Abstract
Adaptation to different ecological environments can, through divergent selection, generate phenotypic and genetic differences between populations, and eventually give rise to new species. The fire salamander (Salamandra salamandra) has been proposed to represent an early stage of ecological speciation, driven by differential habitat adaptation through the deposition and development of larvae in streams versus ponds in the Kottenforst near Bonn (Germany). We set out to test this hypothesis of ecological speciation in an area different from the one where it was raised and we took the opportunity to explore for drivers of genetic differentiation at a landscape scale. A survey over 640 localities demonstrated the species’ presence in ponds and streams across forests, hilly terrain and areas with hedgerows (‘bocage’). Genetic variation at 14 microsatellite loci across 41 localities in and around two small deciduous forests showed that salamander effective population sizes were higher in forests than in the bocage, with panmixia in the forests (Fst < 0.010) versus genetic drift or founder effects in several of the small and more or less isolated bocage populations (Fst > 0.025). The system fits the ‘mainland-island’ metapopulation model rather than indicating adaptive genetic divergence in pond versus stream larval habitats. A reanalysis of the Kottenforst data indicated that microsatellite genetic variation fitted a geographical rather than an environmental axis, with a sharp transition from a western pond-breeding to an eastern, more frequently stream-breeding group of populations. A parallel changeover in mitochondrial DNA exists but remains to be well documented. The data support the existence of a hybrid zone following secondary contact of differentiated lineages, more so than speciation in situ.
Similar content being viewed by others
Introduction
Adaptation to different ecological environments can, through divergent selection, generate phenotypic and genetic differences between populations. These changes may eventually give rise to new species. The speciation process is often quantitative in nature, as illustrated by numerous studies showing that divergence during speciation varies continuously, and the sequence of genetically-based changes that occur as two lineages on the pathway to reproductive isolation diverge from one another has been coined the ‘speciation continuum’1,2. Divergent evolution and reproductive isolation are the primary elements of the speciation continuum, but many have recognized that reproductive isolation is usually a signature effect rather than a primary cause of speciation. Whereas the mechanisms underlying reproductive isolation are by now mostly well understood (such as natural and sexual selection and genetic drift due to founder events, etc.), biologists continue to struggle with understanding how and why these evolutionary processes cause the disjoined genetic connections that are integral to the emergence of new species, in particular in conditions of sympatry3,4,5,6. Organisms that are organized in deme-structured metapopulations, with good population sizes and small to moderate dispersal capabilities, such as many amphibian species, are good models for population genetic research and convincing examples involving the early stages of ecological speciation in this group would be particularly welcome. Unfortunately, amphibians also appear particularly prone to severe losses in recent times e.g.7,8 and this includes the fire salamander Salamandra salamandra Linnaeus, 17589,10.
The fire salamander represents a remarkable study system because from studies in the Kottenforst, near Bonn in Germany, it figures as an example of local ecological and genetic differentiation and adaptation, with pond- and stream-breeding populations possibly representing the first step in the speciation process11,12,13. It would be important to determine whether similar processes can be uncovered in other regions where the species is distributed today. Other regions may serve as replicates despite the fact that each region may have had its own contingent evolutionary specificities. To perform a replicate study, we choose the ‘département’ (department) Mayenne in the west of France where we found the fire salamanders to deposit their larvae in streams as well as in ponds.
The standing genetic variability of populations represents the adaptive potential to changing environmental conditions and acts as a buffer against stochastic and catastrophic events14,15. The protection of genetic diversity, along with that of habitats and species per se is thus a pillar to nature conservation16,17. The landscapes of the French coastal zone represent the archetypal dense hedgerow configuration known as ‘bocage’. From the perspective of biodiversity conservation prime assets of hedgerows are that they act as shelter areas for species unable to exist elsewhere in farmland and that they are usually interconnected into networks which facilitates dispersal across the landscape18. The network of hedgerows possibly supports the presence and dispersal of the fire salamander and that of other primarily forest-dwelling species. The Mayenne study area therewith affords the opportunity to explore other landscape processes in this system, such as supported by continuous forest versus a mosaic of hedgerows. It is important to document if species are reliant on the bocage because a decline through the thinning of the hedgerow network could fragment otherwise continuous species distributions. Accordingly, in the present study we aimed to determine whether adaptive divergence in pond- and stream-breeding populations of the fire salamander could be identified in the west of France. We expect genetic differentiation to be more strongly associated with a pond- versus stream-breeding habitat than with the wider geography. A second aim was to quantify the importance of the bocage as a constituent to the fire salamander habitat and to evaluate what role it plays in the population dynamics of the species. Finally, informed by our new findings, we reanalyzed the Kottenforst data in an explicit geographical context.
Results
Forest and bocage populations in the west of France
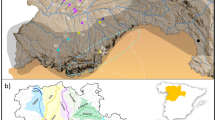
Larvae of S. salamandra were present in 251 localities (39%) and absent in 389 localities (61%). Localities are listed in a .kml file for use with e.g. Google Earth (Supplementary Information I). In logistic regression analysis the presence of the fire salamander is positively associated with forestation (P < 0.001), altitude (P < 0.001) and hedgerows (P < 0.05). At any amphibian breeding site, the probability for the occurrence of the fire salamander (Ps) is estimated by the equation (1/(1 + exp(−0.0303*percent_forest_cover-0.00562*altitude-0.0299*percent_hedgerow_cover + 1.769))). The fit of the model is expressed by the ‘area under the curve’ statistic AUC = 0.721 ± 0.020, indicating that we achieved a good description of favourable versus unfavourable fire salamander terrestrial habitats. When habitat parameter values are standardized the formula is (1/(1 + exp(a*forest_cover + b*altitude + c*hedgerow_cover + 0.0210))), indicating the relative contributions to the model in the order forest (a = −0.910), altitude (b = −0.288) and hedgerows (c = −0.275). Extrapolating the model over the entire department suggests that habitats for the fire salamander outside forests are widespread in the hilly bocage landscape in the northeastern and northwestern corners of the department Mayenne. Low habitat suitability is found in the flat and deforested southern part of Mayenne, in particular the southwest (Fig. 1). The area studied for population genetic characteristics shows low habitat suitability in between and south of the forests Forêt de Bourgon and Bois de Hermet, intermediate habitat suitability surrounding the forests in the other cardinal directions and high habitat suitability further east of Bois de Hermet.
Continental France with the department Mayenne highlighted (A) and habitat model for the Fire salamander in department Mayenne (B). The map represents the habitat suitability model Ps = (1/(1 + exp(−0.0303*percent_forest_cover-0.00562*altitude-0.0299*percent_hedgerow_cover + 1.769))) and was visualized with ILWIS 3.6 software58, available at https://52north.org/software/software-projects/ilwis/. Habitat suitability increases from deep blue with a probability of occurrence of zero to deep red with a probability of occurrence at unity (see colour bar). Prime fire salamander habitats are found at higher altitudes and are forested (in black) or with a dense hedgerow cover. Populations genetically investigated are located in and around the largely deciduous forests Forêt de Bourgon (FB) and Bois de Hermet (BH) and listed in Table 1.The outer geographical coordinates of the department are 1.239–0.049W and 47.733–48.568N.
Localities studied for population genetic differentiation included 25 ponds and 16 streams and were 23 times from inside and 18 times from outside the forest. For the subdivision pond inside – outside and streams inside – outside the forests see Table 1. A high frequency of inferred null alleles (11.7%) and a substantial amount of missing data (5.0%) were observed for the microsatellite locus C2. After exclusion of this marker and the subsequent removal of seven individuals that had data missing for more than one locus (N = 734 remaining) the frequency of null alleles was estimated at 1.6% and 0.11% of data was missing. The genotypic data are presented in Supplementary Information II. The analyses with Colony software suggested that among individuals sampled from the same locality many were full siblings. With just one representative per family group population sample sizes went down by 357 under monogamy (48.6%) and by 85 under polygamy (11.6%). This approach also revealed a high potential for false positives since inferred siblings were frequent among larvae from different localities (N = 315 under monogamy, 42.9% and N = 108 under polygamy, 14.7%). Analytical results for the three data sets are summarized in Table 2.
The number of alleles observed per locus ranged from four in locus SalE5 to 15 in locus Sal3. A single locus significantly deviated from Hardy-Weinberg expectations and there were four instances of pairs of loci showing linkage disequilibrium. The overall Fst was 0.0244. The statistical power to be able to detect the mixture of genetically differentiated populations with genetic clustering methods is dependent on sample size, the number of loci studied (14), the number of alleles per locus (average 9.4) and the proportioning of the samples over habitats, in our case sample size in streams and ponds (285/449 = 0.63) or forest versus outside forest (294/440 = 0.67). Under equation 1 of Jorde et al. (2018: 4021; see Materials and methods) the threshold for a power of 10% is Fst~0.004, which is well below the observed Fst-value of 0.024.
No association was observed between the pairwise distance matrices for genetic differentiation (Fst/(1-Fst)) and the logarithm of geographical distance (non-parametric matrix correlation, ρ = −0.033, P > 0.05), indicating the absence of isolation by distance over the study area. The observed amount of genetic differentiation is substantial but does not have a bimodal distribution as is illustrated by the UPGMA-dendrogram on Fst (Fig. 2A). A tight cluster at Fst < 0.010 is mostly composed of forest populations. Populations that join the dendrogram at higher Fst-values are mostly from outside the forests, which reflects a non-random distribution (G-test for independence, G = 11.34, df = 1, P < 0.001). At Fst > 0.025 just non-forest populations are added. No significant genetic differentiation was found for pond- versus stream inhabiting population (G = 1.35, P > 0.05). Analyses with principal components revealed a wide overlap for the forest and non-forest groups. However, the forest populations form a tight group whereas the non-forest populations are more widely scattered over the bivariate plot (Fig. 2B). Accordingly, average distance to their nadir in the PC-plots is smaller for forest than for non-forest populations (Student’s t-test, t = 3.18, df = 21.1, P < 0.01). No significant difference was found for the pond- versus stream-breeding classification (t = 0.58, df = 39, P > 0.05). The analysis with Structure yielded flat distributions with undifferentiated Q-values (Table 2). It is now realized that the methods employed frequently identify K = 2 as the top level of hierarchical structure, even when more subpopulations are actually present19.
(A) Clustering of pairwise Fst-values of fire salamander populations (Mayenne localities 1–41) with the UPGMA-method. The basal cluster at Fst < 0.010 is mostly composed of forest populations (F, 17/21 = 81%) whereas populations that join the dendrogram at higher Fst-values are mostly from the bocage (B, 14/20 = 70%). At Fst > 0.025 the contribution of the bocage populations is eight out of eight. Note that populations that join the dendrogram at the highest Fst-values are characterized by mostly small effective population sizes (Ňe ≤ 10, indicated by small open dots). (B) Populations plotted along the first and second axis of a principal component analysis. The 23 forest populations are shown by small solid round symbols and the solid ellipse represents the mean ± standard deviation. Eighteen populations from the bocage are shown by large open round symbols, with the mean ± standard deviation shown by the wider ellipse with the interrupted line.
Estimates for effective population size Ne (Ňe) were significantly higher for forest than for non-forest populations (Ňe/forested = 32.0, Ňe/non-forested = 22.4; Mann-Whitney U-test, U = 301, P < 0.05) whereas they were not significantly different for pond and stream populations (Ňe/pond = 27.0, Ňe/stream = 29.0; U = 240.5, P > 0.05). Among just pond populations, those from the forest were larger than those from outside the forest (Ňe/forested = 32.7, Ňe/non-forested = 20.8; Kruskal-Wallis test statistic 5.25, P < 0.05) (Table 1; see also Supplementary Information III). No significant indications for population genetic bottleneck effects were observed.
Mitochondrial DNA sequences were 755–756 bp. The shorter sequence (Genbank accession number MK395359, N = 40) conforms to ‘type 1b’ and the longer one (Genbank accession number MK395358, N = 89) to ‘type 1a’11. Sequences are equivalent to Genbank accession number KU24998920 with an indel at position 599. The distribution of the two mtDNA types bears no correspondence to the forest/non-forest inhabiting or to the stream- versus pond-breeding habitat classification (see Supplementary Information IV). For a compilation of mtDNA data from across northern Europe see Supplementary Information V.
Forest populations in the Kottenforst, Germany
Microsatellite genetic data for the Kottenforst were taken from Hendrix et al.21. All the investigated populations are situated inside the forest. Fire salamander larvae were observed in a variety of aquatic habitats (Table 3), with streams less frequent in the western section of the forest (one stream locality out of 27, 4%) than in the eastern section of the forest (six stream localities out of 20, 30%).
Analyses with Colony software suggested that many of the sampled individuals were full siblings. With only one representative per family group population sample sizes went down by 1628 under monogamy (63.5%) and by 1194 under polygamy (46.6%). Colony also revealed a high potential for false positives since inferred siblings were frequent among larvae from different localities (N = 624 under monogamy, 24.3% and N = 210 under polygamy, 8.2%). Results for genetic (dis)equilibria, genetic bottleneck effects and isolation by distance were markedly different for full and the sibling-excluded data sets. In the former we found 72 cases of significant deviation from Hardy-Weinberg expectations and 941 pairwise locus combinations with significant linkage disequilibrium (Table 2). Numbers were by two orders of magnitude lower in the reduced data sets, suggesting that the significant signals for genetic equilibria are largely to be attributed to the sampling of family groups. In the sibling reduced data set we identified no significant signal for population genetic bottlenecks whereas there were seven in the full data set. Finally, the signal for isolation by distance was not significant in the full data set (non-parametric matrix correlation, ρ = 0.070, P > 0.05) and significant in the reduced data sets (ρ = 0.191, P < 0.001 under monogamy and ρ = 0.119, P < 0.05 under polygamy).
Estimates for effective population size varied widely and were not significantly different for forest sections (Ňe/west = 31.2, range 4–104; Ňe/east = 55.6, range 2–272; Mann-Whitney U-test, U = 249, P > 0.05). Average effective population sizes for the Kottenforst were not significantly different from those in Mayenne (Mann-Whitney U-test, U = 1037, P > 0.05), but showed a wider range in the Kottenforst than in Mayenne (5 < Ňe < 56, see also Supplementary Information III).
The overall Fst for the studied fire salamander populations was 0.0360, which value widely surpassed the threshold for 10% analytical power of 0.0016. The UPGMA-dendrogram of pairwise Fst-values reveals two clusters that are differentiated at Fst = 0.04 (Fig. 3A). One cluster is made up of one eastern plus 14 western populations and the other cluster is made up of two western and 14 eastern populations, which reflects a non-random distribution (G-test for independence, G = 23.54, df = 1, P < 0.001). This signal is stronger than the parallel separation in pond- and stream-breeding populations (G = 4.60, P < 0.05). Populations joining the dendrogram at higher Fst levels are from either forest section. This set of populations is characterized by particularly low Ňe (average Ňe = 7.6 versus Ňe = 60.1 for the remainder; Mann-Whitney U-test, U = 439.5, P < 0.001), suggesting that a founder effect or genetic drift underlies the genetic differentiation of small populations. Analyses with principal components supports the population allocation of two spatial groups, in which the western and eastern group show no overlap along the first PC-axis if indeed small populations are excluded (Fig. 3B). The support for separation of a western versus an eastern group is several orders of magnitude stronger than that for the (more or less parallel) separation of pond- and stream-breeding salamander populations (Student’s t-test, t = 8.58, df = 29.0, P < 0.000001 versus t = 2.81, df = 45, P = 0.007401). In line with these results, the plot of the PC1-scores over a west to east axis describes a genetic transition in a sigmoid curve of the type reminiscent to those analyzed in the classical hybrid zone literature e.g.22. The cline that best fitted the data has a central position at km 365.3 of the longitudinal axis of the Universal Transverse Mercator coordinate system grid (UTM) and a width of 3.95 km (Fig. 4A). Because the PC-analysis extracted only a small proportion of the total variance in the data (2.1% along the first axis) we repeated the cline fitting procedure with the ‘proportion of pond- versus stream-breeding genotypes’ or ‘assignment probabilities’ of Hendrix et al.21 under an inferred optimal number of two genetic clusters (K = 2), derived with Structure software23,24. The cline description that best accommodates these values has a central position at km 365.1 and a width of 1.11 km (Fig. 4B). The full model descriptions are presented in Supplementary Information VI.
(A) Clustering of pairwise Fst-values of Kottenforst fire salamander populations (localities K01-K47) with the UPGMA-method. Numbers K01-K27 represent populations in the western section of the forest and K28-K47 represent populations in the eastern section of the forest. The basal cluster at Fst < 0.04 is composed of two groups (shaded) composed of mostly eastern (14/16 = 88%) or mostly western localities (14/15 = 93%). Populations breeding in streams are shown by the letter S. Note that populations that join the dendrogram at higher Fst-values are characterized by mostly small effective population sizes (Ňe ≤ 10, indicated by small open dots; X – Ňe not determined). B top panel - Populations plotted along the first and second axis of a principal component analysis. Middle panel - Ellipses represent means ± standard deviation for seven stream populations (left ellipse) and 40 non-stream populations (right ellipse). Lower panel - Ellipses represent means ± standard deviation for the western (left) and eastern (right) section of the Kottenforst, for small populations (Ňe ≤ 10) shown by interrupted lines and for larger populations (Ňe > 10) shown by uninterrupted lines. Note that for the larger populations the ellipses for western and eastern localities do not overlap.
Microsatellite population genetic data for the fire salamander in the Kottenforst, Germany21,24 analyzed in the framework of allopatric speciation, i.e. a secondary spatial contact of a western pond-breeding lineage and an eastern stream-breeding lineage. The 95% credible cline regions are shown by grey shading. Solid and open round symbols represent larger (Ňe > 10) and small populations (Ňe ≤ 10), respectively. Note that the stream-breeding populations that gave the composite genotype its name are all located in the eastern section of the Kottenforst (six data points indicated with a forward slash (/). One ‘intermittent stream’ in the western section is indicated by a backward slash. Also note the paucity of data at and around the steepest part of the clines. A – loadings on the first PC axis versus geographical distance. The cline centre is at km 365.3 of the Universal Transverse Mercator (UTM) grid. Cline width is 3952 m. B – frequency of the stream-breeding genotype versus distance (after21). The cline centre is at UTM km 365.1 and the cline width is 1108 m. For model details see Supplementary Information VI.
Discussion
Adult fire salamanders are terrestrial amphibians that deposit larvae in ponds or streams, in where these develop until metamorphosis. The species figures as an example of local ecological and genetic differentiation and adaptation, possibly representing the first step in the speciation process12,25. We set out to replicate the observation of genetic differentiation of stream- and pond-breeding populations and we sought the opportunity to explore for additional drivers of genetic differentiation at a landscape scale. However, the microsatellite data we obtained for the fire salamander in Mayenne did not yield a signal for genetic differentiation of pond- and stream-breeding populations. Instead we found genetic differences for populations from inside and outside forests.
Fire salamander population structure in Mayenne, France
The results of our habitat suitability modelling are in line with the general habitat preference of the fire salamander in Central Europe, namely mixed deciduous forest at 200–400 m a.s.l.26,27. In addition, we document the regular presence of fire salamanders outside forests, in particular in hilly areas where the network of hedges is dense. Such a pastural landscape is known as ‘bocage’. Our data suggest that forests are the primary fire salamander habitat. Firstly, forestation represents the most prominent contribution to the habitat model. Secondly, the estimates for effective population sizes are larger for forest than for bocage populations. Thirdly, forest populations are genetically most similar suggesting ongoing gene flow, presumably covering historical times. In contrast, the bocage populations are frequently genetically differentiated from one another (Fst > 0.025) indicating that they are more or less isolated and that genetic drift and founder effect operate more effectively in these small populations. In metapopulation terminology, the fire salamander system in Mayenne fits the ‘mainland-island’ model28 in which forests are the mainland and the bocage represents an archipelago of islands. In Spain, genetic differentiation was observed to increase from Fst < 0.05 among most continental S. salamandra populations to Fst~0.10 for peninsular populations separated for at least 2000 years, to Fst~0.19 for insular populations separated for 6–13 Kybp (thousands of years before present) (geological data29,30, genetic data31,32). This system, literally fitting the mainland-island model and at a comparable spatial scale to Mayenne, confirms the propensity for genetic drift in small and isolated fire salamander populations.
The two forests in Mayenne may be interconnected by dispersal through the dense but patchy and declining bocage. While population connectivity operating through the network of hedgerows is likely, the study area does not offer a good setting for testing this hypothesis. First, we noted no consistent spatial genetic signal and it will be difficult to assess if this can be attributed to the counteracting forces of isolation by distance and connectivity by habitat. Second, to be able to disentangle these effects requires an explicit spatial configuration. For example, to test for fire salamander dispersal along hedgerows, the hypothesized corridor is to deviate from a straight line. If not, genetic differentiation will, by default, be associated to geographical, not ecological distance.
We found no strong evidence for population genetic bottlenecking, suggesting that fire salamander populations may be stable over time. Another line of evidence for the long-term persistence of fire salamander populations are several occurrences in the northwest of Mayenne (Supplementary Information I)33. This area is currently devoid of forests (Fig. 2B) and also 18th century maps by the Cassini family (accessible at https://www.geoportail.gouv.fr/donnees/carte-de-cassini) show no forest at the localities where fire salamanders were actually observed, suggesting that extant populations persisted in the bocage over the centuries and providing ample opportunity for genetic drift to operate. While we could not detect significant departures from panmixia in forests, the bocage populations are more or less isolated. Yet the bocage localities may serve as ‘islands’ or ‘stepping stones’. Even a loose network may help to preserve population connectivity at a wider spatial scale and eventually promote the genetic exchange between forest populations, such as those of Forêt de Bourgon and Bois de Hermet (Fig. 2B). However, the pastural landscape in western France rapidly deteriorates from the perspective of wildlife, mostly through field size enlargements and agricultural reform, with the concomitant loss of small landscape elements such as hedgerows, spinneys and ponds18,33,34,35,36. A further deterioration of the bocage is likely to isolate the forest populations from one another. To illustrate this point, in the flat, southern part of Mayenne the bocage has largely disappeared since the Second World War37 and the fire salamander is locally rare with some scattered occurrences33 (Supplementary Information I).
Fire salamander population structure in the Kottenforst
The German Kottenforst supports two fire salamander population groups that are differentiated at Fst = 0.030 and that are proposed to represent pond- versus stream-breeders. However, the ecological data in support of this interpretation are meagre, simply because stream-breeding populations are a minority in the Kottenforst system. Moreover, focusing on large and presumably healthy populations that are locally more frequently found in streams than in ponds (Table 3) strengthens the eco-geographical distinction more so than it sharpens the genetic differentiation.
The west to east distinction is to some degree obscured by a suite of 16 populations that are more deeply genetically differentiated (0.030 < Fst < 0.153; Fig. 3). These latter populations are characterized by significantly smaller effective population sizes than average and 11 of them have Ňe ≤ 10. The small population sizes suggest that genetic drift might be prominent, but given the more or less unhampered dispersal of adult fire salamanders across the forest, populations are unlikely to be isolated and a genetic founder effect is a more plausible explanation.
With an average minimum distance of neighbouring populations of ca. 400 m the grid of localities studied in the Kottenforst is tighter than in Mayenne. Yet, unlike Mayenne, the pond- and stream-breeding populations appear to be spatially, behaviourally and genetically separated. This separation takes effect along a roughly longitudinal axis. The by approximation sigmoid shaped transition is 1.11–3.95 km wide, with inflection points that are 730–1440 m apart (Fig. 4). A cline this narrow cannot result from neutral processes exclusively. In the absence of selection, the width (w) of the cline can be predicted from a diffusion model as a function of dispersal distance (d) and the length of time since contact (t), as w = 2.51d √(t)38. Generation time is reported as six years for both sexes39 and also as three years for males and four years for females40. At an average dispersal of 200 m per generation, which is at the lower limit because it just covers the regular home range, cline widths would exceed the measured width in a couple of generations and at higher dispersal, such as distances of up to 2000 m24, the collapse of the cline would be near-instantaneous. However, the Kottenforst genetic transition is sharp, suggesting that intrinsic selection operates against the two lineages’ mixed offspring. The center of the transition is further characterized by a paucity of material studied, presumably caused by lack of aquatic habitat suitable for the deposition of larvae. This observation fits classical theory where a hybrid zone may be ‘captured’ where an area of low population density acts as a barrier to further movement41,42. If dispersal is high, selection against hybrids is to be strong. Evidence for genetic incompatibility of the lineages may have been overlooked by Caspers et al.13. These authors were surprised to find that offspring numbers in the Kottenforst increased with the degree of genetic relatedness between females and their sires. Puzzling as this observation might be for within-lineage data, in the context of selection against hybrids it makes direct sense.
The postglacial range expansion of S. salamandra has been linked to the spread of the beech, Fagus sylvatica L11,12,43. The contact zone, cq. intra-specific hybrid zone in the Kottenforst may have originated ca. 8 Kybp, by colonization from southern European glacial refuge areas, possibly involving the south of France and the northern Balkans, but more northerly locations cannot be excluded44,45. Regardless of the location of the glacial refugia, the hybrid zone in the Kottenforst appears to have been kept in check over hundreds or thousands of years, in which selection against hybrids counteracted dispersal into the zone. As an alternative explanation to in situ ecological speciation, we suggest that the Kottenforst is an area of secondary contact of a pond-breeding western lineage and a stream-breeding eastern lineage. Pond-breeding in the Kottenforst is considered a recent adaptation25,46, but we consider this unlikely because the condition is widespread in western Europe26,28,35,47,48.
An argument against the secondary contact scenario may be that the Kottenforst fire salamander populations are more similar to one another than to other, nearby populations for mitochondrial11 and nuclear genetic markers49. The fire salamander in northern Europe is characterized by two mitochondrial DNA lineages with ‘type 1’ in the west and ‘type 2’ in the east. According to Weitere et al.11 the pond- and stream-breeding populations in the Kottenforst are all descendants of the western mitochondrial DNA lineage. However, a sharp and complete transition from type 1 to type 2 is found not far away (at 70–80 km north, in between the localities Felderbachtal and Bochum; Supplementary Information V). We propose that the mtDNA cline is displaced relative to the cline from the nuclear genetic markers, a phenomenon regularly observed, especially for uniparentally transmitted markers50,51,52. Unfortunately, a wide sampling gap (>350 km) precludes measuring position, width and shape of the mtDNA transition east of the Kottenforst. With microsatellite data Steinfartz et al.49 also found that the Kottenforst populations are the most closely related in the wider area. This conclusion, however, depends on the position of the ‘root’ of the graph (which is not provided) and the supporting evidence is not unambiguous given the graph’s short internal branches. Future research might reveal a continuation of the fire salamander contact zone outside the Kottenforst and possibly show that the western and eastern lineages represent the currently described subspecies S. s. terrestris Lacépède, 1788 and S. s. salamandra. A wider phylogeographic survey is required to solve these issues.
Concluding remarks
The fire salamander is a species prone to population genetic differentiation. Substantial genetic drift has been reported for small and isolated populations across its range31,53,54,55,56. We analyzed microsatellite genetic data in two metapopulations of the fire salamander in the northwest of Europe, with contrasting results. In France we documented small, isolated, persisting and genetically differentiated populations in the bocage versus panmixia in two small deciduous forests. This population structure fits the classical mainland-island model. A mainland-island population structure may have been paramount to the fire salamander prior to the recent intensification of agriculture, the field size enlargements that came with it and the widespread conversion from pasture for cattle-breeding to arable for growing crops. We did not find a bimodal distribution coinciding with stream- and pond-reproduction as detected in the Kottenforst12,25,57. Accordingly, the fire salamander system in France offers no support for habitat driven genetic differentiation other than through drift and founder effects operating in more or less isolated bocage populations. In Germany we found small and genetically differentiated populations scattered over the Kottenforst, in addition to larger populations occupying ponds in the western section and ponds and streams in the eastern section of the forest. The finding is somewhat puzzling, given the unrestricted gene flow that we documented for the French forest populations and the rampant migration documented for the Kottenforst per se, with large home ranges and frequent long-distance dispersal24. An ad hoc explanation, supported by the high frequencies of full-siblings and low effective population sizes, is that these localities represent a particularly intensive sampling scheme, with larvae included from unusual aquatic habitats such as puddles and wheel ruts (Table 3). These small and frequently impermanent water bodies may represent sub-optimal conditions for larval growth, survival and metamorphosis and be transient satellites to the core fire salamander populations. Finally, to explain the significant spatial component in the remaining genetic variation, we propose that secondary contact between a western and an eastern fire salamander lineage better explains the available data than an ecological sympatric speciation scenario.
Materials and Methods
We recorded the presence or absence of fire salamander larvae in 640 amphibian pond and stream breeding sites across the department Mayenne. Ponds represent focal points of amphibian presence and harbour more or less isolated populations (or ‘demes’) that together constitute a metapopulation. The deme structure in streams is less obvious, but clearly not all sections of a stream are equally suitable for reproduction, for example in sections with and without predatory fish, or sections falling dry. Yet, for consistency, we consider the amphibian sites that we study to represent local populations. Locality data on altitude (m above sea level, a.s.l.), forestation and hedgerows (percent cover) were extracted from digital IGN maps (Institute Géographique Nationale) with a 25 m spatial resolution, provided by department officials. The area considered around each pond or stream sampling locality had a radius of 200 m. This scale should represent a typical fire salamander home range because 200 m is the median distance travelled by adults in a capture-recapture study in the Kottenforst24. The biological and environmental data were analyzed by logistic regression with a weight parameter, so that in analysis the number of fire salamander presences equaled the number of absences. Habitat models were visualized with ILWIS 3.658. We did not observe a disease-born population decline that was reported ca. 500 km to the northwest, in The Netherlands10.
The area selected for population genetic research is situated around two small, largely deciduous forests and positioned at the transition from flat and deforested in the south of Mayenne, to hilly with a bocage landscape in the north (Fig. 1). Larvae were captured with dip nets in 41 ponds and streams which were located inside as well as outside the forests. The larvae were released on the spot directly after tail tip tissue sampling. Altogether 741 salamander larvae were genotyped for 15 polymorphic microsatellite loci. The loci Sal3, Sal29, SalE11, SalE5, SalE6, SalE7 and SalE8 were studied following Steinfartz et al.59 and the loci B11, C2, C3, E11, G6, G9, IA6, IIA6 were studied following Hendrix et al.60. Locus SalE5 is a dinucleotide microsatellite locus and the others are composed of tetramer motifs. After a quality check (see Results) data for the locus C2 were excluded so that 14 markers remained. A total of 129 individuals from 14 localities was sequenced for the mitochondrial D-loop (control region, 756 bp) as described in Steinfartz et al.61. Nuclear genetic data from the German Kottenforst involved 2563 larval fire salamanders studied at 17 polymorphic microsatellite loci and were retrieved from Hendrix et al.21. The 47 Kottenforst sampling localities were classified in two spatial groups (‘west’ with localities K01-K27 and ‘east’ with localities K28-K47) that represent opposite sections of a sharp genetic transition (see Results).
Computer programs employed for the genetic data were as follows. FreeNA62 to analyze for the presence of null-alleles. FreeNA was run with 1000 replicates using the EM algorithm. GenePop version 4.263 to estimate Fst-values and analyze for Hardy-Weinberg and linkage (dis)equilibria. GenePop was run with dememorization number 1000, 100 batches and 1000 iterations per batch. Linkage disequilibrium was determined using the log likelihood ratio statistic. Hardy-Weinberg equilibrium and linkage disequilibrium results were interpreted under the Benjamini-Hochberg correction for multiple comparisons. Statistical power for the detection of genetically differentiated yet sympatric populations was estimated following Jorde et al.64. The genetic diversity among populations was summarized by clustering with the unweighted pair group method with arithmetic mean with Primer-e software (UPGMA65) and by principal component analysis (PC) with Adegenet version 2.0.0 following the manuals66,67. We also carried out a Bayesian assignment analysis with Structure68, to which we adopted the program settings for the similar analyses by Hendrix et al.21. The parameter K (the number of genetic groups suggested by the data) was evaluated under the ‘Evanno-criterion’ that is implemented in StructureHarvester69. We used Colony version 2.0.6.270 to analyze family groups and to estimate effective population size (Ňe) from the frequency of siblings. Settings for Colony were both sexes either monogamous or polygamous (see below), no allelic dropout or typing errors, diploid, codominant markers, no inbreeding, scaled full sibship, unknown population allele frequency, ten ‘medium long’ or ‘very long’ runs under full-likelihood with ‘medium’ precision and a ‘strong/optimal’ sibship prior for effective population size. Results for Ňe reported are under α is zero, assuming that deviations from Hardy-Weinberg equilibrium are negligible. We used Bottleneck71 to test for reductions in effective population size by considering that alleles are generally lost faster than heterozygosity and thus, populations that have experienced a reduction in effective population size are expected to have excess heterozygosity relative to that expected under mutation-drift equilibrium. Samples smaller than for ten individuals were ignored. Statistical relevance of the results were determined with the Wilcoxon signed-rank test. We applied the two phase model with 93.6% single step mutations, variance 30 and 10,000 replications. The proportion of single-step mutations was determined with Misat version 1.072 with dimer code 2, tetramer code 4, gridsize 40, moments estimated under the one-step model, 100,000 runs through the Markov chain, with continuously updating the value of θ0. We used HZAR73 to formalize the geographical cline suggested by the first PC-axis for the Kottenforst. The chain length was 100,000 with a burn-in of 10%, randomized seeds and model selection based on AICc scores. For details on the cline fitting procedure and model selection see e.g.74. Mitochondrial DNA sequences were aligned with ClustalW version 275 under default settings. Matrix correlations were done with Primer-e65 under 10,000 permutations. Other statistical analyses were with SPSS v. 2076.
Reproduction in the fire salamander may be monogamous or polygamous. The available data indicate that both systems occur in nature about equally frequent49 and we explored both possibilities. Including siblings in analyses of population diversity and structure can introduce a variety of biases (77 and references therein). Clutch sizes in the fire salamander are ca. 3027 and the more or less frequent sampling of full- and half-siblings in our study cannot be excluded. However, excluding all but one members of a family group is not exempt from problems neither and will cause other biases78,79. Considering the moderate number of markers and alleles in either of the two data sets we tested for the presence of full-siblings in the material at two levels. Firstly, we reconstructed family groups per population. The numbers of inferred full-siblings were substantial. Secondly, we compared all individuals across localities. Although female fire salamanders may deposit larvae in more than one water body13 this phenomenon will be rare in our sampling of disparate localities with average minimum distances of ca. 1.1 km in Mayenne and 400 m in the Kottenforst. Consequently, inferred full-siblings from different localities possibly qualify as false positives. It is altogether not straightforward to determine the balance between redundant information from siblings included versus independent information from false positives excluded. We here present results from the full data set not to hamper comparability with published results24, with the parallel results for the siblings excluded data set in Table 2. The mitochondrial DNA data considered include80.
Ethics statement
Methods were carried out in accordance with the relevant guidelines and regulations. The tissue sampling protocol was approved by the Research Coordination Office of Naturalis Biodiversity Center. Fieldwork was carried under out under license from the ‘Service Aménagement Environnement-Chasse, Direction départementale de l’Agriculture et de la Forêt, Mayenne arrêt no. 2003-A-207.
Data availability
The genotypic data for fire salamanders from Mayenne, France are presented in Supplementary Information II. The data for the Kottenforst, Germany are accessible at https://doi.org/10.5061/dryad.h0r6q.
Change history
24 April 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Hendry, A. P., Bolnick, D. I., Berner, D. & Peichel, C. L. Along the speciation continuum in sticklebacks. Journal of Fish Biology 75, 2000–2036, https://doi.org/10.1111/j.1095-8649.2009.02419.x (2009).
Nosil, P. Ecological Speciation. Oxford University Press: Oxford, United Kingdom, https://doi.org/10.1093/acprof:osobl/9780199587100.001.0001 (2012).
Harrison, R. G. Molecular changes at speciation. Annual Review of Ecology and Systematics 22, 281–308, https://doi.org/10.1146/annurev.ecolsys.22.1.281 (1991).
Coyne, J. A. & Price, T. D. Little evidence for sympatric speciation in island birds. Evolution 54, 2166–2171, https://doi.org/10.1554/0014-3820(2000)054[2166:LEFSSI]2.0.CO;2 (2000).
Shaw, K. L. & Mullen, S. P. Genes versus phenotypes in the study of speciation. Genetica 139, 649–661, https://doi.org/10.1007/s10709-011-9562-4 (2011).
Shaw, K.L., Mullen, S.P. Speciation continuum. Journal of Heredity 105, (Special Issue), 741–742, https://doi.org/10.1093/jhered/esu060 (2014).
Allentoft, M. E. & O’Brien, J. Global amphibian declines, loss of genetic diversity and fitness: a review. Diversity 2, 47–71, https://doi.org/10.3390/d2010047 (2010).
McCartney‐Melstad, E. & Shaffer, H. B. Amphibian molecular ecology and how it has informed conservation. Molecular Ecology 24, 5084–5109, https://doi.org/10.1111/mec.13391 (2015).
Martel, A. et al. Recent introduction of a chytrid fungus endangers Western Palearctic salamanders. Science 346, 630–631, https://doi.org/10.1126/science.1258268 (2014).
Spitzen-van der Sluijs, A. et al. Rapid enigmatic decline drives the fire salamander (Salamandra salamandra) to the edge of extinction in the Netherlands. Amphibia-Reptilia 34, 233–239, https://doi.org/10.1163/15685381-00002891 (2013).
Weitere, M., Tautz, D., Neumann, D. & Steinfartz, S. Adaptive divergence vs. environmental plasticity: tracing local genetic adaptation of metamorphosis traits in salamanders. Molecular Ecology 13, 1665–1677, https://doi.org/10.1111/j.1365-294X.2004.02155.x (2004).
Steinfartz, S., Weitere, M. & Tautz, D. Tracing the first step to speciation: ecological and genetic differentiation of a salamander population in a small forest. Molecular Ecology 16, 4550–4561, https://doi.org/10.1111/j.1365-294X.2007.03490.x (2007).
Caspers, B. A. et al. The more the better – polyandry and genetic similarity are positively linked to reproductive success in a natural population of terrestrial salamanders (Salamandra salamandra). Molecular Ecology 23, 239–250, https://doi.org/10.1111/mec.12577 (2014).
Frankham, R.; Ballou, J. D., Briscoe, D.A. Introduction to Conservation Genetics. Cambridge University Press: Cambridge, United Kingdom, https://doi.org/10.1017/CBO9780511808999 (2002).
Willi, Y., van Buskirk, J. & Hoffmann, A. A. Limits to the adaptive potential of small populations. Annual Review of Ecology and Systematics 37, 433–458, https://doi.org/10.1146/annurev.ecolsys.37.091305.110145 (2006).
Balkenhol, N. et al. Identifying future research needs in landscape genetics: where to from here? Landscape Ecology 24, 455–463, https://doi.org/10.1007/s10980-009-9334-z (2009).
Hoban, S. M. et al. Bringing genetic diversity to the forefront of conservation policy and management. Conservation Genetics Resources 5, 593–598, https://doi.org/10.1007/s12686-013-9859-y (2013).
Baudry, J., Bunce, R. G. & Burel, F. Hedgerows: an international perspective on their origin, function and management. Journal of Environmental Management 60, 7–22, https://doi.org/10.1006/jema.2000.0358 (2000).
Janes, J. K. et al. The K = 2 conundrum. Molecular Ecology 26, 3594–3602, https://doi.org/10.1111/mec.14187 (2017).
Pereira, R. J., Martínez‐Solano, I. & Buckley, D. Hybridization during altitudinal range shifts: nuclear introgression leads to extensive cyto‐nuclear discordance in the fire salamander. Molecular Ecology 25, 1551–1565, https://doi.org/10.1111/mec.13575 (2016).
Hendrix, R., Schmidt, B. R., Schaub, M., Krause, E. T., Steinfartz, S. Data from: Differentiation of movement behavior in an adaptively divergng salamander population. Dryad Digital Repository., https://doi.org/10.5061/dryad.h0r6q (2017).
Pannekoek, A. J. The Ria problem. The role of antecedence, deep weathering, and Pleistocene slope-wash in the formation of the west-Galician Rias. Tijdschrift Koninklijk Nederlands Aardrijkskundig Genootschap 83, 289–297 (1966).
Szymura, J. M. & Barton, N. H. Genetic analysis of a hybrid zone between the fire‐bellied toads, Bombina bombina and B. variegata, near Cracow in southern Poland. Evolution 40, 1141–1159, https://doi.org/10.1111/j.1558-5646.1986.tb05740.x (1986).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Hendrix, R., Schmidt, B. R., Schaub, M., Krause, E. T. & Steinfartz, S. Differentiation of movement behaviour in an adaptively diverging salamander population. Molecular Ecology 26, 6400–6413, https://doi.org/10.1111/mec.14345 (2017).
Caspers, B. A., Steinfartz, S. & Krause, E. T. Larval deposition behaviour and maternal investment of females reflect differential habitat adaptation in a genetically diverging salamander population. Behavioral Ecology and Sociobiology 69, 407–413, https://doi.org/10.1007/s00265-014-1853-1 (2015).
Thiesmeier, B. Ökologie des Feuersalamanders. Westarp Wissenschaften: Essen, Germany (1992).
Thiesmeier, B. & Grossenbacher, K. Salamandra salamandra (Linnaeus, 1758) —Feuersalamander. Pp. 1059-1132 in: Handbuch der Reptilien und Amphibien Europas: Schwanzlurche IIB. Eds. Thiesmeier, B. and K. Grossenbacher. Aula Verlag: Wiebelsheim, Germany (2004).
Hanski, I. & Gilpin, M. Metapopulation dynamics: brief history and conceptual domain. Biological Journal of the Linnean Society 42, 3–16, https://doi.org/10.1111/j.1095-8312.1991.tb00548.x (1991).
Dias, J. M. A., Boski, T., Rodrigues, A. & Magalhaes, F. Coast line evolution in Portugal since the Last Glacial Maximum until present – a synthesis. Marine Geology 170, 177–186, https://doi.org/10.1016/S0025-3227(00)00073-6 (2000).
Velo-Antón, G., Zamudio, K. R. & Cordero-Rivera, A. Genetic drift and rapid evolution of viviparity in insular fire salamanders (Salamandra salamandra). Heredity 108, 410–418, https://doi.org/10.1038/hdy.2011.91 (2012).
Lourenço, A., Sequeira, F., Buckley, D. & Velo‐Antón, G. Role of colonization history and species‐specific traits on contemporary genetic variation of two salamander species in a Holocene island‐mainland system. Journal of Biogeography 45, 1054–1066, https://doi.org/10.1111/jbi.13192 (2018).
Baudin, B. Amphibiens et Reptiles de la Mayenne. Mayenne Nature Environnement: Laval, France (2010).
Schoorl, J. & Zuiderwijk, A. Ecological isolation in Triturus cristatus and Triturus marmoratus (Amphibia: Salamandridae). Amphibia-Reptilia 1, 235–252, https://doi.org/10.1163/156853881X00357 (1980).
Arntzen, J. W., Abrahams, C., Meilink, W. R. M., Iosif, R. & Zuiderwijk, A. Amphibian decline, pond loss and reduced population connectivity under agricultural intensification over a 38 year period. Biodiversity and Conservation 26, 1411–1430, https://doi.org/10.1007/s10531-017-1307-y (2017).
Boissinot, A., Besnard, A. & Lourdais, O. Amphibian diversity in farmlands: combined influences of breeding-site and landscape attributes in western France. Agriculture Ecosystems and Environment 269, 51–61, https://doi.org/10.1016/j.agee.2018.09.016 (2019).
Visser, M., de Leeuw, M., Zuiderwijk, A. & Arntzen, J. W. Stabilization of a salamander moving hybrid zone. Ecology and Evolution 7, 689–696, https://doi.org/10.1002/ece3.2676 (2017).
Barton, N. H., Gale, K. S. Genetic analysis of hybrid zones. Pp. 13-45 in Harrison RG. Hybrid Zones and the Evolutionary Process. Oxford University Press, Oxford, United Kingdom(1993).
Seifert, D. Untersuchungen an einer ostthüringischen Population des Feuersalamanders. Salamandra salamandra. Artenschutzreport 1, 1–6 (1991).
Trochet, A. et al. A database of life-history traits of European amphibians. Biodiversity Data Journal 2, https://doi.org/10.3897/BDJ.2.e4123 (2014).
Barton, N. H. The dynamics of hybrid zones. Heredity 43, 341–359, https://doi.org/10.1038/hdy.1979.87 (1979).
Barton, N. H. & Hewitt, G. M. Analysis of hybrid zones. Annual Review of Ecology and Systematics 16, 113–148, https://doi.org/10.1146/annurev.es.16.110185.000553 (1985).
Eiselt, J. Der Feuersalamander Salamandra salamandra (L.). Beiträge zu einer taxonomischen Synthese. Abhandlungen und Berichte für Naturkunde und Vorgeschichte Magdeburg 10, 77–154 (1959).
Bhagwat, S. A. & Willis, K. J. Species persistence in northerly glacial refugia of Europe: a matter of chance or biogeographical traits? Journal of Biogeography 35, 464–482, https://doi.org/10.1111/j.1365-2699.2007.01861.x (2008).
Magri, D. et al. A new scenario for the Quaternary history of European beech populations: palaeobotanical evidence and genetic consequences. New Phytologist 171, 199–221, https://doi.org/10.1111/j.1469-8137.2006.01740.x (2006).
Reinhardt, T. S., Steinfartz, S., Paetzold, A. & Weitere, M. Linking the evolution of habitat choice to ecosystem functioning: direct and indirect effects of pond-reproducing fire salamanders on aquatic-terrestrial subsidies. Oecologia 173, 281–291, https://doi.org/10.1007/s00442-013-2592-0 (2013).
Thiesmeier, B. & Mutz, T. Zur Laichzeit und Larvalentwicklung des Feuersalamanders (Salamandra salamandra terrestris) im nordwestdeutschen Tiefland. Zeitschrift für Feldherpetologie 4, 115–125 (1997).
Duguet, R., Melki, F. (eds) Les Amphibiens de France, Belgique et Luxembourg. Biotope: Mèze, France (2003).
Steinfartz, S., Stemshorn, K., Kuesters, D. & Tautz, D. Patterns of multiple paternity within and between annual reproduction cycles of the fire salamander (Salamandra salamandra) under natural conditions. Journal of Zoology 268, 1–8, https://doi.org/10.1111/j.1469-7998.2005.00001.x (2006).
Toews, D. P. L. & Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Molecular Ecology 21, 3907–3930, https://doi.org/10.1111/j.1365-294X.2012.05664.x (2012).
Sloan, D. B., Havird, J. C. & Sharbrough, J. The on‐again‐off‐again relationship between mitochondrial genomes and species boundaries. Molecular Ecology 26, 2212–2236, https://doi.org/10.1111/mec.13959 (2016).
Bonnet, T., Leblois, R., Rousset, F. & Crochet, P. ‐A. A reassessment of explanations for discordant introgressions of mitochondrial and nuclear genomes. Evolution 71, 2140–2158, https://doi.org/10.1111/evo.13296 (2017).
Álvarez, D., Lourenço, A., Oro, D. & Velo-Antón, G. Assessment of census (N) and effective population size (Ne) reveals consistency of Ne single-sample estimators and a high Ne/N ratio in an urban and isolated population of fire salamanders. Conservation Genetics Resources 7, 705–712, https://doi.org/10.1007/s12686-015-0480-0 (2015).
Lourenço, A., Alvarez, D., Wang, I. J. & Velo-Antón, G. Trapped within the city: integrating demography, time since isolation and population-specific traits to assess the genetic effects of urbanization. Molecular Ecology 26, 1498–1514, https://doi.org/10.1111/mec.14019 (2017).
Lourenço, A., Antunes, B., Wang, I. J. & Velo-Antón, G. Fine-scale genetic structure in a salamander with two reproductive modes: Does reproductive mode affect dispersal? Evolutionary Ecology 32, 699–732, https://doi.org/10.1007/s10682-018-9957-0 (2018).
Konowalik, A., Najbar, A., Babik, W., Steinfartz, S. & Ogielska, M. Genetic structure of the fire salamander Salamandra salamandra in the Polish Sudetes. Amphibia-Reptilia 37, 405–415, https://doi.org/10.1163/15685381-00003071 (2016).
Krause, E. T. & Caspers, B. A. The influence of a water current on the larval deposition pattern of females of a diverging fire salamander population (Salamandra salamandra). Salamandra 51, 156–60 (2015).
ILWIS. Integrated Land and Water Information System (ILWIS). Open software version 3.6. ITC: Enschede, The Netherlands (2009).
Steinfartz, S., Kuesters, D. & Tautz, D. Isolation and characterization of polymorphic tetranucleotide microsatellite loci in the Fire salamander Salamandra salamandra (Amphibia: Caudata). Molecular Ecology Notes 4, 626–628, https://doi.org/10.1111/j.1471-8286.2004.00716.x (2004).
Hendrix, R., Hauswaldt, S., Veith, M. & Steinfartz, S. Strong correlation between cross‐amplification success and genetic distance across all members of ‘True Salamanders’(Amphibia: Salamandridae) revealed by Salamandra salamandra‐specific microsatellite loci. Molecular Ecology Resources 10, 1038–1047, https://doi.org/10.1111/j.1755-0998.2010.02861.x (2010).
Steinfartz, S., Veith, M. & Tautz, D. Mitochondrial sequence analysis of Salamandra taxa suggests old splits of major lineages and postglacial recolonizations of Central Europe from distinct source populations of Salamandra salamandra. Molecular Ecology 9, 397–410, https://doi.org/10.1046/j.1365-294x.2000.00870.x (2000).
Chapuis, M.-P. & Estoup, A. Microsatellite null alleles and estimation of population differentiation. Molecular Biology and Evolution 24, 621–631, https://doi.org/10.1093/molbev/msl191 (2007).
Rousset, F. GenePop’007: a complete re‐implementation of the GenePop software for Windows and Linux. Molecular Ecology Resources 8, 103–106, https://doi.org/10.1111/j.1471-8286.2007.01931.x (2008).
Jorde, P. E., Andersson, A., Ryman, N. & Laikre, L. Are we underestimating the occurrence of sympatric populations? Molecular Ecology 27, 4011–4025, https://doi.org/10.1111/mec.14846 (2018).
Clarke, K. R., Gorley, R. N. Primer v6: User Manual/Tutorial. Primer-e: Plymouth, United Kingdom (2006).
Jombart, T. Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405, https://doi.org/10.1093/bioinformatics/btn129 (2008).
Jombart, T., Pontier, D. & Dufour, A. B. Genetic markers in the playground of multivariate analysis. Heredity 102, 330, https://doi.org/10.1038/hdy.2008.130 (2009).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Earl, D. A. & vonHoldt, B. M. Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources 4, 359–361, https://doi.org/10.1007/s12686-011-9548-7 (2012).
Jones, O. R. & Wang, J. COLONY: a program for parentage and sibship inference from multilocus genotype data. Molecular Ecology Resources 10, 551–555, https://doi.org/10.1111/j.1755-0998.2009.02787.x (2010).
Piry, S., Luikart, G. & Cornuet, J. M. BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. Journal of Heredity 90, 502–503, https://doi.org/10.1093/jhered/90.4.502 (1999).
Nielsen, R. A maximum likelihood approach to population samples of microsatellite alleles. Genetics 146, 711–716 (1997).
Derryberry, E. P., Derryberry, G. E., Maley, J. M. & Brumfield, R. T. HZAR: hybrid zone analysis using an R software package. Molecular Ecology Resources 14, 652–663, https://doi.org/10.1111/1755-0998.12209 (2014).
Prada, C. & Hellberg, M. E. Strong natural selection on juveniles maintains a narrow adult hybrid zone in a broadcast spawner. American Naturalist 184, 702–713, https://doi.org/10.1086/678403 (2014).
Larkin, M. A. et al. ClustalW and ClustalX version 2. Bioinformatics 23, 2947–2948, https://doi.org/10.1093/bioinformatics/btm404 (2007).
IBM SPSS. Statistical Package for the Social Sciences. SPSS Inc., Chicago, USA (2016).
Goldberg, C. S. & Waits, L. P. Quantification and reduction of bias from sampling larvae to infer population and landscape genetic structure. Molecular Ecology Resources 10, 304–313, https://doi.org/10.1111/j.1755-0998.2009.02755.x (2010).
Sánchez-Montes, G., Ariño, A. H., Vizmanos, J. L., Wang, J. & Martínez-Solano, I. Effects of sample size and full sibs on genetic diversity characterization: a case study of three syntopic Iberian pond-breeding amphibians. Journal of Heredity 108, 535–543, https://doi.org/10.1093/jhered/esx038 (2017).
O’Connell, K. A., Mulder, K. P., Maldonado, J., Currie, K. L. & Ferraro, D. M. Sampling related individuals within ponds biases estimates of population structure in a pond‐breeding amphibian. Ecology and Evolution 9, 3620–3636, https://doi.org/10.1002/ece3.4994 (2019).
Najbar, A., Babik, W., Najbar, B. & Ogielska, M. Genetic structure and differentiation of the fire salamander Salamandra salamandra at the northern margin of its range in the Carpathians. Amphibia-Reptilia 36, 301–331, https://doi.org/10.1163/15685381-00003005 (2015).
Acknowledgements
We thank Maarten de Leeuw, Jaap Schoorl, Machiel Visser, Pieter de Wijer and Annie Zuiderwijk for field observations and Mrs. I. Brugioni of the ‘Conseil départemental de la Mayenne’, Laval, for environmental data in digital format. J.v.B. thanks Ralf Hendrix, Gabrielle Keunecke and Sebastian Steinfartz for hospitality and advice in the laboratory and the Jan-Joost ter Pelkwijk fund for a travel grant.
Author information
Authors and Affiliations
Contributions
J.W.A. conceived and designed the study, and organized the species inventory. The files SI I and SI V are also available at https://www.repository.naturalis.nl/record/707616. J.v.B. collected the tissue samples and performed the laboratory work in the laboratory of S. Steinfartz, Braunschweig. J.W.A. analyzed the data and wrote the manuscript with the help of J.v.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arntzen, J.W., van Belkom, J. ‘Mainland-island’ population structure of a terrestrial salamander in a forest-bocage landscape with little evidence for in situ ecological speciation. Sci Rep 10, 1700 (2020). https://doi.org/10.1038/s41598-020-58551-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58551-0
This article is cited by
-
Fine scale genetic structure in fire salamanders (Salamandra salamandra) along a rural-to-urban gradient
Conservation Genetics (2021)
-
Life history modelling reveals trends in fitness and apparent survival of an isolated Salamandra salamandra population in an urbanised landscape
European Journal of Wildlife Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.