Abstract
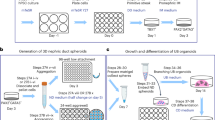
Protocols have been established to direct the differentiation of human induced pluripotent stem (iPS) cells into nephron progenitor cells and organoids containing many types of kidney cells, but it has been difficult to direct the differentiation of iPS cells to form specific types of mature human kidney cells with high yield. Here, we describe a detailed protocol for the directed differentiation of human iPS cells into mature, post-mitotic kidney glomerular podocytes with high (>90%) efficiency within 26 d and under chemically defined conditions, without genetic manipulations or subpopulation selection. We also describe how these iPS cell–derived podocytes may be induced to form within a microfluidic organ-on-a-chip (Organ Chip) culture device to build a human kidney Glomerulus Chip that mimics the structure and function of the kidney glomerular capillary wall in vitro within 35 d (starting with undifferentiated iPS cells). The podocyte differentiation protocol requires skills for culturing iPS cells, and the development of a Glomerulus Chip requires some experience with building and operating microfluidic cell culture systems. This method could be useful for applications in nephrotoxicity screening, therapeutic development, and regenerative medicine, as well as mechanistic study of kidney development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Thomson, J. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Benam, K. H. et al. Engineered in vitro disease models. Annu. Rev. Pathol. 10, 195–262 (2015).
Tabar, V. & Studer, L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat. Rev. Genet. 15, 82–92 (2014).
Musah, S. et al. Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip. Nat. Biomed. Eng. 1, 0069 (2017).
Ingber, D. E. & Musah, S. Methods for generation of podocytes from pluripotent stem cells and cells produced by the same. US patent application no. 14/950859 (2015).
Mooney, D. J., Langer, R. & Ingber, D. E. Cytoskeletal filament assembly and the control of cell spreading and function by extracellular matrix. J. Cell Sci. 108, 2311–20 (1995).
Jones, L. & Wagers, A. No place like home: anatomy and function of the stem cell niche. Nat. Rev. Mol. Cell Biol. 9, 11–21 (2008).
Ingber, D. Mechanical control of tissue morphogenesis during embryological development. Int. J. Dev. Biol. 50, 255–266 (2006).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681–686 (2007).
Mummery, C. et al. Differentiation of human embryonic stem cells and induced pluripotent stem cells to cardiomyocytes: a methods overview. Circ. Res. 111, 344–358 (2012).
Derda, R. et al. High-throughput discovery of synthetic surfaces that support proliferation of pluripotent cells. J. Am. Chem. Soc. 132, 1289–1295 (2010).
Musah, S. et al. Glycosaminoglycan-binding hydrogels enable mechanical control of human pluripotent stem cell self-renewal. ACS Nano 6, 10168–10177 (2012).
Musah, S. et al. Substratum-induced differentiation of human pluripotent stem cells reveals the coactivator YAP is a potent regulator of neuronal specification. Proc. Natl Acad. Sci. USA 111, 13805–13810 (2014).
Li, D. et al. Role of mechanical factors in fate decisions of stem cells. Regen. Med. 6, 229–240 (2011).
Kanasaki, K. et al. Integrin beta1-mediated matrix assembly and signaling are critical for the normal development and function of the kidney glomerulus. Dev. Biol. 313, 584–593 (2008).
Pozzi, A. et al. β1 integrin expression by podocytes is required to maintain glomerular structural integrity. Dev. Biol. 316, 288–301 (2008).
Rodin, S. et al. Long-term self-renewal of human pluripotent stem cells on human recombinant laminin-511. Nat. Biotechnol. 28, 611–615 (2010).
Miyazaki, T. et al. Laminin E8 fragments support efficient adhesion and expansion of dissociated human pluripotent stem cells. Nat. Commun. 3, 1236 (2012).
Mae, S.-I. et al. Monitoring and robust induction of nephrogenic intermediate mesoderm from human pluripotent stem cells. Nat. Commun. 4, 1367 (2013).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015).
Morizane, R. et al. Nephron organoids derived from human pluripotent stem cells model kidney development and injury. Nat. Biotechnol. 33, 1193–1200 (2015).
Kriz, W., Hähnel, B., Rösener, S. & Elger, M. Long-term treatment of rats with FGF-2 results in focal segmental glomerulosclerosis. Kidney Int. 48, 1435–1450 (1995).
Floege, J. et al. Visceral glomerular epithelial cells can proliferate in vivo and synthesize platelet-derived growth factor B-chain. Am. J. Pathol. 142, 637–650 (1993).
Takeuchi, A. et al. Basic fibroblast growth factor promotes proliferation of rat glomerular visceral epithelial cells in vitro. Am. J. Pathol. 141, 107–116 (1992).
Shirato, I. Podocyte process effacement in vivo. Microsc. Res. Techn. 57, 241–246 (2002).
Huh, D. et al. Microfabrication of human organs-on-chips. Nat. Protoc. 8, 2135–2157 (2013).
Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010).
Peti-Peterdi, J., Kidokoro, K. & Riquier-Brison, A. Novel in vivo techniques to visualize kidney anatomy and function. Kidney Int. 88, 44–51 (2015).
Hendry, C. et al. Direct transcriptional reprogramming of adult cells to embryonic nephron progenitors. J. Am. Soc. Nephrol. 24, 1424–1434 (2013).
Lam, A. Q. et al. Rapid and efficient differentiation of human pluripotent stem cells into intermediate mesoderm that forms tubules expressing kidney proximal tubular markers. J. Am. Soc. Nephrol. 25, 1211–1225 (2014).
Morizane, R. & Bonventre, J. V. Generation of nephron progenitor cells and kidney organoids from human pluripotent stem cells. Nat. Protoc. 12, 195–207 (2017).
Takasato, M. et al. Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nat. Cell Biol. 16, 118–126 (2014).
Reiser, J. & Sever, S. Podocyte biology and pathogenesis of kidney disease. Annu. Rev. Med. 64, 357–366 (2013).
Büscher, A. & Weber, S. Educational paper: the podocytopathies. Eur. J. Pediatr. 171, 1151–1160 (2012).
Friedman, D. & Pollak, M. Genetics of kidney failure and the evolving story of APOL1. J. Clin. Invest. 121, 3367–3374 (2011).
Devuyst, O., Knoers, N. V., Remuzzi, G. & Schaefer, F. Rare inherited kidney diseases: challenges, opportunities, and perspectives. Lancet 383, 1844–1859 (2014).
Edwards, J. K. Glomerular disease: novel candidate genes implicated in FSGS. Nat. Rev. Nephrol. 12, 256 (2016).
Greek, R. & Menache, A. Systematic reviews of animal models: methodology versus epistemology. Int. J. Med. Sci. 10, 206–221 (2013).
Yang, L., Yang, J., Byrne, S., Pan, J. & Church, G. CRISPR/Cas9‐directed genome editing of cultured cells. Curr. Protoc. Mol. Biol. 31.1.1–31.1.17 (2014).
Jang, K.-J. et al. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol. 5, 1119–1129 (2013).
Nieskens, T. & Wilmer, M. Kidney-on-a-chip technology for renal proximal tubule tissue reconstruction. Eur. J. Pharmacol. 790, 46–56 (2016).
Benam, K. H. et al. Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro. Nat. Methods 13, 151–157 (2016).
Maschmeyer, I. et al. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab Chip 15, 2688–2699 (2015).
Chang, S. Y., Weber, E. J., Ness, K. V., Eaton, D. L. & Kelly, E. J. Liver and kidney on chips: microphysiological models to understand transporter function. Clin. Pharmacol. Ther. 100, 464–478 (2016).
Materne, E.-M. et al. The multi-organ chip–a microfluidic platform for long-term multi-tissue coculture. J. Vis. Exp. (98), e52526 (2015).
Kim, H., Li, H., Collins, J. & Ingber, D. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc. Natl Acad. Sci. USA 113, E7–15 (2016).
Torisawa, Y. et al. Bone marrow-on-a-chip replicates hematopoietic niche physiology in vitro. Nat. Methods 11, 663–669 (2014).
Ingber, D. Reverse engineering human pathophysiology with organs-on-chips. Cell 164, 1105–1109 (2016).
Bhatia, S. & Ingber, D. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
Prantil-Baun, R. et al. Physiologically-based pharmacokinetic and pharmacodynamic analysis enabled by microfluidically linked organs-on-chips. Annu. Rev. Pharmacol. Toxicol. 58, 37–64 (2018).
Des Rochers, T. M. et al. Bio-engineered human kidney for the study of nephrotoxicity and kidney disease. J. Tissue Eng. Regen. Med. 6, 1–429 (2012).
Miller, J. et al. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat. Mater. 11, 768–774 (2012).
Kolesky, D. et al. 3D bioprinting of vascularized, heterogeneous cell‐laden tissue constructs. Adv. Mater. 26, 3124–3130 (2014).
Chung, H., Ko, I., Atala, A. & Yoo, J. Cell-based therapy for kidney disease. Korean J. Urol. 56, 412–421 (2015).
Ni, L., Saleem, M. & Mathieson, P. Podocyte culture: tricks of the trade. Nephrology 17, 525–531 (2012).
Saleem, M. A. One hundred ways to kill a podocyte. Nephrol. Dial. Transplant. 30, 1266–1271 (2015).
Shankland, S. J., Pippin, J. W., Reiser, J. & Mundel, P. Podocytes in culture: past, present, and future. Kidney Int. 72, 26–36 (2007).
Song, B. et al. The directed differentiation of human iPS cells into kidney podocytes. PLoS ONE 7, e46453 (2012).
Ciampi, O. et al. Generation of functional podocytes from human induced pluripotent stem cells. Stem Cell Res. 17, 130–139 (2016).
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011).
Sances, S. et al. Modeling ALS with motor neurons derived from human induced pluripotent stem cells. Nat. Neurosci. 19, 542–553 (2016).
Johnson, A., Weick, J., Pearce, R. & Zhang, S.-C. Functional neural development from human embryonic stem cells: accelerated synaptic activity via astrocyte coculture. J. Neurosci. 27, 3069–3077 (2007).
Busskamp, V. et al. Rapid neurogenesis through transcriptional activation in human stem cells. Mol. Syst. Biol. 10, 760 (2014).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Murphy, W., McDevitt, T. & Engler, A. Materials as stem cell regulators. Nat. Mater. 13, 547–557 (2014).
Mammoto, T. & Ingber, D. Mechanical control of tissue and organ development. Development 137, 1407–1420 (2010).
Borysiak, M. et al. Simple replica micromolding of biocompatible styrenic elastomers. Lab Chip 13, 2773–2784 (2013).
Domansky, K. et al. Clear castable polyurethane elastomer for fabrication of microfluidic devices. Lab Chip 13, 3956–3964 (2013).
Rayat, C. S., Joshi, K., Sakhuja, V. & Datta, U. Glomerular basement membrane thickness in normal adults and its application to the diagnosis of thin basement membrane disease: an Indian study. Indian J. Pathol. Microbiol. 48, 453–458 (2005).
Puleo, C., Ambrose, W., Takezawa, T., Elisseeff, J. & Wang, T.-H. Integration and application of vitrified collagen in multilayered microfluidic devices for corneal microtissue culture. Lab Chip 9, 3221–3227 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schneider, C., Rasband, W. & Eliceiri, K. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Abrahamson, D. Role of the podocyte (and glomerular endothelium) in building the GBM. Semin. Nephrol. 32, 342–349 (2012).
Abrahamson, D., Hudson, B., Stroganova, L., Borza, D.-B. & John, P. Cellular origins of type IV collagen networks in developing glomeruli. J. Am. Soc. Nephrol. 20, 1471–1479 (2009).
Miner, J. Organogenesis of the kidney glomerulus: focus on the glomerular basement membrane. Organogenesis 7, 75–82 (2011).
Acknowledgements
This work was supported by the Defense Advanced Research Projects Agency under Cooperative Agreement No. W911NF-12-2-0036 and the Wyss Institute for Biologically Inspired Engineering at Harvard University. S.M. was supported by a Dean’s Postdoctoral Fellowship from Harvard Medical School, a UNCF-Merck Postdoctoral Fellowship, a Postdoctoral Enrichment Program Award from the Burroughs Wellcome Fund, and an NIH/NIDDK Nephrology Training Grant (4T32DK007199-39). We thank the Wyss Institute Microfabrication Team for engineering the microfluidic devices, S. Jeanty for providing photographs of the setup for microfluidic Organ Chip culture, P.K. Tetteh for helpful suggestions, and O. Levy and R. Prantil-Baun for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
S.M., G.M.C., and D.E.I. conceived the strategy for this study; S.M. designed and performed the experiments; S.M. and D.E.I. wrote the manuscript; N.D. and D.M.C. independently analyzed the microarray data; S.M. interpreted the results; and N.D. generated heatmaps and corresponding statistical datasets. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.E.I. and S.M. declare that they are authors on a patent pending for methods for the generation of kidney glomerular podocytes from pluripotent stem cells (US patent application 14/950859). D.E.I. declares that he is a founder, holds equity and chairs the scientific advisory board at Emulate Inc. The remaining authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
1. Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip: https://doi.org/10.1038/s41551-017-0069
2. Monitoring and robust induction of nephron intermediate mesoderm from human pluripotent stem cells: https://doi.org/10.1038/ncomms2378
3. Microfabrication of human organs-on-chips: https://doi.org/10.1038/nprot.2013.137
Integrated supplementary information
Supplementary Figure 1 Expression of integrin subtypes in human iPS cells and an established human glomerular podocyte cell line.
Expression of (a) alpha and (b) beta integrin subtypes in human iPS cells and the immortalized human podocyte cell line (PCL) determined by their ability to bind to surfaces presenting antibodies specific for the indicated integrin receptors. Error bars denote standard deviation of the mean (n=3), and open circles represent individual data points. Immunofluorescence staining of β1 integrin in (c) human iPS cells and (d) PCL. Scale bars, 100 µm
Supplementary Figure 2 Heatmap of top 300 variant genes.
Heatmap of top 300 variant genes between triplicate samples of undifferentiated human iPS cells, human iPS-derived podocytes (hiPS-podocytes) and the immortalized human podocyte cell line (PCL). Each replicate represents an independent experiment. Red color indicate higher expression Z-score. Hierarchical clustering was performed using complete agglomeration method and a Euclidean distance metric
Supplementary information
Combined Supplementary Information
Supplementary Figures 1 and 2 and Supplementary Methods
Supplementary Dataset 1
Lineage characterization of genes up- or downregulated in human iPS-cell–derived podocytes, relative to expression in undifferentiated human iPS cells (PGP1 line)
Supplementary Dataset 2
Lineage characterization of genes up or downregulated in human iPS-cell–derived podocytes, relative to expression in the human immortalized podocyte cell line (PCL)
Rights and permissions
About this article
Cite this article
Musah, S., Dimitrakakis, N., Camacho, D.M. et al. Directed differentiation of human induced pluripotent stem cells into mature kidney podocytes and establishment of a Glomerulus Chip. Nat Protoc 13, 1662–1685 (2018). https://doi.org/10.1038/s41596-018-0007-8
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0007-8
This article is cited by
-
Urine-derived podocytes from steroid resistant nephrotic syndrome patients as a model for renal-progenitor derived extracellular vesicles effect and drug screening
Journal of Translational Medicine (2024)
-
The potential of organoids in renal cell carcinoma research
BMC Urology (2024)
-
Role of biophysics and mechanobiology in podocyte physiology
Nature Reviews Nephrology (2024)
-
Advances and challenges in organ-on-chip technology: toward mimicking human physiology and disease in vitro
Medical & Biological Engineering & Computing (2024)
-
Organoids and organs-on-chips: insights into predicting the efficacy of systemic treatment in colorectal cancer
Cell Death Discovery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.