical comparison between Ab-treated pen2 and Ab-treated pen2 bak1-5 at same time point samples was conducted using two-tailed Student’s t tests and did not show significant differences. (B) The Ab invasion-triggered accumulations of 6-hydroxy-ICA (6OGlcICA) and camalexin were not canceled by the bak1–5 mutation. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto the tested mutant lines. The means of metabolites (nmol/g FW) and SDs from four biological independent samples are shown in the graph. The statistical comparison between pen2 and pen2 bak1-5 at same timepoint was conducted using two-tailed Student’s t tests (**P < 0.01).
The bak1–5 mutation reduces the Ab invasion-triggered expression of defense-related genes, including GLIP1.
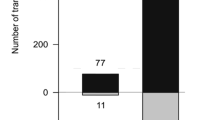
We further investigated the bak1–5 sensitive pathways for postinvasive resistance against Ab. We performed comparative expression profiling experiments in pen2 and pen2 bak1–5 plants following Ab invasion using microarray analysis. Ab was inoculated to each plant, and RNA isolated from inoculated leaves at 24 hpi was subjected to microarray analysis. We focused on differentially expressed genes associated with the immune response based on Gene Ontology (GO) data (GO term 0006955: http://www.informatics.jax.org). As a result, we found that 14 genes had greater than a 2.5-fold change in expression. Interestingly, 11 were down-regulated, and three were up-regulated (Supplementary Table S1), implying that, compared with the pen2 plants, the pen2 bak1–5 plants exhibited a trend for reduced immune responses upon Ab invasion. Among the 11 down-regulated genes, four were shown previously to be involved in the Arabidopsis immune system using functional analyses such as the analysis of corresponding knockout mutants, including AED1 (APOPLASTIC, EDS1-DEPENDENT 1)31, BGL2/PR232, GLIP133,34, and RLP23 (RECEPTOR-LIKE PROTEIN 23)35. Notably, the glip1 plants were reported to be more susceptible to Ab than the WT plant33. Subsequently, we performed reverse transcription quantitative polymerase chain reaction (RT–qPCR) analyses to investigate the expression levels of these four genes (AED1, BGL2/PR2, GLIP1, and RLP23) at 4, 12, 24, and 48 hpi in pen2 and pen2 bak1–5 plants inoculated with Ab (Fig. 6). As a control, we also investigated gene expression in the mock-treated plants. We confirmed that these four genes exhibited lower expression in the pen2 bak1–5 than in the pen2 mutant at 24 hpi, consistent with the array data. Furthermore, the RT–qPCR analysis revealed that the expression of AED1, BGL2/PR2, GLIP1, and RLP23 was lower at 4 hpi than at 12, 24, and 48 hpi (Fig. 6). Together with our finding that Ab did not invade at 4 hpi, but started to invade at 12 hpi (Fig. 2B), our results indicate that these genes are induced upon Ab invasion to function in postinvasive resistance. Such induced expression was continuously suppressed in the pen2 bak1–5 plants (Fig. 6), further suggesting that this invasion-triggered expression depends on putative PRRs with function impaired in the bak1–5 mutant. It is also noteworthy that the invasion-triggered expressions of AED1, BGL2/PR2, GLIP1, and RLP23 were time-dependent, i.e., induced expression started to be down-regulated at 12 hpi (RLP23) or 24 hpi (AED1, BGL2/PR2, GLIP1) (Fig. 6), which is in contrast to the invasion-triggered expressions of PAD3 and CYP71A12, which exhibited sustained elevations for up to 48 hpi (Figs. 2A, 5A).
The Ab invasion-triggered expression levels of AED1, BGL2/PR2, GLIP1, and RLP23 were reduced by the bak1–5 mutation. Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto 4–5-week-old pen2 and pen2 bak1–5 plants, and then kept at 100% humidity. The samples were collected at 4, 12, 24, and 48 hpi. Each gene transcript was quantified by RT–qPCR using the gene-specific primers listed in Supplementary Table S2. Values were normalized to the expression level of UBC21. The means and SDs were calculated from three independent experiments. The statistical analysis was conducted by a two-tailed t test. The expression levels of each gene between pen2 and pen2 bak1–5 were compared at the same time points and treatment (*P < 0.05; **P < 0.01).
Discussion
We recently reported that CYP71A12 is indispensable for postinvasive resistance to the hemibiotrophic pathogen Ctro that is not adapted to Arabidopsis15. However, it remained obscure whether CYP71A12 is also involved in this invasion-triggered resistance of Arabidopsis against other fungal pathogens.
Here we found that the absence of functional CYP71A12 enhanced lesion development during colonization of Arabidopsis leaves with the necrotrophic pathogen Ab, indicating that this enzyme is required for the immune response of Arabidopsis against this fungus. CYP71A12 was not induced at 4 hpi with Ab when the pathogen had not yet invaded, but it started to be induced upon Ab invasion (Fig. 2A). We also found that CYP71A12 was dispensable for preinvasive resistance against Ab (Fig. 2C). Collectively, these results demonstrate that CYP71A12 is required for postinvasive resistance against Ab as well as Ctro. Metabolic analyses showed that the enhanced accumulation of ICAs triggered by Ab invasion was reduced in the absence of functional CYP71A12. Thus, this enzyme is involved in the accumulation of ICAs following Ab invasion (Fig. 3B). Therefore, we consider that the CYP71A12-dependent synthesis of ICA and/or its derivatives upon Ab invasion is critical for postinvasive resistance to this fungus.
We also showed that CYP71A13 contributes to the postinvasive resistance against Ab via synthesis of camalexin, but not of ICAs (Fig. 3B). The result suggests the importance of camalexin for this second layer of defense, which is further supported by our phenotypic analyses of mutants defective in PAD3 (Fig. 1 and Supplementary Fig. S2).
PEN2 is involved in preinvasive resistance against Ctro6, but we here revealed that it is dispensable for preinvasive resistance against Ab (Fig. 2C,D). Furthermore, PEN1 is known to be required for preinvasive resistance against nonadapted powdery mildews36, but dispensable for the Colletotrichum fungi and nonadapted Alternaria alternata8,37. It is also noteworthy that PEN2 is involved in preinvasive resistance against A. alternata, in contrast with Ab37. Thus, the molecular components that underlie preinvasive resistance vary between fungal pathogens, probably because pathogens have evolved various strategies for plant entry. By contrast, once these pathogens enter Arabidopsis, they commonly develop invasive structures inside the plants; thus, the hosts might deploy an universal defense systems to terminate further fungal growth.
We found that the cyp79B2 cyp79B3 mutant is more susceptible to Ab than is the pen2 cyp71A12 cyp71A13 mutant (Fig. 1 and Supplementary Fig. S1). Interestingly, this phenomenon is also observed in the infection by Ctro and P. cucumerina15. Metabolite analyses of plants upon Ab invasion revealed that camalexin accumulation in the pen2 cyp71A12 cyp71A13 plants was almost the same as that in the cyp79B2 cyp79B3 plants (Fig. 3B). Importantly, the pen2 cyp71A1 cyp71A13 plants reduced their accumulation of ICAs, but still produced them to some degree, whereas the cyp79B2 cyp79B3 plants were entirely defective in this regard (Fig. 3B, Supplementary Fig. S4). Consistent with this finding, several independent metabolic branches are supposed to contribute to endogenous ICAs levels, including contribution of Arabidopsis aldehyde oxidase 1 and CYP71B615,38,39.
We also compared the phenotype of pen2 cyp71A12 cyp71A13 mutants with that of myb34 myb51 myb122 cyp71A12 cyp71A13 following Ab invasion and found no detectable differences in either mutants in terms of postinvasive resistance against Ab, indicating that the difference between pen2 cyp71A12 cyp71A13 and cyp79B2 cyp79B3 mutants is not caused by PEN2-unrelated IG metabolism products (Supplementary Figs. S5, S6). Collectively, these findings suggest that the lower susceptibility in the former mutants is caused by residual ICAs, i.e., ICAs contribute to postinvasive resistance in a dose-dependent manner. This supports the idea that ICAs or their derivatives work as antifungal compounds as opposed to functioning as signaling molecules for plant immune responses. However, we cannot exclude a possibility that accumulation of so far not-reported IAOx-derivatives whose biosynthesis is not dependent on CYP71A12 and CYP71A13 contribute to the difference observed in susceptibility of pen2 cyp71A12 cyp71A13 and cyp79B2 cyp79B3 plants.
We also investigated how Arabidopsis recognizes the invasion of fungal pathogens and then mounts its postinvasive resistance. We found that the two bak1 mutations, especially the bak1–5 mutation, reduce postinvasive resistance against Ab (Fig. 4). The finding that the negative effects of bak1-5 on the postinvasive resistance towards Ab was higher than that of bak1-4 is likely consistent with the previous works25.
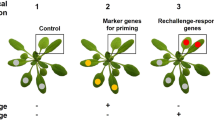
Surprisingly, we found that the bak1–5 mutation did not hamper the invasion-triggered expressions of PAD3 and CYP71A12 and the subsequent accumulation of ICAs and camalexin (Fig. 5). Thus, we postulate existence of another defense mechanism that is affected by the bak1–5 mutation and required for postinvasive resistance against Ab (Fig. 7). Our further analyses revealed that this pathogen activates the expression of distinct defense-related genes, including AED1, BGL2/PR2, GLIP1, and RLP23 (Supplementary Table S1 and Fig. 6). Notably, expressions of these genes were induced following Ab invasion, and these were canceled in bak1–5 mutants (Fig. 6). Therefore, we suggest that Arabidopsis deploys a PRR system to sense the invasion of Ab and subsequently activate antifungal defense pathways that are uncoupled from the Trp-metabolism (Fig. 7).
Summarized model for Arabidopsis postinvasive resistance against Ab. The CYP71A12-dependent production of ICAs is required for postinvasive resistance against A. brassicicola. The CYP71A13- and PAD3-dependent production of camalexin is also required for postinvasive resistance against Ab. The bak1 mutations (especially bak1-5) reduced the postinvasive resistance, however, invasion-triggered activation of these Trp-related pathways is not canceled by bak1-5. The bak1-5 sensitive pathway control the expression of antifungal protein genes (e.g. GLIP1).
Notably, it has been reported that the glip1 mutant plants exhibit enhanced susceptibility to Ab33. The recombinant GLIP1 protein exhibits antimicrobial activity that disrupts the Ab spores and hyphae33 and triggers systemic acquired resistance against bacterial pathogens (e.g., Erwinia carotovora and Pseudomonas syringae) as well as Ab34. Thus, the enhanced susceptibility of Arabidopsis to Ab in the presence of bak1–5 might be partially caused by the reduced expression of GLIP1.
It remains unclear how Arabidopsis plant recognizes pathogen invasion and then activates Trp-related metabolite accumulation as key immune responses in postinvasive resistance. Because PAD3 and CYP71A12 were commonly induced by the invasion of diverse fungal pathogens such as Ab and Ctro, we suggest that Arabidopsis probably recognizes the cell damage that is commonly caused by pathogen invasion and then activates Trp-related metabolism. Further studies are needed to explore a recognition mechanism of pathogen invasion that activates these secondary metabolic pathways for antifungal defense.
Materials and methods
Fungal materials
C. tropicale (Ctro) (formerly Colletotrichum gleosporioides S9275) was provided by Shigenobu Yoshida (National Institute for Agro-Environmental Sciences, Japan); and A. brassicicola (Ab) strain Ryo-1 was provided by Akira Tohyama. Cultures of Ab were maintained on 3.9% (w/v) potato dextrose agar medium (PDA; Nissui Pharmaceutical Co., Ltd., Tokyo, Japan) at 24 °C in the dark. Ctro was cultured on 2.5% (w/v) PDA (Difco, Detroit, MI, USA) at 24 °C under a cycle of 16 h black light (FS20S/BLB 20 W; Toshiba, Tokyo, Japan) illumination and 8 h dark.
Arabidopsis lines and growth conditions
The A. thaliana accession Col-0 was used as the WT plant. The mutants pen2-1, pen2-22, pad3-140, cyp71A12, cyp71A12 cyp71A1341, cyp82C214, bak1–419, bak1–525, cyp79B2 cyp79B311, pen2 pad310, pen2 cyp82C215, pen2 cyp71A1215, pen2 cyp71A12 cyp71A1315, cyp71A12 cyp71A13 myb34 myb51 myb12215, pen2 bak1–430 and pen2 bak1–5 (generated in this study) were used in this study. Arabidopsis seeds were sown on rockwool (Grodan; http://www.grodan.com) and kept at 4 °C in the dark for 2 days, and later grown at 25 °C with a cycle of 16 h light and 8 h dark in Hoagland medium.
Pathogen inoculation, lesion development analysis and trypan blue viability staining assay
For spray inoculation assays of Ab, 5 × 105 conidia/mL of conidial suspension was spray-inoculated on 4–5-week-old plants. For drop-inoculation, 5 μL of conidial suspensions of Ab (1 × 105 conidia/mL) were placed onto each leaf. The inoculated plants were kept at 25 °C with a cycle of 16 h light and 8 h dark and maintained at 100% relative humidity. For analysis of lesion development following the inoculation of Ab, four drops of 5 μL conidial suspension of each pathogen were drop-inoculated on each leaf, and 24–50 lesions were evaluated in each experiment. The developed lesions were quantified using ImageJ image analysis software (http://imagej.net) and relative values to WT (Col-0) plants were calculated. To measure lesion areas, yellowish areas were included as lesions. Trypan blue staining was conducted according to the method previously described42. For the trypan blue assay, at least 50 lesions were investigated in each experiment. The Ab invasion ratio (%) was calculated by using the following formula: Ab invasion ratio (%) = (number of germinating conidia that developed invasive hyphae / number of germinating conidia) × 100.
Generation of mutant plants
The generation of pen2 bak1–5 line used in this study was generated by crossing the pen2–1 mutant with bak1–5 plants. The genotype was checked with the corresponding specific primers for the derived cleaved-amplified polymorphic sequence (dCAPS) markers using dCAPS Finder 2.0 (http://helix.wustl.edu/dcaps/dcaps.html), and the PCR products (WT or mutant types) were cleaved with appropriate restriction enzymes (Supplementary Table S2).
RT–qPCR analysis
Seven Arabidopsis leaves inoculated with Ab (5 × 105 conidia/mL) were collected from each of seven different plants of either WT Col-0 or mutant plants at corresponding time points. Total RNA was extracted using PureLink (TRIzol plus RNA purification kits, Life Technologies/Thermo Fisher Scientific, Waltham, MA, USA) and treated with DNase (RQ1 RNase-free DNase; Promega, Madison, WI, USA; http://www.promega.com) to remove DNA contamination. Takara Prime Script RT kits (Takara Bio Inc., Shiga, Japan; http://www.takara-bio.com) was used for the cDNA synthesis. Takara TB Green Premix Ex Taq I was used for RT–qPCR, performed using the primers listed in Supplementary Table S2. Arabidopsis UBC21 (At5g25760) was used as an internal control for normalizing the level of cDNA43. RT–qPCR analysis was performed using a Thermal Cycler Dice Real Time System TP800 (Takara). The expression levels of genes of interest were normalized relative to those of UBC21.
Metabolite analysis
Conidial suspensions (5 × 105 conidia/mL) of Ab were spray-inoculated onto 4–5-week-old plants and kept at 100% relative humidity. Leaf samples (100–200 mg fresh weight) were collected at corresponding time points and frozen immediately in liquid nitrogen. The plant extracts containing Trp derivatives were extracted using DMSO and metabolite analyses were performed as described4,15.
Microarray analysis
Ab conidial suspensions (5 × 105 conidia/mL) were spray-inoculated onto 4–5-week-old plants of the pen2 and pen2 bak1–5 mutants. For each sample, five leaves were collected at 24 hpi and frozen immediately in liquid nitrogen. In total, eight samples (four biological replicates each of the mock and Ab-treated samples) were used for RNA extraction. Total RNA was extracted using Plant RNA Isolation Mini kits (Agilent Technologies., Santa Clara, CA, USA). Aliquots of 200 ng of total RNA were used to prepare Cy3-labeled cRNA using Agilent Low Input Quick Amp labeling kits. The labeled samples were hybridized onto an Agilent Arabidopsis thaliana microarray (ver. 4.0; 4 × 44 K format). After hybridization and washing, the arrays were scanned using an Agilent microarray scanner (G2565BA). The images were analyzed using Agilent Feature Extraction software (ver. 10.7.3.1), and further analysis was performed using Agilent GeneSpring GX12.1 software. Signal normalization was based on the expression ratio of pen2 bak1–5 to pen2. Differentially upregulated genes were defined as having a greater than 2.5-fold increase in expression, and differentially downregulated genes were defined as having at least a 0.4-fold decrease in expression. Microarray data have been deposited in the NCBI Gene Expression Omnibus (GEO) database GSE 124921.
References
Heath, M. C. Nonhost resistance C. Nonhost resistance and nonspecific plant defenses. Curr. Opin. Plant. Biol. 3, 315–319 (2000).
Lipka, V. et al. Pre- and postinvasion defenses both contribute to nonhost resistance in Arabidopsis. Science 310, 1180–1183 (2005).
Stein, M. et al. Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell 18, 731–746 (2006).
Bednarek, P. et al. A glucosinolate metabolism pathway in living plant cells mediates broad-spectrum antifungal defense. Science 323, 101–106 (2009).
Clay, N. K., Adio, A. M., Denoux, C., Jander, G. & Ausubel, F. M. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323, 95–101 (2009).
Hiruma, K. et al. Entry mode-dependent function of an indole glucosinolate pathway in Arabidopsis for nonhost resistance against anthracnose pathogens. Plant Cell 22, 2429–2443 (2010).
Kosaka, A. & Takano, Y. Nonhost resistance of Arabidopsis thaliana against Colletotrichum species. J. Gen. Plant Pathol. 84, 305–311 (2018).
Shimada, C. et al. Nonhost resistance in Arabidopsis-Colletotrichum interactions acts at the cell periphery and requires actin filament function. Mol. Plant Microbe Interact. 19, 270–279 (2006).
Hiruma, K. et al. Arabidopsis ENHANCED DISEASE RESISTANCE 1 is required for pathogen-induced expression of plant defensins in nonhost resistance, and acts through interference of MYC2-mediated repressor function. Plant J. 67, 980–992 (2011).
Hiruma, K. et al. Glutathione and tryptophan metabolism are required for Arabidopsis immunity during the hypersensitive response to hemibiotrophs. Proc. Natl. Acad. Sci. USA 110, 9589–9594 (2013).
Zhao, Y. et al. Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes Dev. 16, 3100–3112 (2002).
Bednarek, P. Sulfur-containing secondary metabolites from Arabidopsis thaliana and other Brassicaceae with function in plant immunity. Chem. Bio Chem. 13, 1846–1859 (2012).
Böttcher, C. et al. The multifunctional enzyme CYP71B15 (PHYTOALEXIN DEFICIENT3) converts cysteine-indole-3-acetonitrile to camalexin in the indole-3-acetonitrile metabolic network of Arabidopsis thaliana. Plant Cell 21, 1830–1845 (2009).
Rajniak, J., Barco, B., Clay, N. K. & Sattely, E. S. A new cyanogenic metabolite in Arabidopsis required for inducible pathogen defence. Nature 525, 376–379 (2015).
Pastorczyk, M. et al. The role of CYP71A12 monooxygenase in pathogen-triggered tryptophan metabolism and Arabidopsis immunity. New Phytol. 225, 400–412 (2020).
Nafisi, M. et al. Arabidopsis cytochrome P450 monooxygenase 71A13 catalyzes the conversion of indole-3-acetaldoxime in camalexin synthesis. Plant Cell 19, 2039–2052 (2007).
Thomma, B. P. H. J., Nelissen, I., Eggermont, K. & Broekaert, W. F. Deficiency in phytoalexin production causes enhanced susceptibility of Arabidopsis thaliana to the fungus Alternaria brassicicola. Plant J. 19, 163–171 (1999).
Sellam, A., Lacomi-Vasilescu, B., Hudhomme, P. & Simoneau, P. In vitro antifungal activity of brassinin, camalexin and two isothiocyanates against the crucifer pathogens Alternaria brassicicola and Alternaria brassicae. Plant Pathol. 56, 296–330 (2007).
Chinchilla, D. et al. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448, 497–500 (2007).
Heese, A. et al. The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl. Acad. Sci. USA 104, 12217–12222 (2007).
Roux, M. et al. The Arabidopsis leucine-rich repeat receptor-like kinases BAK1/SERK3 and BKK1/SERK4 are required for innate immunity to hemibiotrophic and biotrophic pathogens. Plant Cell 23, 2440–2455 (2011).
Ma, X., Xu, G., He, P. & Shan, L. SERKing coreceptors for receptors. Trends Plant Sci. 21, 1017–1033 (2016).
Li, J. et al. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 110, 213–222 (2002).
Nam, K. H. & Li, J. BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110, 203–212 (2002).
Schwessinger, B. et al. Phosphorylation-dependent differential regulation of plant growth, cell death, and innate immunity by the regulatory receptor-like kinase BAK1. PLoS Genet. 7, e1002046 (2011).
Narusaka, Y. et al. The cDNA microarray analysis using an Arabidopsis pad3 mutant reveals the expression profiles and classification of genes induced by Alternaria brassicicola attack. Plant Cell Physiol. 44, 377–387 (2003).
Frerigmann, H. et al. Regulation of pathogen-triggered tryptophan metabolism in Arabidopsis thaliana by MYB transcription factors and indole glucosinolate conversion products. Mol. Plant 9, 682–695 (2016).
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signaling in plants. Nat. Rev. Immunol. 16, 537–552 (2016).
Saijo, Y., Loo, E. P. & Yasuda, S. Pattern recognition receptors and signaling in plant–microbe interactions. Plant J. 93, 592–613 (2018).
Takahashi, T., Shibuya, H. & Ishikawa, A. SOBIR1 contributes to non-host resistance to Magnaporthe oryzae in Arabidopsis. Biosci. Biotechnol. Biochem. 80, 1577–1579 (2016).
Breitenbach, H. H. et al. Contrasting roles of apoplastic aspartyl protease Apoplastic, enhanced disease susceptibility1-dependent1 and Legume Lectin-like Protein1 in Arabidopsis systemic acquired resistance. Plant Physiol. 165, 791–809 (2014).
Leubner-Metzger, G. & Meins, F. Jr. Functions and regulation of plant ß 1,3-glucanases (PR2). In Pathogenesis-Related Proteins in Plants (eds Datta, S. K. & Muthukrishnan, S.) 49–76 (CRC Press LLC, Boca Raton, 1999).
Oh, I. S. et al. Secretome analysis reveals an Arabidopsis lipase involved in defense against Alternaria brassicicola. Plant Cell 17, 2832–2847 (2005).
Kwon, S. J. et al. GDSL lipase-like 1 regulates systemic resistance associated with ethylene signaling in Arabidopsis. Plant J. 58, 235–245 (2009).
Albert, I. et al. An RLP23–SOBIR1–BAK1 complex mediates NLP-triggered immunity. Nat. Plants 1, 15140 (2015).
Collins, N. C. et al. SNARE-protein-mediated disease resistance at the plant cell wall. Nature 425, 973–977 (2003).
Egusa, M., Miwa, T., Kaminaka, H., Takano, Y. & Kodama, M. Nonhost resistance of Arabidopsis thaliana against Alternaria alternata involves both pre- and postinvasive defenses but is collapsed by AAL-toxin in the absence of LOH2. Phytopathology 103, 733–740 (2013).
Böttcher, C. et al. The biosynthetic pathway of indole-3-carbaldehyde and indole-3-carboxylic acid derivatives in Arabidopsis. Plant Physiol. 165, 841–853 (2014).
Müller, T. M., Böttcher, C. & Glawischnig, E. Dissection of the network of indolic defence compounds in Arabidopsis thaliana by multiple mutant analysis. Phytochemistry 161, 11–20 (2019).
Glazebrook, J. & Ausubel, F. M. Isolation of phytoalexin-deficient mutants of Arabidopsis thaliana and characterization of their interactions with bacterial pathogens. Proc. Natl Acad. Sci. USA 91, 8955–8959 (1994).
Müller, T. M. et al. Transcription activator-like effector nuclease-mediated generation and metabolic analysis of camalexin-deficient cyp71a12 cyp71a13 double knockout lines. Plant Physiol. 168, 849–858 (2015).
Koch, E. & Slusarenko, A. Arabidopsis is susceptible to infection by a downy mildew fungus. Plant Cell 2, 437–445 (1990).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Acknowledgements
We thank ABRC for providing Arabidopsis seeds. We also thank Shigenobu Yoshida (Ctro) and Akira Tohyama (Ab) for providing fungal pathogens. We also thank Yoshihiro Inoue for supports on the statistical analyses. This work was supported by Grants-in-Aid for Scientific Research (15H05780, 18H02204, 18H04780, 18K19212) (KAKENHI), by grants from the Project of the NARO Bio-oriented Technology Research Advancement Institution (Research program on development of innovative technology), and by the Asahi Glass Foundation. Work in the PB laboratory was supported by the National Science Centre SONATA BIS Grant (UMO-2012/07/E/NZ2/04098).
Author information
Authors and Affiliations
Contributions
Y.T. and P.B. designed this research. A.K., M.P., M.P-B., T.N., E.O., H.S., A.I. and H.F. performed the experiments and analyzed the data. A.K., Y.T., P.B., E.O., M.K. and K.M. wrote the manuscript and prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kosaka, A., Pastorczyk, M., Piślewska-Bednarek, M. et al. Tryptophan-derived metabolites and BAK1 separately contribute to Arabidopsis postinvasive immunity against Alternaria brassicicola. Sci Rep 11, 1488 (2021). https://doi.org/10.1038/s41598-020-79562-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79562-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.