Abstract
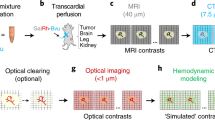
The pathogenesis of cancer and cardiovascular diseases is subjected to spatiotemporal regulation by the tissue microenvironment. Multiplex visualization of the microenvironmental components, including immune cells, vasculature and tissue hypoxia, provides critical information underlying the disease progression and therapy resistance, which is often limited by imaging depth and resolution in large-volume tissues. To this end, light sheet fluorescence microscopy, following tissue clarification and immunostaining, may generate three-dimensional high-resolution images at a whole-organ level. Here we provide a detailed description of light sheet fluorescence microscopy imaging analysis of immune cell composition, vascularization, tissue perfusion and hypoxia in mouse normal brains and hearts, as well as brain tumors. We describe a procedure for visualizing tissue vascularization, perfusion and hypoxia with a transgenic vascular labeling system. We provide the procedures for tissue collection, tissue semi-clearing and immunostaining. We further describe standard methods for analyzing tissue immunity and vascularity. We anticipate that this method will facilitate the spatial illustration of structure and function of the tissue microenvironmental components in cancer and cardiovascular diseases. The procedure requires 1–2 weeks and can be performed by users with expertise in general molecular biology.
Key points
-
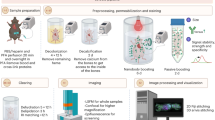
The protocol covers tissue preparation, polymerization, tissue semi-clearing, immunostaining, refractive index matching and mounting, imaging, and data processing and analysis.
-
The mild processing of samples ensures structural and molecular stability, preserving most of the proteins of interest for antibody-based immunostaining.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
References
Anderson, N. M. & Simon, M. C. The tumor microenvironment. Curr. Biol. 30, R921–R925 (2020).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550 (2018).
Sun, K., Li, Y. Y. & Jin, J. A double-edged sword of immuno-microenvironment in cardiac homeostasis and injury repair. Signal Transduct. Target Ther. 6, 79 (2021).
Jain, R. K. et al. Angiogenesis in brain tumours. Nat. Rev. Neurosci. 8, 610–622 (2007).
Jain, R. K. Normalization of tumor vasculature: an emerging concept in antiangiogenic therapy. Science 307, 58–62 (2005).
Lamplugh, Z. & Fan, Y. Vascular microenvironment, tumor immunity and immunotherapy. Front. Immunol. 12, 811485 (2021).
Zhang, D. et al. PHGDH-mediated endothelial metabolism drives glioblastoma resistance to chimeric antigen receptor T cell immunotherapy. Cell Metab. 35, 517–534 e518 (2023).
Ma, W. et al. Targeting PAK4 to reprogram the vascular microenvironment and improve CAR-T immunotherapy for glioblastoma. Nat. Cancer 2, 83–97 (2021).
Wu, X., Reboll, M. R., Korf-Klingebiel, M. & Wollert, K. C. Angiogenesis after acute myocardial infarction. Cardiovasc Res. 117, 1257–1273 (2021).
Harada, K. et al. Vascular endothelial growth factor administration in chronic myocardial ischemia. Am. J. Physiol. 270, H1791–H1802 (1996).
Huang, M. et al. Endothelial plasticity drives aberrant vascularization and impedes cardiac repair after myocardial infarction. Nat. Cardiovasc. Res. 1, 372–388 (2022).
Ntziachristos, V. Going deeper than microscopy: the optical imaging frontier in biology. Nat. Methods 7, 603–614 (2010).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Renier, N. et al. IDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Cai, R. et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull–meninges connections. Nat. Neurosci. 22, 317–327 (2019).
Qi, Y. et al. FDISCO: advanced solvent-based clearing method for imaging whole organs. Sci. Adv. 5, eaau8355 (2019).
Hahn, C. et al. High-resolution imaging of fluorescent whole mouse brains using stabilised organic media (sDISCO). J. Biophotonics https://doi.org/10.1002/jbio.201800368 (2019).
Pan, C. et al. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat. Methods 13, 859–867 (2016).
Tainaka, K. et al. Chemical landscape for tissue clearing based on hydrophilic reagents. Cell Rep. 24, 2196–2210.e2199 (2018).
Tainaka, K., Kuno, A., Kubota, S. I., Murakami, T. & Ueda, H. R. Chemical principles in tissue clearing and staining protocols for whole-body cell profiling. Annu. Rev. Cell. Dev. Biol. 32, 713–741 (2016).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Murakami, T. C. et al. A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nat. Neurosci. 21, 625–637 (2018).
Susaki, E. A. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 10, 1709–1727 (2015).
Hama, H. et al. ScaleS: an optical clearing palette for biological imaging. Nat. Neurosci. 18, 1518–1529 (2015).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Richardson, D. S. et al. Tissue clearing. Nat. Rev. Methods Primers https://doi.org/10.1038/s43586-021-00080-9 (2021).
Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Ranchoux, B. et al. Endothelial-to-mesenchymal transition in pulmonary hypertension. Circulation 131, 1006–1018 (2015).
Huang, M. et al. c-Met-mediated endothelial plasticity drives aberrant vascularization and chemoresistance in glioblastoma. J. Clin. Invest. 126, 1801–1814 (2016).
Kisanuki, Y. Y. et al. Tie2–Cre transgenic mice: a new model for endothelial cell-lineage analysis in vivo. Dev. Biol. 230, 230–242 (2001).
De Palma, M. et al. Tie2 identifies a hematopoietic lineage of proangiogenic monocytes required for tumor vessel formation and a mesenchymal population of pericyte progenitors. Cancer Cell 8, 211–226 (2005).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Cormack, B. P., Valdivia, R. H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Dodt, H. U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Hutchinson, M. H., Dorgan, J. R., Knauss, D. M. & Hait, S. B. Optical properties of polylactides. J. Polym. Environ. 14, 119–124 (2006).
Eliat, F. et al. Tissue clearing may alter emission and absorption properties of common fluorophores. Sci. Rep. 12, 5551 (2022).
Acknowledgements
This work was supported in part by National Institutes of Health grants R01NS094533 (to Y.F.), R01NS106108 (to Y.F.), R01CA241501 (to J.F.D. and Y.F.) and R01HL155198 (to Y.G. and Y.F.), and by American Heart Association grants Innovative Project Award (to Y.F.), Transformational Project Award (to Y.G.) and Predoctoral Fellowship (to D.Z.).
Author information
Authors and Affiliations
Contributions
D.Z. and Y.F. initiated the project. D.Z. performed most of the experiments and originally developed the protocol. D.Z., A.H.C., E.K., K.H., A.L.S. and J.F.D. designed the experiments. D.Z., C.Y. and W.Z. contributed to imaging analysis. Y.G. and Y.F. supervised the project. D.Z., E.K. and Y.F. wrote the manuscript. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Hiroki Ueda and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Wang, Q. et al. Nat. Commun. 9, 559 (2018): https://doi.org/10.1038/s41467-018-03050-0
Ma, W. et al. Nat. Cancer 2, 83–97 (2021): https://doi.org/10.1038/s43018-020-00147-8
Huang, M. et al. Nat. Cardiovasc. Res. 1, 372–388 (2022): https://doi.org/10.1038/s44161-022-00047-3
Zhang, D. et al. Cell Metab. 35, 517–534 (2023): https://doi.org/10.1016/j.cmet.2023.01.010
Extended data
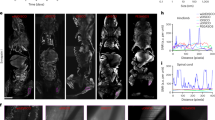
Extended Data Fig. 1 LSFM analysis of healthy mouse brain left hemisphere.
Healthy Cdh5-CreERT2;LSL-tdTomato mice were perfused with DyLight 649-lectin. Brain tissue was excised, followed by tissue clearing and LSFM imaging. Each minor tick on the grid represents 1 mm.
Extended Data Fig. 2 LSFM analysis of tissue samples after long-term storage.
a, GBM was induced in Cdh5-CreERT2;LSL-tdTomato mice, followed by perfused with hypoxyprobe. Tissue samples were subjected to tissue clearing, and stored in RIMS at 4°C for 2.5 years and imaged by LSFM. Each minor tick on the grid represents 1 mm. b, MI was induced in Cdh5-CreERT2;LSL-tdTomato mice, followed by perfused with lectin. Tissue samples were subjected to tissue clearing, and stored in RIMS at 4°C for 2.5 years and imaged by LSFM. Each minor tick on the grid represents 0.5 mm.
Extended Data Fig. 3 Key steps of image processing and analysis in Imaris.
a, Voxel size correction in “Image Properties” (Step 42). b, Crop out ROI in “Crop 3D” (Step 43). c, “Image Processing” module (Step 44). d, Setting deconvolution parameters in the “Image Processing” module (Step 44). e, Preprocessing the image using “Normalize Layers” and “Background Subtraction “ (Steps 45, 46). f, Creating objects for feature registration (Step 48). g, Initial parameters for creating surface objects (Step 48A(ii)). h, Thresholding of surface object creation (Step 48A(iii)). i, Filtering generated surface objects by size (Step 48A(iv)). j, Multiscale Points method of generating filament seed points (Step 48B(ii)). k, Seed point filtering by vessel diameter (Step 48B(ii)). l, Segment classification and filtering by machine learning-based model (Step 48B(iii)). m, Initial parameters for creating spot objects (Step 48C(i)). n, Spot detection by feature diameter (Step 48C(ii)). o, Filter Spot feature by quality and diameter/size (Step 48C(iii–iv)).
Supplementary information
Supplementary Software 1
3D printing file for ETC sample holder.
Supplementary Software 2
3D printing file for sample imaging holder.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, D., Cleveland, A.H., Krimitza, E. et al. Spatial analysis of tissue immunity and vascularity by light sheet fluorescence microscopy. Nat Protoc 19, 1053–1082 (2024). https://doi.org/10.1038/s41596-023-00941-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00941-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.